|
Size: 12867
Comment:
|
Size: 20944
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| The Brainstorm window includes a graphical batching interface. With the two tabs Process1 and Process2 in the lower part of the window, you can easily select files from the database explorer and assemble a processing pipeline. Most of the operations available in the interface can also be executed this way, including everything we've been doing with Brainstorm so far. On the other hand, some features are only available this way. It is the case for the frequency filters we will need for the pre-processing of our auditory recordings. This tutorial is a parenthesis to explain how to select files and run processes on them, the cleaning of the recordings will resume in the next tutorial. |
The Brainstorm window includes a graphical batching interface. With the two tabs Process1 and Process2 in the lower part of the window, you can select files from the database explorer and assemble a processing pipeline. Most of the operations available in the interface can also be executed this way, including everything we've been doing with Brainstorm so far. On the other hand, some features are only available this way. It is the case for the frequency filters we will need for the pre-processing of our auditory recordings. This tutorial is a parenthesis to explain how to select files and run processes, we will resume with the cleaning of the recordings in the next tutorial. |
| Line 11: | Line 11: |
| The tab '''Process1''' contains a empty box in which you can drag and drop any number of files or folders from the database explorer. The easier way to understand how it works is to try it. First thing to do is to define the files you are going to process. This is done easily by picking files or folders in the database explorer and dropping them in the empty list of the Process1 tab. 1. Drag and drop the following nodes in the Process1 list: Right/ERF (recordings), Right (condition), and Subject01 (subject) . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=files1.gif|files1.gif|class="attachment"}} * The number in the brackets next to each node represents the number of data files that where found in each of them. The node ERF "contains" only itself (1), Subject01/Right contains ERF and Std files (2), and Subject01 contains 2 conditions x 2 recordings (4). * The total number of files, ie. the sum of all those values, appears in the title of the panel "Files to process [7]". 1. The buttons on the left side allow you to select what type of file you want to process: Recordings, sources, time-frequency, other. Now select the second button "Sources". All the counts are updated and now reflect the number of sources files that are found for each node. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=files2.gif|files2.gif|class="attachment"}} 1. If you select the third button "Time-frequency", you would see "0" everywhere because there are no time-frequency decompositions in the database yet. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=files3.gif|files3.gif|class="attachment"}} 1. Now clear the list from all the files. You may either right-click on the list (popup menu ''Clear list''), or select all the nodes (holding ''Shift ''or ''Crtl ''key) and then press the ''Delete ''key. 1. Select both files Left/ERF and Right/ERF in the tree (holding ''Ctrl ''key), and put the in Process list. We are going to apply some functions on those two files. You cannot distinguish them after they are dropped in the list, because they are both referred as "ERP". If at some point you need to know what is in the list, just leave you mouse over a node for a few seconds, and a tooltip would give you information about it. Just like in the database explorer. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=files4.gif|files4.gif|class="attachment"}} = Selecting files in the database = ''Authors: Francois Tadel'' When you have lots of files in a folder, like multiple source reconstructions of time-frequency files for each trial, it is difficult to grab just the ones you are interested in. Two mechanisms can help you: the Filter box and the process "File > Select files with tag". ==== Filter text box ==== You want to select only the averages in the data files in the protocol TutorialRaw. Drag and drop all your files there, then use the Filter search box at the bottom-right of the Brainstorm window. Type the string "Avg" to keep only the files that contain this string. {{http://neuroimage.usc.edu/brainstorm/SelectFiles?action=AttachFile&do=get&target=allFiles.gif|allFiles.gif|class="attachment"}} |
The tab '''Process1''' contains a empty box in which you can drag and drop any number of files or folders from the database explorer. The easiest way to understand how it works is to try it. * Try to drag and drop, in Process1, all the nodes you currently have in your database explorer. * You will see that it accepts all the folders and all the recordings, but not the channel files. * When you add a new node, the interface evaluates the number of files of the selected type that each of them contain. The number in the brackets next to each node represents the number of data files that were found in them. * On top of the list, a comment shows the total number of files that are currently selected. <<BR>><<BR>> {{attachment:process1_example.gif}} * The buttons on the left side allow you to select what type of file you want to process: Recordings, sources, time-frequency, other. When you select another button, all the counts are updated to reflect the number of files of the selected type that are found for each node. * Right now, if you select another file type, it would show only "0" everywhere because there are no sources or time-frequency decompositions available in the database yet. <<BR>><<BR>> {{attachment:process1_sources.gif}} * To remove files from the Process1 list: * Select the nodes to remove (holding Shift or Ctrl key) and press the '''Delete''' key. * Right-click on the list > '''Clear list''' == Filter by name == When you have lots of files in a folder, like multiple source reconstructions or time-frequency files for each trial, it is difficult to grab just the ones you are interested in. After selecting your folders in the Process1 box, you can refine the selection with the '''Filter search box''' at the bottom-right corner of the window. * The example below shows how to select the data files corresponding to the noise recordings: by typing "Noise" in the search box and selecting the option "Search file paths". We cannot perform the search "by name" because all the data files have the same name "Link to raw file".<<BR>><<BR>> {{attachment:process1_search.gif}} * Reminder: To see the file name corresponding to a node in the database, leave your mouse over it for a few seconds. You can do this both in the database explorer and the Process1 list.<<BR>><<BR>> {{attachment:process_filename.gif}} |
| Line 39: | Line 33: |
