T1-MRI Segmentation with SPM12 / CAT12
Authors: Francois Tadel
CAT is a SPM12 toolbox that is fully interfaced with Brainstorm. It can replace efficiently FreeSurfer for generating the cortical surface from any T1 MRI. It runs on any OS in about 1 hour, instead of the typical 24hr FreeSurfer recon-all processing. The surfaces are registered to the templates with the FreeSurfer spheres, and include many anatomical and functional atlases.
You can either run the CAT segmentation from Brainstorm, or run it separately and import its outputs as you would do with FreeSurfer. The software is not packaged with Brainstorm and does not work with the compiled version of Brainstorm, you must install SPM12 and CAT separately.
Contents
Install SPM12
Download SPM12: https://www.fil.ion.ucl.ac.uk/spm/software/download/
Unzip it anywhere on your computer, but not in any of the Brainstorm folder.
- Do not add the spm12 folder to your path if you are not a SPM user, Brainstorm will do this automatically when needed.
Start Brainstorm, set the SPM12 path in the Brainstorm preferences (File > Edit preferences).
Install CAT12
Download CAT: http://www.neuro.uni-jena.de/cat/index.html#DOWNLOAD
- Unzip the downloaded file.
Move the cat12 folder to the spm12/toolbox directory
Citing CAT12
If you use CAT from Brainstorm for MRI segmentation, please cite the following article in your publications:
...
Running CAT from Brainstorm
- Switch to the anatomy side of the database explorer.
- Create a subject, import the T1 MRI for this subject.
Set the fiducial points manually (NAS/LPA/RPA) or compute the MNI transformation.
Right-click on the MRI > CAT12 MRI registration,
or use the process Import anatomy > Segment MRI with SPM12/CAT12
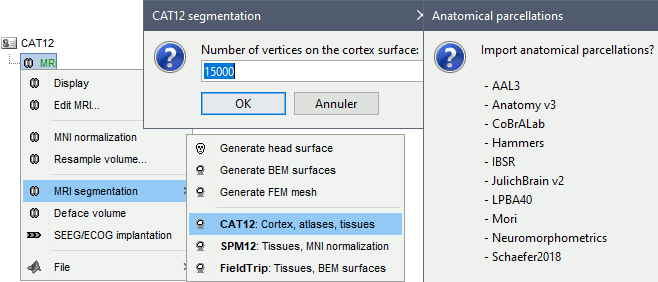
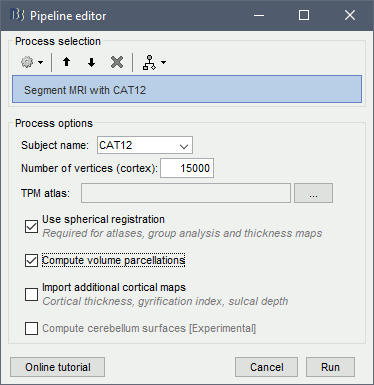
In interactive mode, you will be prompted for the number of vertices you want in the final cortex surface. This will by extension define the number of dipoles to estimate during the source estimation process. By default we set this value to 15000 for the entire brain (it means 7500 for each hemisphere).
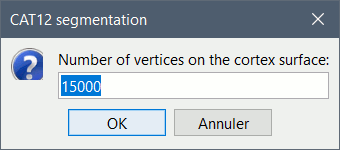
Brainstorm saves the MRI to process in a temporary folder, then calls CAT as an SPM batch:
$HOME/.brainstorm/tmp/cat12/spm_cat12.nii- All the output from CAT will be saved in the same temporary folder. At the end, Brainstorm imports the output folder as the anatomy folder for the selected subject. If the process crashes, you can inspect the contents of this folder for indices on how to solve the problem. If the segmentation and the import is successful, the temporary folder is deleted.
A report is displayed by CAT, and saved as an image and a PDF file in tmp/cat12/report.