| * '''Search comments''': Look for the string in the comments of the files, ie. what is displayed in the database explorer to represent them (the .Comment field) * '''Search file names''': Look for the string in the full file names (including their relative path) * '''Select files''': Only the files that contain the string are selected * '''Exclude files''': Only the files that DO NOT contain the string are selected * '''Reset filters''': Removes the current file filters applied on Process1 and Process2 * '''Case insensitive''': Note that the search is not sensitive to case. If you try to select the constrained sources using the string "Constr" in the comments, it would return all the files, because this matches at the same time "Constr" and "Unconstr". But you can do the opposite instead: exclude the files that contain "Unconstr". ==== Process: Select files with tag ==== You can get to the exact same result by adding the process "File > Select files with tag" before the process you want to execute, to keep only a subset of the files that were placed in the Process1 list. It is less convenient in interactive mode because you don't see immediately the effect of your file filter, but it can be very useful when writing scripts. {{http://neuroimage.usc.edu/brainstorm/SelectFiles?action=AttachFile&do=get&target=selectFiles.gif|selectFiles.gif|class="attachment"}} To see the list of files that was selected by the process, you can open the report viewer (File > Report viewer). This lists the input files (all the files) and the final files (the selected files), with a summary of how many files where selected on the top. {{http://neuroimage.usc.edu/brainstorm/SelectFiles?action=AttachFile&do=get&target=selectReport.gif|selectReport.gif|class="attachment"}} ==== After ==== This process is useless if it is not followed immediately by another process that would do something with the selected files. An application can be the deletion of some files, by adding the process "File > Delete files". ==== How to control the output file names ==== If you are running two processes with different parameters but that produce exactly the same file names and file comments, you wouldn't be able to select them with this process. But immediately after calling any process, you can add the process "File > Add tag" to tag one specific set of files, so that you can re-select them easily later. Example: you run twice the time-frequency decomposition with different options on the same files, tag the files after calculating them with different tags. {{http://neuroimage.usc.edu/brainstorm/SelectFiles?action=AttachFile&do=get&target=addTag.gif|addTag.gif|class="attachment"}} == Creating a pipeline == === List of processes === 1. Click on Run. The Process selection window appears, with which you can create an analysis pipeline (ie. a list of process that are applied on the selected files one after the other). The first button in the toolbar shows the list of processed that are currently available. If you click on a menu, it's added to the list. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=pipeline1.gif|pipeline1.gif|class="attachment"}} 1. Some menus appear in grey (example: Sources > Spatial smoothing). This means that they are not meant to be applied to the type of data that you have in input, or at the end of the current pipeline. The "spatial smoothing" process may only be run on source files. 1. When you select a process, a list of options specific to this process is shown in the window. * To delete a process: Select it and press the ''Delete'' key, or the big cross in the toolbar. * With the "up arrow" and "down arrow" buttons in the toolbar, you can move up/down a process in the pipeline. 1. Now add the following processes, and set their options: * '''Pre-process > Band-pass filter''': 2Hz - 30Hz * In some processes, you can specify the type(s) of sensors on which you want to apply the process. This way you can for instance apply different filters on the EEG and the MEG, if you have both in the same files. * '''Extract > Extract time''': 40.00ms - 49.60ms, overwrite initial file * This will extract from each file a small time window around the main response peak. * Selecting the overwrite option will replace the previous file (bandpass), with the output of this process (bandpass+extract). This option is usually unselected for the first process in the list, then selected automatically. * '''Average > Average over time''': Overwrite initial file * Compute the average over this small time window. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=pipeline2.gif|pipeline2.gif|class="attachment"}} 1. Save your pipeline: Click on the last button in the toolbar > Save > New... > Type "process_avg45". === Saving/exporting a pipeline === The last button in the the toolbar offers a list of menus to save, load and export the pipelines. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=pipeline3.gif|pipeline3.gif|class="attachment"}} * '''Load''': List of processes that are saved in the user preferences * '''Load from file''': Import a pipeline from a pipeline_...mat file (previously saved with the menu "Save as Matlab matrix") * '''Save''': Save the pipeline in the user preferences, to be able to access it really fast after * '''Save as Matlab matrix''': Exports the pipeline for a Matlab structure in a .mat file. Allows different users to exchange their analysis pipelines (or a single user between different computers). * '''Generate .m script''': This option generates a human-readable Matlab script that can be re-used for other purposes or modified. |
* '''Search file paths''': Look for the string in the full file paths (including their relative path). * '''Search names''': Look for the string in the names of the files, ie. what is displayed in the database explorer to represent them (the .Comment field). * '''Search parent names''': Extends the search to the name of the parent files (applicable only to source and time-frequency files, which can depend on a data file). * '''Select files''': Only the files that contain the string are selected. * '''Exclude files''': Only the files that DO NOT contain the string are selected. * '''Reset filters''': Removes the current file filters applied on Process1 and Process2. * '''Case insensitive''': Note that the search is not sensitive to case. * '''Boolean logic''': You can combine different keywords to make a more precise search using advanced search queries. See the following section for more information. == Selecting processes == * Clear the file list and the search filters. * Select all three datasets we have linked to our protocol. <<BR>>You can select the three "link to raw file" nodes, the three folders or the entire subject node. * Click on the '''[Run]''' button at the bottom-left corner of the Process1 tab. * The '''Pipeline editor''' window appears. You can use it to create an analysis pipeline, i.e., a list of processes that are applied to the selected files one after the other. The first button in the toolbar shows the list of processes that are currently available. If you click on a menu, it's added to the list. * Some menus appear in grey. This means that they are not designed to be applied to the type of data that you have in input, or at the end of the current pipeline. * In the current example, we have a file with the type "continuous raw recordings", so we have access mostly to menus to manipulate event markers, run cleaning procedures and import data blocks. You can recognize a few operations that we executed in the previous tutorials: "Event > Read from channel" and "Event > Detect analog triggers". <<BR>><<BR>> {{attachment:process1_addprocess.gif||width="473",height="480"}} * When you select a process, a list of options specific to this process is shown in the window. * To delete a process: Select it and press the '''Delete''' key, or use the '''[X]''' button in the toolbar. * After selecting a first process, you can add another one. The output of the first process will be passed to the second process without giving back the control to the user. This is how you can build a full analysis pipeline with the interface. * After adding a few processes, you can move a process up or down in the pipeline with the [up arrow] and [down arrow] buttons in the toolbar. Click on a process in the pipeline to edit its options. * Select and delete a few processes to understand how this interface works. Just do not click on RUN. == Plugin structure == All the menus available in the pipeline editor are actually plugins for Brainstorm. The processes are functions that are independent from each other and automatically detected when starting Brainstorm. Any Matlab script that is added to the plugin folder (brainstorm3/toolbox/process/functions/) and has the right format will automatically be detected and made available in the GUI. This mechanism makes it easy for external contributors