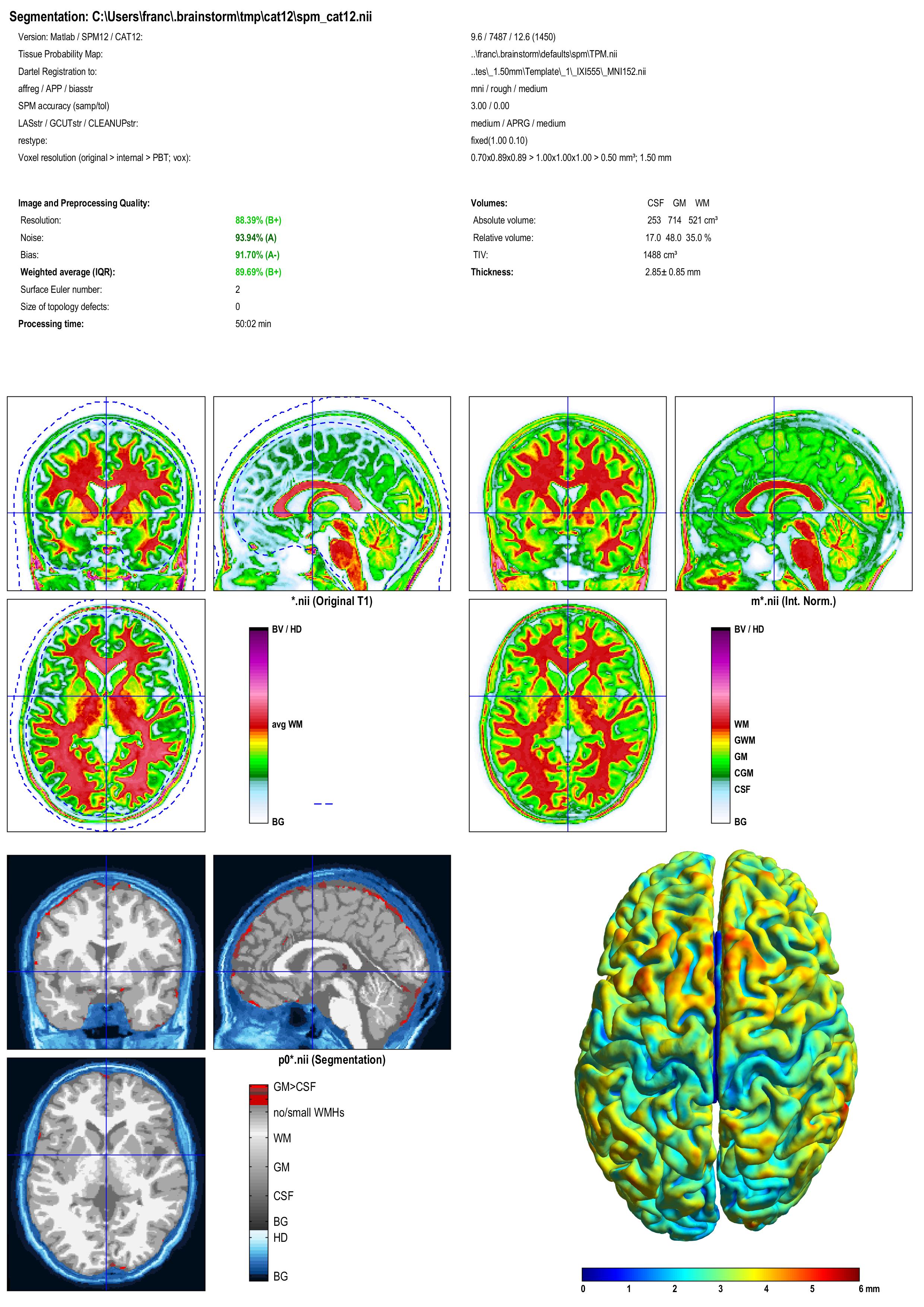
Importing the results in Brainstorm
If you run the segmentation process from Brainstorm, the import will be done automatically. Otherwise, if you need to import an existing CAT segmentation, here is the following procedure.
Right-click on the subject > Import anatomy folder.
Select the file format CAT12 folder or CAT12 folder + thickness maps.
- Enter the number of vertices in the final cortex surface.
- The files that are imported from the segmentation output folder are the following:
/*.nii (T1 MRI volume - only one .nii file allowed in the top-level folder)
/surf/?h.central.*.gii (left/right hemisphere of the central surface)
/surf/?h.sphere.reg.*.gii (left/right FreeSurfer registered spheres)
/surf/?h.thickness (texture with the cortical thickness at each vertex)
Additional atlases are imported from the CAT12 program folder (spm12/toolbox/cat12):
/templates_surfaces/?h.central.freesurfer.gii (FSAverage central surfaces, high resolution)
/templates_surfaces_32k/?h.central.freesurfer.gii (FSAverage surfaces, medium resolution)
/atlases_surfaces/?h.*.annot (cortical surface-based atlases)
/atlases_surfaces_32k/?h.*.annot (cortical surface-based atlases)
All these atlases available on the FSAverage template are projected to the subject native surfaces using the FreeSurfer registered spheres, with functions bst_project_scouts.m and tess_interp_tess2tess.m.
The central surfaces generated by CAT are meshes half-way between the grey-white interface and the external pial surface.