to develop their own code and integrate it in the interface. More information: [[Tutorials/TutUserProcess|How to write your own process]] To see where the function corresponding to a process is on the hard drive: select the process in the pipeline editor, then leave your mouse for a few seconds over its title. . {{attachment:process_path.gif||width="387",height="227"}} == Note for beginners == Everything below is advanced documentation, you can skip it for now. <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?skip=Tutorials/ArtifactsFilter")>> <<TAG(Advanced)>> == Search Database == Sometimes when working with huge protocols, you can get lost in the size of your database tree. While filtering from the process box as introduced in the previous section is one way to select the files you are looking for, we have introduced a more straightforward approach to search for file(s) in your database. At the right below the protocol selection dropdown, you can click on the magnifying glass to open up the search dialog. . {{attachment:search_new.png}} From there, you can create a new search query from the GUI, or type / paste an existing search query string (see the following section for more details). Let's select "New Search" to create a new query from the GUI. . {{attachment:search_dialog.png}} From this menu, you can create a search query to apply on your active protocol. It has different options: * '''Search by''': The file metadata to use for the search. * Name: Name of the file in Brainstorm * File type: Type of the file, see dropdown when selected for possible values * File path: Path of the file in the Brainstorm database folder * Parent name: Name of any parent file in the database tree (e.g. Subject or Folder) * '''Equality''': Type of equality to apply. * Contains: File metadata contains the entered value * Contains (case): Same as contains, but case sensitive * Equals: Exact equality, the file metadata is equal to the entered value * Equals (case): Same as equals, but case sensitive * '''Not''': Whether to invert the selected equality, e.g. DOES NOT CONTAIN vs CONTAINS. * '''Search for''': The value to search for. * '''Remove''': To remove the search row if not needed anymore. * '''+ and''': To add a search row, with the AND boolean logic. If you have two rows A and B, then the returned files will match both search A and B. * '''+ or''': To add a search row, with the OR boolean logic. If you have two rows A and B, then the returned files will match both search A or B. In the above example, we are looking for raw files (File type = Raw data) whose parent name contains the word "noise". This allows us to search for raw noise recordings. . {{attachment:search_noise_result.png}} Notice that you now have multiple tabs in your Brainstorm database. The "Database" tab contains all files in your protocol, whereas the "noise" tab only contains the files that pass the search and their parents. You can have multiple searches/tabs active so that you can easily create pipelines by dragging and dropping different search results in the process box. Do keep in mind that if you drag and drop a parent object in the process box (e.g. Subject01) with an active search, __only files that pass the active search will be processed by the pipeline__. Once a search is created, you can interact with it in different ways. You can right click on the tab and Edit the search on the fly from the GUI, Copy the search to clipboard as a query string to use it in a script, or Close the search. . {{attachment:search_right_click_tab.png}} You can also click on the magnifying glass when a search is active to get more options such as Saving the search for later use and Generating a process call to apply this search in a script. . {{attachment:search_edit_options.png}} If you click Generate process call, a line of script will be generated for you to use your search query as a process in a script. It will also be copied to clipboard. . {{attachment:search_process.png}} Notice that your search was created to a query string: . ([parent CONTAINS "noise"] AND [type EQUALS "RawData"]) This advanced query syntax is described in the following section. === Advanced search queries === For advanced users, you can write more complex search queries that can combine multiple keywords and types of keywords using boolean logic. You can do this using the Brainstorm search GUI and then copy your search as text to re-use later. These queries work for both database searches and process filters. The syntax is rigid such that the order of the commands is important, so we recommend you use the search GUI whenever possible to avoid errors. Search queries can contain the following types of elements: * '''Search parameters''': These are simple searches that are on a specific type of value. They need to be written in [square brackets]. They look like the following: . [searchFor EQUALITY NOT "value"] * __SearchFor__: Which field of the files metadata to search for It can have the following values, in lower case: * Name: Searches using the file name in Brainstorm * Type: Searches using the file type in Brainstorm * Path: Searches using the file path in the Brainstorm database folder * Parent: Searches using the parents name in the Brainstorm database tree * __Equality__: The type of equality you want to use to compare the file value to the searched value. It can have the following values, in upper case: * CONTAINS: Whether the searchFor field contains the text "value" * CONTAINS_CASE: Same as CONTAINS, but case sensitive * EQUALS: Whether the searchFor field exactly equals the text "value" * EQUALS_CASE: Same as EQUALS, but case sensitive * __NOT__: (optional) add this reserved keyword to return the opposite results of the search, so for example, all files that do NOT CONTAIN the text "value". * __"value"__: the text you want to search for, in double quotes. * '''Boolean operators''': These are used to group together search parameters and search blocks using boolean logic. Considering search parameters a, b and c, the following will return files that pass searches a and a, or does not pass search c: . (a AND b) OR NOT c * '''AND''': This combines search parameters and blocks such that both conditions have to be met. * '''OR''': This combines search parameters and blocks such that either conditions have to be met * '''NOT''': This precedes a search block or parameter such that the condition result is reversed. So if a condition had to be met, it now has to not be met. * '''Important note''': AND and OR operators cannot be mixed together (you cannot have both in the same search block), because otherwise it creates uncertainties. * '''Search blocks''': These are combinations of search parameters and boolean operators, wrapped in (round brackets). You cannot have different boolean operators in the same block ==== Example ==== (([name CONTAINS "test1"] AND [type EQUALS "Matrix"]) OR NOT [parent CONTAINS "test2"]) __Effect__: This will match all matrix files containing text "test1" or all files whose parent docontains the text "test2". ==== Limitations of the GUI ==== The GUI does not support multiple nested search blocks. It only allows for one OR block followed by one AND block. If your query is more advanced than this, you will not be able to edit it with the search GUI. We recommend you use the process filter box instead. <<TAG(Advanced)>> == Saving a pipeline == After preparing your analysis pipeline by listing all the operations to run on your input files, you can either click on the [Run] button, or save/export your pipeline. The last button in the the toolbar offers a list of menus to save, load and export the pipelines. . {{attachment:pipeline_example.gif||width="334",height="340"}} * '''Load''': List of pipelines that are saved in the user preferences on this computer. * '''Load from .mat file''': Import a pipeline from a pipeline_...mat file. * '''Save''': Save the pipeline in the user preferences. * '''Save as .mat matrix''': Exports the pipeline as Matlab structure in a .mat file. Allows different users to exchange their analysis pipelines, or a single user between different computers. * '''Generate .m script''': This option generates a Matlab script. |
| Line 97: | Line 173: |
| * '''Reset options''': Brainstorm saves automatically for each user the options of all the processes. This menu removes all the saved options, and set them back to the default values. Here is the Matlab script that is generated automatically for this pipeline. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=script.gif|script.gif|class="attachment"}} . Click on Ok, in the pipeline window. After a few seconds, you will see two new files in the database, and the "Report viewer" window. === Report viewer === Each time the pipeline editor is used to executed to run a list of processes, a report is generated and saved in the user home folder (/home/username/reports/). The report viewer shows as an HTML page some of the information saved in this report structure: the date and duration of execution, the list of processes, the input and output files. It reports all the warning and errors that happen during the execution. The report viewer does not necessarily appear automatically at the end of the last process: it is shown only when more than one processes were executed, or when any of the processes returned an error or a warning. When running processes manually from a script, the calls bst_report(Start, Save, Open) explicitly indicate when the logging of the events should start and stop. You can add images to the reports for quality control using the process "File > Save snapshot". {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutProcesses?action=AttachFile&do=get&target=report.gif|output.gif|class="attachment"}} After you close the report window, you can re-open the last report with the main menu of the Brainstorm window: '''File > Report viewer'''. |
* '''Reset options''': Brainstorm automatically saves the options of all the processes in the user preferences. This menu removes all the saved options and sets them back to the default values. <<TAG(Advanced)>> == Automatic script generation == Here is the Matlab script that is generated for this pipeline. {{attachment:pipeline_script.gif||width="703",height="509"}} Reading this script is easy: input files at the top, one block per process, one line per option. You can also modify them to add personal code, loops or tests. Many features are still missing in the pipeline editor, but the generated scripts are easy enough for users with basic Matlab knowledge to edit and improve them. Running this script from Matlab or clicking on the [Run] button of the pipeline editor produce exactly the same results. In both cases you will not have any interaction with the script, it could be executed without any direct supervision. You just get a report in the end that describes everything that happened during the execution. These scripts cannot be reloaded in the pipeline editor window after being generated. If you work on a long analysis pipeline, save it in your user preferences before generating the corresponding Matlab script. <<TAG(Advanced)>> == Process: Select files with tag == Since we are discussing the file selection and the pipeline execution, we can explore a few more available options. We have seen how to filter the files in the Process1 box using the Filter search box. We can get to the exact same result by using the process '''File > Select files: By tag''' before the process you want to execute, to keep only a subset of the files that were placed in the Process1 list. It is less convenient in interactive mode because you don't immediately see the effect of your file filter, but it can be very useful when writing scripts. You can also combine search constraints by adding the same process multiple times in your pipeline, which is not possible with the search box. * Make sure you still have the three datasets selected in the Process1 list. * Select the process: '''File > Select files: By tag''' * Select the options: Search: "Noise", Search the file names, Select only the files with the tag. * Click on [Run] to execute the process. <<BR>><<BR>> {{attachment:process1_select.gif}} * This process is useless if not followed immediately by another process that does something with the selected files. It does nothing but selecting the file, but we can observe that the operation was actually executed with the report viewer. <<TAG(Advanced)>> == Report viewer == Everytime the pipeline editor is used to run a list of processes, a report is created and logs all the messages that are generated during the execution. These reports are saved in the user home folder: $HOME/.brainstorm/reports/. The report viewer shows, as an HTML page, some of the information saved in this report structure: the date and duration of execution, the list of processes, and the input and output files. It reports all the warnings and errors that occurred during the execution. The report is displayed at the end of the execution only if there were more than one process executed, or if an error or a warning was reported. In this case, nothing is displayed. You can always explicitly open the report viewer to show the last reports: '''File > Report viewer'''. {{attachment:reports.gif||width="711",height="379"}} When running processes manually from a script, the calls to '''bst_report''' explicitly indicate when the logging of the events should start and stop. You can add images to the reports for quality control using the process '''File > Save snapshot'''. |
| Line 121: | Line 220: |
| PROCESSES AND PLUG-INS The available processes are organized in a plug-in structure. Any Matlab script that is added to the plug-in folder (brainstorm3/toolbox/process/functions/) and has the right format will be automatically detected and made available in the GUI. This mechanism makes the contribution from other developers to Brainstorm very easy. REPORT: OTHER TUTORIALS |
More information: [[Tutorials/Scripting|Complete tutorial script]] <<TAG(Advanced)>> == Error management == * Select the same files and same process: '''File > Select files: By tag''' * Note that the options you used during the previous call are now selected by default. * Instead of "Noise", now search for a string that doesn't exist in the file name, such as "XXXX". <<BR>><<BR>> {{attachment:search_xxxx.gif}} * Click on [Run] to execute the process. You will get the following error. <<BR>><<BR>> {{attachment:search_error.gif||width="253",height="129"}} * If you open the report viewer, it should look like this. <<BR>><<BR>> {{attachment:search_report.gif||width="512",height="283"}} <<TAG(Advanced)>> == Control the output file names == If you are running two processes with different parameters but that produce exactly the same file paths and file names, you wouldn't be able to select them with this process. But immediately after calling any process, you can add the process '''File > Add tag '''to tag one specific set of files, so that you can easily re-select them later. Example: You run the time-frequency decomposition twice with different options on the same files, tag the files after calculating them with different tags. . {{attachment:addTag.gif}} == Additional documentation == * Tutorial: [[Tutorials/Scripting|Scripting]] * Tutorial: [[Tutorials/TutUserProcess|How to write your own process]] <<HTML(<!-- END-PAGE -->)>> |
Tutorial 9: Select files and run processes
Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet
The Brainstorm window includes a graphical batching interface. With the two tabs Process1 and Process2 in the lower part of the window, you can select files from the database explorer and assemble a processing pipeline. Most of the operations available in the interface can also be executed this way, including everything we've been doing with Brainstorm so far.