- The files you can see in the database explorer at the end:
MRI: The T1 MRI of the subject, imported from the .nii file at the top-level folder.
head mask (10000,0,2): Scalp surface generated by Brainstorm. The numbers indicate the parameters that were automatically used for this head: vertices=10000, erode factor=0, fill holes=2 (these are detailed later)
cortex_250000V: High-resolution cortex surface that was generated by CAT.
cortex_15000V: Low-resolution cortex surface, downsampled using the reducepatch function from Matlab (it keeps a meaningful subset of vertices from the original surface). It appears in green in the database explorer, ie. it is going to be used as the default by the processes that require a cortex surface.
A figure is automatically shown at the end of the process, to check visually that the cortex and head surfaces were properly generated and imported. If it doesn't look like the following picture, do not go any further in your source analysis, fix the anatomy first.
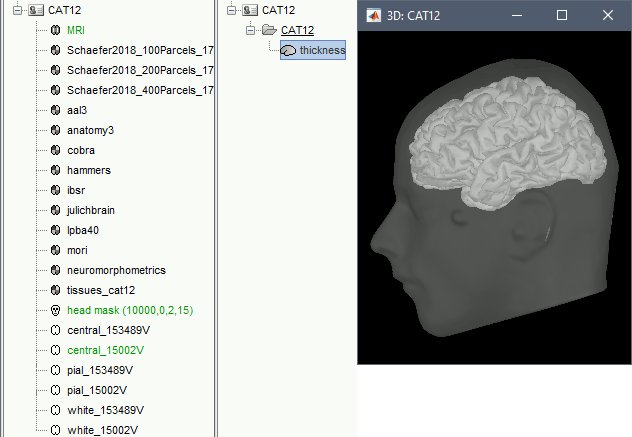
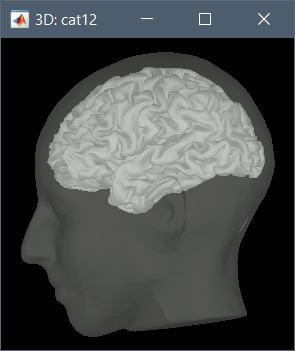
Cortical parcellations
FreeSurfer's FSAverage subject includes parcellations of the cortical surface in anatomical regions. The description of this feature is available here:
http://freesurfer.net/fswiki/CorticalParcellation
Using CAT12 from Brainstorm, you have access to 15 all the individual brains:
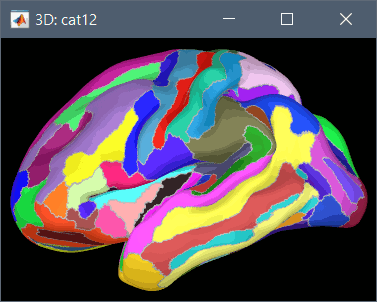
Destrieux atlas (atlases_surfaces/?h.aparc_a2009s.freesurfer.annot): more info
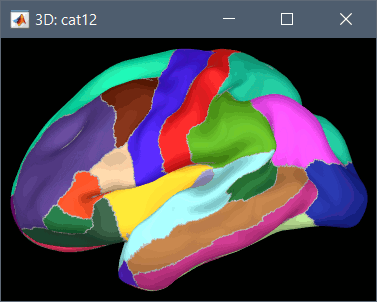
Desikan-Killiany atlas (atlases_surfaces/?h.aparc_DK40.freesurfer.annot): more info
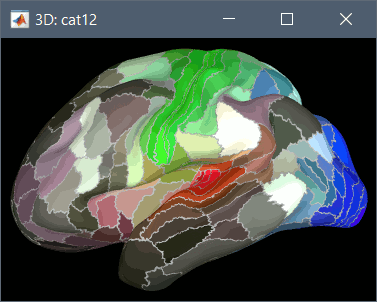
HCP MMP1 atlas (atlases_surfaces/?h.aparc_HCP_MMP1.freesurfer.annot): more info
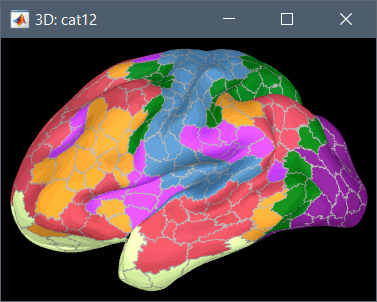
Schaefer 2018 atlases (atlases_surfaces_32k/?h.Schaefer2018_*.annot): more info
These atlases are imported in Brainstorm as scouts (cortical regions of interest), and saved directly in the surface files. To check where they are saved: right-click on the low-resolution cortex file > File > View file contents. You can see that 17 structures "Atlas" are available, the first one that has Name='User scouts', and the second one Name='Destrieux'.