On the other hand, some features are only available this way. It is the case for the frequency filters we will need for the pre-processing of our auditory recordings. This tutorial is a parenthesis to explain how to select files and run processes, we will resume with the cleaning of the recordings in the next tutorial.
Contents
Selecting files to process
The tab Process1 contains a empty box in which you can drag and drop any number of files or folders from the database explorer. The easiest way to understand how it works is to try it.
- Try to drag and drop, in Process1, all the nodes you currently have in your database explorer.
- You will see that it accepts all the folders and all the recordings, but not the channel files.
- When you add a new node, the interface evaluates the number of files of the selected type that each of them contain. The number in the brackets next to each node represents the number of data files that were found in them.
On top of the list, a comment shows the total number of files that are currently selected.
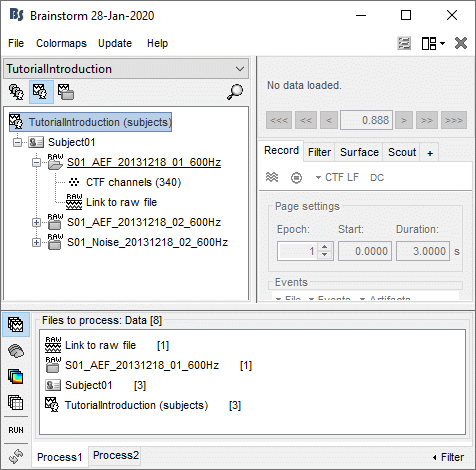
- The buttons on the left side allow you to select what type of file you want to process: Recordings, sources, time-frequency, other. When you select another button, all the counts are updated to reflect the number of files of the selected type that are found for each node.
Right now, if you select another file type, it would show only "0" everywhere because there are no sources or time-frequency decompositions available in the database yet.
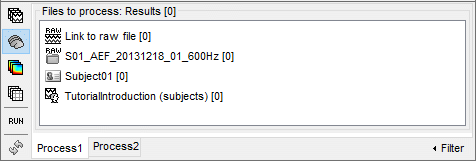
- To remove files from the Process1 list:
Select the nodes to remove (holding Shift or Ctrl key) and press the Delete key.
Right-click on the list > Clear list
Filter by name
When you have lots of files in a folder, like multiple source reconstructions or time-frequency files for each trial, it is difficult to grab just the ones you are interested in. After selecting your folders in the Process1 box, you can refine the selection with the Filter search box at the bottom-right corner of the window.
The example below shows how to select the data files corresponding to the noise recordings: by typing "Noise" in the search box and selecting the option "Search file paths". We cannot perform the search "by name" because all the data files have the same name "Link to raw file".
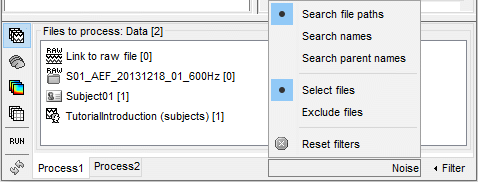
Reminder: To see the file name corresponding to a node in the database, leave your mouse over it for a few seconds. You can do this both in the database explorer and the Process1 list.
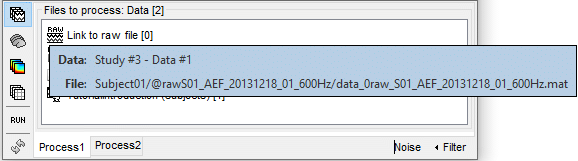
The options offered in the Filter menu are:
Search file paths: Look for the string in the full file paths (including their relative path).
Search names: Look for the string in the names of the files, ie. what is displayed in the database explorer to represent them (the .Comment field).
Search parent names: Extends the search to the name of the parent files (applicable only to source and time-frequency files, which can depend on a data file).
Select files: Only the files that contain the string are selected.
Exclude files: Only the files that DO NOT contain the string are selected.
Reset filters: Removes the current file filters applied on Process1 and Process2.
Case insensitive: Note that the search is not sensitive to case.
Boolean logic: You can combine different keywords to make a more precise search using advanced search queries. See the following section for more information.
Selecting processes
- Clear the file list and the search filters.
Select all three datasets we have linked to our protocol.
You can select the three "link to raw file" nodes, the three folders or the entire subject node.Click on the [Run] button at the bottom-left corner of the Process1 tab.
The Pipeline editor window appears. You can use it to create an analysis pipeline, i.e., a list of processes that are applied to the selected files one after the other. The first button in the toolbar shows the list of processes that are currently available. If you click on a menu, it's added to the list.
- Some menus appear in grey. This means that they are not designed to be applied to the type of data that you have in input, or at the end of the current pipeline.
In the current example, we have a file with the type "continuous raw recordings", so we have access mostly to menus to manipulate event markers, run cleaning procedures and import data blocks. You can recognize a few operations that we executed in the previous tutorials: "Event > Read from channel" and "Event > Detect analog triggers".
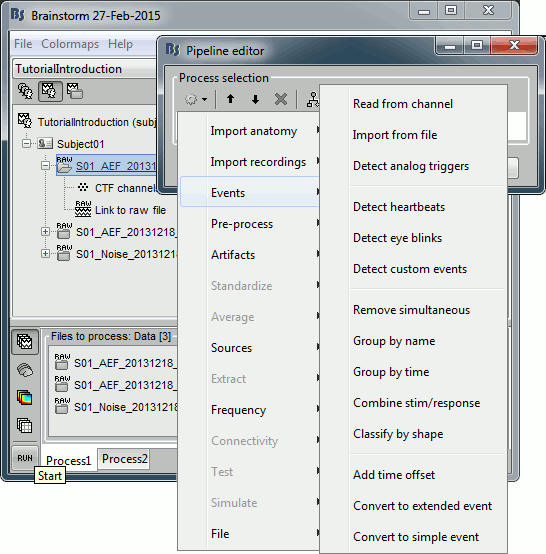
- When you select a process, a list of options specific to this process is shown in the window.
To delete a process: Select it and press the Delete key, or use the [X] button in the toolbar.
- After selecting a first process, you can add another one. The output of the first process will be passed to the second process without giving back the control to the user. This is how you can build a full analysis pipeline with the interface.
- After adding a few processes, you can move a process up or down in the pipeline with the [up arrow] and [down arrow] buttons in the toolbar. Click on a process in the pipeline to edit its options.
- Select and delete a few processes to understand how this interface works. Just do not click on RUN.
Plugin structure
All the menus available in the pipeline editor are actually plugins for Brainstorm. The processes are functions that are independent from each other and automatically detected when starting Brainstorm.
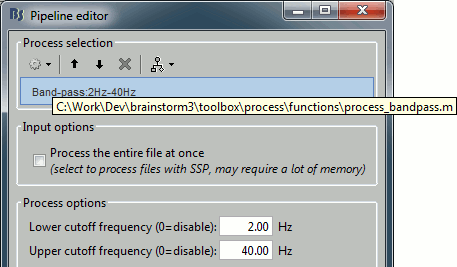
Any Matlab script that is added to the plugin folder (brainstorm3/toolbox/process/functions/) and has the right format will automatically be detected and made available in the GUI. This mechanism makes it easy for external contributors to develop their own code and integrate it in the interface.
More information: How to write your own process
To see where the function corresponding to a process is on the hard drive: select the process in the pipeline editor, then leave your mouse for a few seconds over its title.
Note for beginners
Everything below is advanced documentation, you can skip it for now.
Search Database
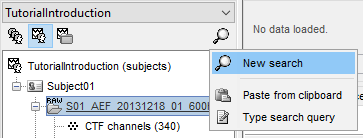
Sometimes when working with huge protocols, you can get lost in the size of your database tree. While filtering from the process box as introduced in the previous section is one way to select the files you are looking for, we have introduced a more straightforward approach to search for file(s) in your database. At the right below the protocol selection dropdown, you can click on the magnifying glass to open up the search dialog.
From there, you can create a new search query from the GUI, or type / paste an existing search query string (see the following section for more details). Let's select "New Search" to create a new query from the GUI.
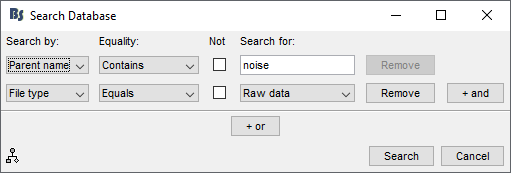
From this menu, you can create a search query to apply on your active protocol. It has different options:
Search by: The file metadata to use for the search.
- Name: Name of the file in Brainstorm
- File type: Type of the file, see dropdown when selected for possible values
- File path: Path of the file in the Brainstorm database folder
- Parent name: Name of any parent file in the database tree (e.g. Subject or Folder)
Equality: Type of equality to apply.
- Contains: File metadata contains the entered value
- Contains (case): Same as contains, but case sensitive
- Equals: Exact equality, the file metadata is equal to the entered value
- Equals (case): Same as equals, but case sensitive
Not: Whether to invert the selected equality, e.g. DOES NOT CONTAIN vs CONTAINS.
Search for: The value to search for.
Remove: To remove the search row if not needed anymore.
+ and: To add a search row, with the AND boolean logic. If you have two rows A and B, then the returned files will match both search A and B.
+ or: To add a search row, with the OR boolean logic. If you have two rows A and B, then the returned files will match both search A or B.
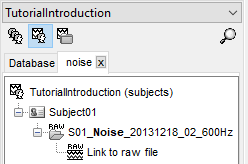
In the above example, we are looking for raw files (File type = Raw data) whose parent name contains the word "noise". This allows us to search for raw noise recordings.
Notice that you now have multiple tabs in your Brainstorm database. The "Database" tab contains all files in your protocol, whereas the "noise" tab only contains the files that pass the search and their parents. You can have multiple searches/tabs active so that you can easily create pipelines by dragging and dropping different search results in the process box. Do keep in mind that if you drag and drop a parent object in the process box (e.g. Subject01) with an active search, only files that pass the active search will be processed by the pipeline.
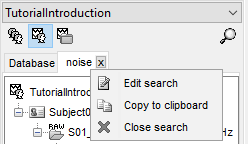
Once a search is created, you can interact with it in different ways. You can right click on the tab and Edit the search on the fly from the GUI, Copy the search to clipboard as a query string to use it in a script, or Close the search.
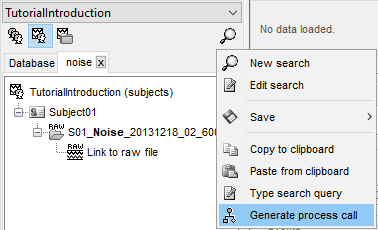
You can also click on the magnifying glass when a search is active to get more options such as Saving the search for later use and Generating a process call to apply this search in a script.
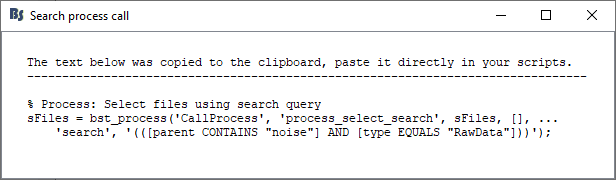
If you click Generate process call, a line of script will be generated for you to use your search query as a process in a script. It will also be copied to clipboard.
Notice that your search was created to a query string:
([parent CONTAINS "noise"] AND [type EQUALS "RawData"])
This advanced query syntax is described in the following section.