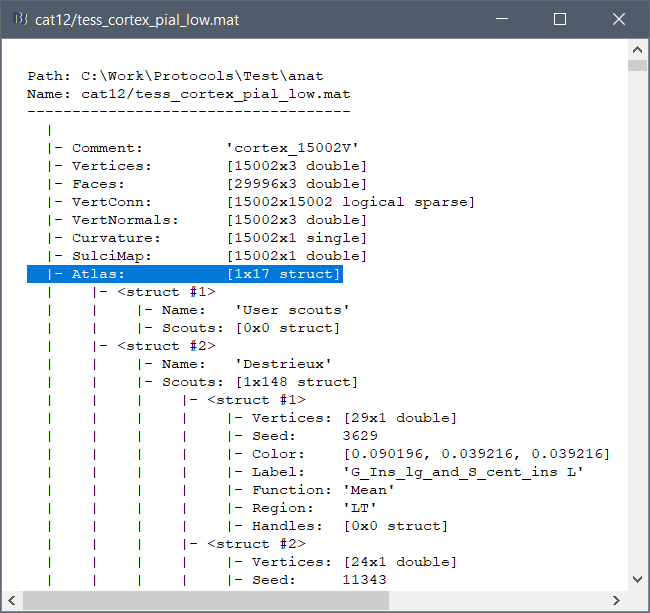
To access them from the interface: Double-click on the cortex and go to the Scout tab, and click on the drop-down list to select another Atlas (ie group of scouts):
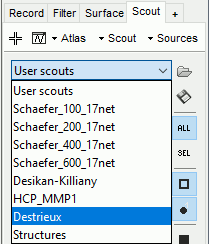
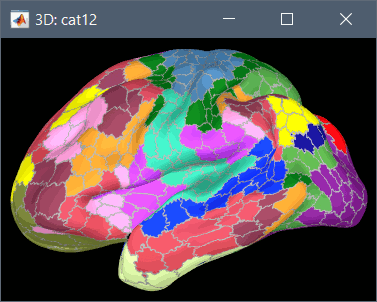
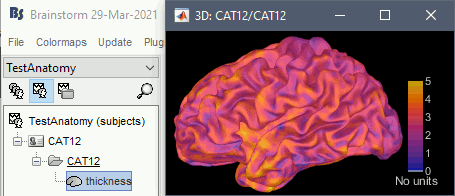
Cortical thickness
The cortical thickness can be saved as a cortical map in the database (a "results" file). This result is generated when using the file format "CAT12 folder + Thickness maps" in the Import anatomy folder selection.
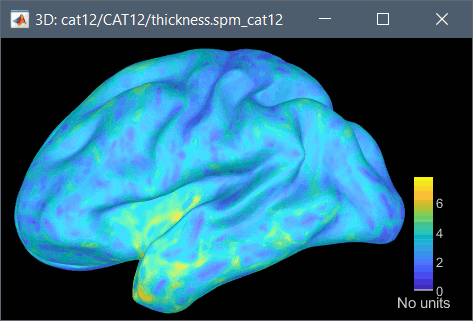
SPM batch
The CAT segmentation is executed with the following SPM12 batch:
% Create SPM batch
matlabbatch{1}.spm.tools.cat.estwrite.data = {[NiiFile ',1']};
matlabbatch{1}.spm.tools.cat.estwrite.nproc = 0;
matlabbatch{1}.spm.tools.cat.estwrite.opts.tpm = {bst_get('SpmTpmAtlas')};
matlabbatch{1}.spm.tools.cat.estwrite.opts.affreg = 'mni';
matlabbatch{1}.spm.tools.cat.estwrite.opts.biasstr = 0.5;
matlabbatch{1}.spm.tools.cat.estwrite.opts.accstr = 0.5;
matlabbatch{1}.spm.tools.cat.estwrite.extopts.APP = 1070;
matlabbatch{1}.spm.tools.cat.estwrite.extopts.LASstr = 0.5;
matlabbatch{1}.spm.tools.cat.estwrite.extopts.gcutstr = 2;
matlabbatch{1}.spm.tools.cat.estwrite.extopts.registration.dartel.darteltpm = {dartelTpm};
matlabbatch{1}.spm.tools.cat.estwrite.extopts.vox = 1.5;
matlabbatch{1}.spm.tools.cat.estwrite.extopts.restypes.fixed = [1 0.1];
matlabbatch{1}.spm.tools.cat.estwrite.output.surface = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.ROImenu.atlases.neuromorphometrics = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.ROImenu.atlases.lpba40 = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.ROImenu.atlases.cobra = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.ROImenu.atlases.hammers = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.GM.native = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.GM.mod = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.GM.dartel = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.WM.native = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.WM.mod = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.WM.dartel = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.labelnative = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.bias.warped = 1;
matlabbatch{1}.spm.tools.cat.estwrite.output.jacobianwarped = 0;
matlabbatch{1}.spm.tools.cat.estwrite.output.warps = [0 0];
% Run SPM batch
spm_jobman('initcfg');
spm_jobman('run',matlabbatch);