Advanced search queries
For advanced users, you can write more complex search queries that can combine multiple keywords and types of keywords using boolean logic. You can do this using the Brainstorm search GUI and then copy your search as text to re-use later. These queries work for both database searches and process filters. The syntax is rigid such that the order of the commands is important, so we recommend you use the search GUI whenever possible to avoid errors. Search queries can contain the following types of elements:
Search parameters: These are simple searches that are on a specific type of value. They need to be written in [square brackets]. They look like the following:
- [searchFor EQUALITY NOT "value"]
SearchFor: Which field of the files metadata to search for It can have the following values, in lower case:
- Name: Searches using the file name in Brainstorm
- Type: Searches using the file type in Brainstorm
- Path: Searches using the file path in the Brainstorm database folder
- Parent: Searches using the parents name in the Brainstorm database tree
Equality: The type of equality you want to use to compare the file value to the searched value. It can have the following values, in upper case:
- CONTAINS: Whether the searchFor field contains the text "value"
- CONTAINS_CASE: Same as CONTAINS, but case sensitive
- EQUALS: Whether the searchFor field exactly equals the text "value"
- EQUALS_CASE: Same as EQUALS, but case sensitive
NOT: (optional) add this reserved keyword to return the opposite results of the search, so for example, all files that do NOT CONTAIN the text "value".
"value": the text you want to search for, in double quotes.
Boolean operators: These are used to group together search parameters and search blocks using boolean logic. Considering search parameters a, b and c, the following will return files that pass searches a and a, or does not pass search c:
- (a AND b) OR NOT c
AND: This combines search parameters and blocks such that both conditions have to be met.
OR: This combines search parameters and blocks such that either conditions have to be met
NOT: This precedes a search block or parameter such that the condition result is reversed. So if a condition had to be met, it now has to not be met.
Important note: AND and OR operators cannot be mixed together (you cannot have both in the same search block), because otherwise it creates uncertainties.
Search blocks: These are combinations of search parameters and boolean operators, wrapped in (round brackets). You cannot have different boolean operators in the same block
Example
(([name CONTAINS "test1"] AND [type EQUALS "Matrix"]) OR NOT [parent CONTAINS "test2"])
Effect: This will match all matrix files containing text "test1" or all files whose parent docontains the text "test2".
Limitations of the GUI
The GUI does not support multiple nested search blocks. It only allows for one OR block followed by one AND block. If your query is more advanced than this, you will not be able to edit it with the search GUI. We recommend you use the process filter box instead.
Saving a pipeline
After preparing your analysis pipeline by listing all the operations to run on your input files, you can either click on the [Run] button, or save/export your pipeline. The last button in the the toolbar offers a list of menus to save, load and export the pipelines.
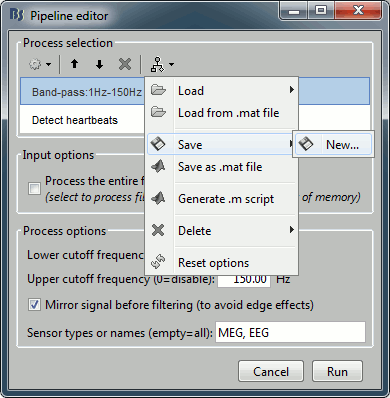
Load: List of pipelines that are saved in the user preferences on this computer.
Load from .mat file: Import a pipeline from a pipeline_...mat file.
Save: Save the pipeline in the user preferences.
Save as .mat matrix: Exports the pipeline as Matlab structure in a .mat file. Allows different users to exchange their analysis pipelines, or a single user between different computers.
Generate .m script: This option generates a Matlab script.
Delete: Remove a pipeline that is saved in the user preferences.
Reset options: Brainstorm automatically saves the options of all the processes in the user preferences. This menu removes all the saved options and sets them back to the default values.
Automatic script generation
Here is the Matlab script that is generated for this pipeline.
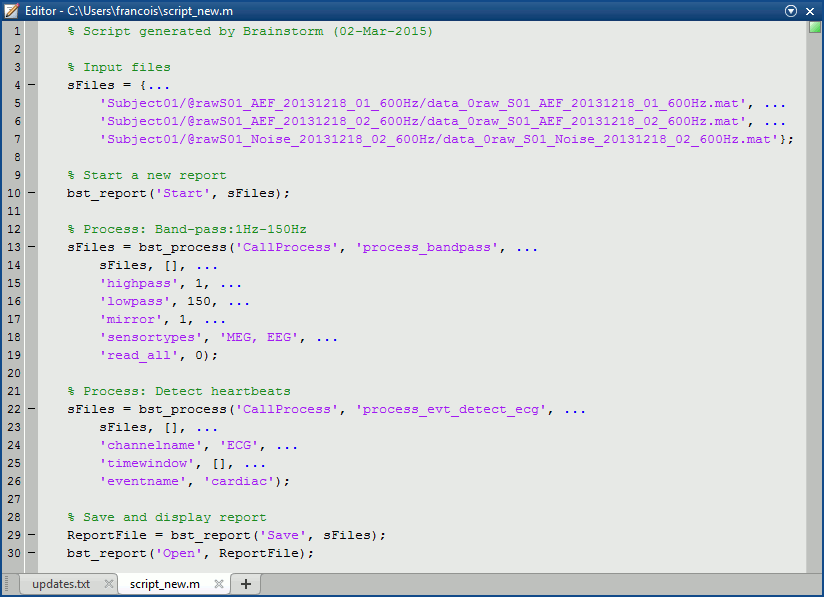
Reading this script is easy: input files at the top, one block per process, one line per option. You can also modify them to add personal code, loops or tests. Many features are still missing in the pipeline editor, but the generated scripts are easy enough for users with basic Matlab knowledge to edit and improve them.
Running this script from Matlab or clicking on the [Run] button of the pipeline editor produce exactly the same results. In both cases you will not have any interaction with the script, it could be executed without any direct supervision. You just get a report in the end that describes everything that happened during the execution.
These scripts cannot be reloaded in the pipeline editor window after being generated. If you work on a long analysis pipeline, save it in your user preferences before generating the corresponding Matlab script.
Process: Select files with tag
Since we are discussing the file selection and the pipeline execution, we can explore a few more available options. We have seen how to filter the files in the Process1 box using the Filter search box. We can get to the exact same result by using the process File > Select files: By tag before the process you want to execute, to keep only a subset of the files that were placed in the Process1 list.
It is less convenient in interactive mode because you don't immediately see the effect of your file filter, but it can be very useful when writing scripts. You can also combine search constraints by adding the same process multiple times in your pipeline, which is not possible with the search box.
- Make sure you still have the three datasets selected in the Process1 list.
Select the process: File > Select files: By tag
- Select the options: Search: "Noise", Search the file names, Select only the files with the tag.
Click on [Run] to execute the process.
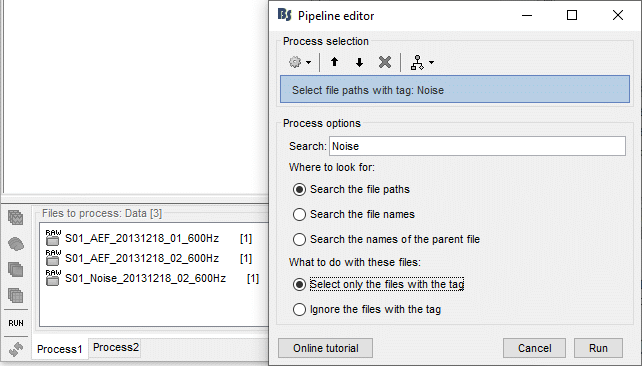
- This process is useless if not followed immediately by another process that does something with the selected files. It does nothing but selecting the file, but we can observe that the operation was actually executed with the report viewer.
Report viewer
Everytime the pipeline editor is used to run a list of processes, a report is created and logs all the messages that are generated during the execution. These reports are saved in the user home folder: $HOME/.brainstorm/reports/.
The report viewer shows, as an HTML page, some of the information saved in this report structure: the date and duration of execution, the list of processes, and the input and output files. It reports all the warnings and errors that occurred during the execution.
The report is displayed at the end of the execution only if there were more than one process executed, or if an error or a warning was reported. In this case, nothing is displayed.
You can always explicitly open the report viewer to show the last reports: File > Report viewer.
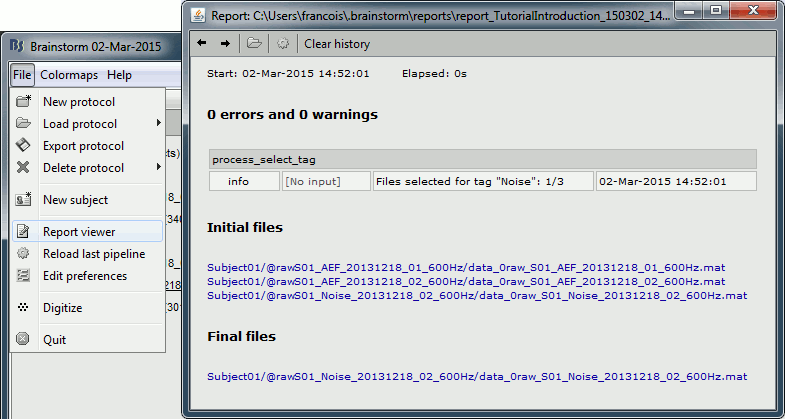
When running processes manually from a script, the calls to bst_report explicitly indicate when the logging of the events should start and stop.
You can add images to the reports for quality control using the process File > Save snapshot.
With the buttons in the toolbar, you can go back to the previous reports saved from the same protocol.
More information: Complete tutorial script
Error management
Select the same files and same process: File > Select files: By tag
- Note that the options you used during the previous call are now selected by default.
Instead of "Noise", now search for a string that doesn't exist in the file name, such as "XXXX".
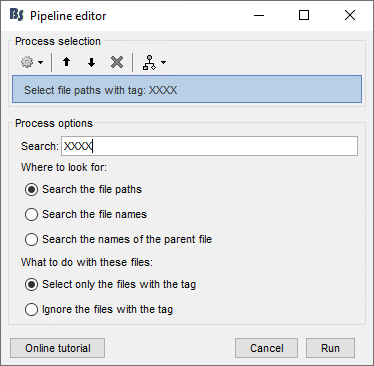
Click on [Run] to execute the process. You will get the following error.
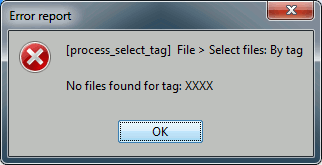
If you open the report viewer, it should look like this.
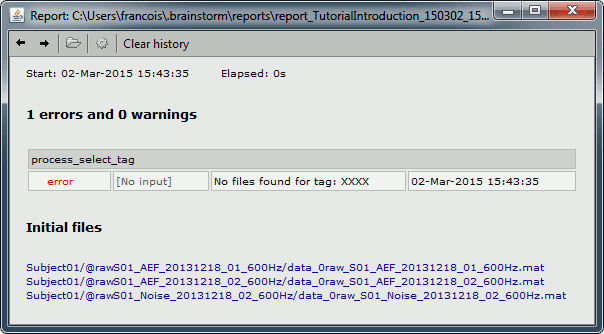
Control the output file names
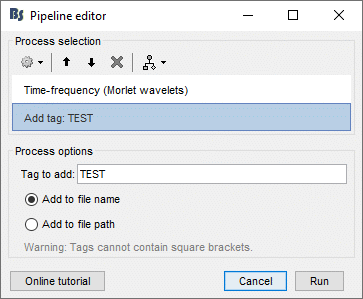
If you are running two processes with different parameters but that produce exactly the same file paths and file names, you wouldn't be able to select them with this process. But immediately after calling any process, you can add the process File > Add tag to tag one specific set of files, so that you can easily re-select them later.
Example: You run the time-frequency decomposition twice with different options on the same files, tag the files after calculating them with different tags.
Additional documentation
Tutorial: Scripting
Tutorial: How to write your own process