|
Size: 8730
Comment:
|
Size: 8104
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Tutorial: Compute a head model = This tutorial is still based on Sabine Meunier's somatotopy experiment, called ''TutorialCTF ''in your Brainstorm database. The recordings have already been imported and analyzed at the sensors level, and they are now ready for source estimation. |
'''[WARNING: This page is part of the old tutorials, please refer to the [[Tutorials|new documentation]]]'''<<BR>><<BR>><<BR>> = Tutorial 6: Computing a head model = ''Authors: Francois Tadel, John C Mosher, Richard Leahy, Sylvain Baillet'' This tutorial is still based on Sabine Meunier's somatotopy experiment, called ''TutorialCTF ''in your Brainstorm database. The recordings have already been imported and analyzed at the sensors level, they are ready for source estimation. <<TableOfContents(2,2)>> |
| Line 9: | Line 15: |
| * In Brainstorm software, we consider that the electric or magnetic activity which is recorded by the sensors is produced mainly by a set of electric dipoles located at the surface of the cortex. * Contrary to a dipole localization method, we fix the dipoles locations (cortex surface) and orientations (perpendicular to the cortex), and try to estimate the activity of each dipole at each time sample. * The grid of sources (dipoles) that is used is defined by the cortex surface we have imported in one of the previous tutorials; each vertex of this surface is considered as dipole. * The default surface distributed with Brainstorm have around 15.000 vertices. So we will have 15.000 dipole amplitudes to estimate. Using less vertices would just lower the resolution of the results; using more produces too much data and might lead to memory issues. * What we expect to get at the end of this process is a matrix which size is [Number of sensors x Number of sources] * For computing this matrix, two methods are available for MEG recordings in Brainstorm, and will be computed in this tutorial: * Single sphere model: the head is considered a homogeneous sphere * Overlapping spheres method: * BEM was implemented but still need to be validated. Coming soon. |
* In the Brainstorm software, we consider by default that the electric or magnetic activity which is recorded by the sensors is produced mainly by a set of electric dipoles located at the surface of the cortex.<<BR>> * The grid of sources (dipoles) that is used is defined by the cortex surface we have imported in one of the previous tutorials; each vertex of this surface is considered as a dipole. * The default surface distributed with Brainstorm have around 15,000 vertices. So we will have 15,000 dipole amplitudes to estimate. Using less vertices would just lower the resolution of the results; using more produces too much data and might lead to memory issues. * What we expect to get at the end of this process is a matrix whose size is [Number of sensors x Number of sources] * For computing this matrix, three methods are available for MEG recordings in Brainstorm: * '''Single sphere''': the head is considered a homogeneous sphere * '''Overlapping spheres''': Refining the previous model by fitting one local sphere for each sensor * '''OpenMEEG BEM''': Symmetric Boundary Element Method from the open-source software OpenMEEG. Described in an advanced tutorial: [[Tutorials/TutBem|BEM head model]]. |
| Line 19: | Line 24: |
| To understand properly what is a forward model, why this approach was chosen, what is the best method, etc. please visit the [[Theory]] section. For now we will only focus on how you compute a head model with the Brainstorm interface. | {{attachment:forwardInverse.gif}} |
| Line 24: | Line 29: |
| 1. Right-click on the ''!StimRightThumb'' condition and select ''Compute head model''. The ''Head modeler'' window will appear.<<BR>><<BR>> {{attachment:popupHeadModel.gif}} --- {{attachment:headModeler.gif}} 1. The only thing you can choose here is the forward model you will use: ''Single sphere'' or ''Overlapping spheres''. * Choose ''Single sphere''. * You can also edit the Comment field of the file that will be created. * If you had MEG and EEG recorded in the same file, you have had the option to compute a forward matrix for either MEG, EEG or both. * The ''Head compartments'' panel just shows the surfaces that are going to be used to compute this model. If you did your work while importing the subject's anatomy, you wouldn't even have to check this. |
1. Right-click on the ''Right'' condition and select ''Compute head model''. The ''Head modeler'' window will appear.<<BR>><<BR>> {{attachment:popupHeadModel.gif}} --- {{attachment:headModeler.gif}} 1. Set the options for your head model: * Source space: '''Cortex surface'''. <<BR>>The MRI volume option will be introduced in an advanced tutorial: [[Tutorials/TutVolSource|Volume source estimation]]. * Forward modeling method: '''Single sphere'''. * You can also edit the '''Comment '''field of the file that will be created (the string that will be representing the head model in the database explorer). |
| Line 31: | Line 35: |
| 1. Two other windows appear, to help you define the sphere. Estimating the best fitting sphere for a head is not always that easy as it looks like, for many reasons: a human head is usually not spherical, and the sets of point we can use at this point are not always well adapted for an automatic sphere estimation. <<BR>><<BR>> {{attachment:editBfsPanel.gif}} --- {{attachment:editBfsFigure.gif}} * Read and follow the instructions in those windows. * Click on ''Scalp ''surface, move and resize the sphere manually, just to see how it works. * Click again on Scalp: here we will use directly the estimation of the sphere based on the vertices of the ''Scalp ''surface (a simple least-squares fitting using all the vertices of the surface). * In that case, you may just use (click on ''Scalp ''button) * For EEG 3-shell spheres models, you just estimate and manipulate the outer sphere, and then use the ''Edit properties...'' button to defined the relative radii of the 2 other spheres, and their respective conductivities. This will be decribed in an other tutorial. * Click on ''Ok''. And wait for a few moments, the forward model computation is started. |
1. Two other windows appear, to help you define the sphere. Estimating the best fitting sphere for a head is not always as easy as it looks like, because a human head is usually not spherical. <<BR>><<BR>> {{attachment:helpBfs.gif|editBfs.gif}} <<BR>><<BR>> {{attachment:editBfsFigure.gif}} * Read and follow the instructions in the help window. * Click on the ''Scalp ''button, move and resize the sphere manually, just to see how it works. * Click again on ''Scalp'': here we will use directly the estimation of the sphere based on the vertices of the ''Scalp ''surface (a simple least-squares fitting using all the vertices of the surface). * For EEG 3-shell spheres models, you just estimate and manipulate the largest sphere (scalp), and then use the ''Edit properties...'' button in the toolbar to define the relative radii of the 2 other spheres, and their respective conductivities. This will be described in another tutorial. * Click on ''Ok'', and wait for a few seconds. |
| Line 40: | Line 44: |
| * You cannot do many interesting things only with this file, as it is only a matrix that converts the cortical sources into MEG/EEG recordings, and we don't have any sources information yet. * You may just check the sphere(s) that were used to compute the head model, with the ''Check spheres'' menu. * This process created 3 files on the hard drive; the head model is the only information in Brainstorm database that is stored in a separate binary file instead of a .mat file: * '''headmodel_Single_sphere_CD.mat''': description, parameters, pointers to the binary files * '''headmodel_Single_sphere_CD.bin''': binary file that just contains the forward matrix (constrained orientation of the sources, perperpendicular with the cortex) * '''headmodel_Single_sphere_CD_xyz.bin''': another matrix, but with unconstrained orientations (3 values instead of one for each vertex, for each direction) |
* There is not much you can do with this file, as it is only a matrix that converts the cortical sources into MEG/EEG recordings, and we do not have any sources information yet. * You may just check the sphere(s) that were used to compute the head model. |
| Line 48: | Line 48: |
| Let's compute now an other forward model. The overlapping spheres method is based on the estimation of a different sphere for each sensor. Instead of using only one sphere for the whole head, it estimates a sphere that fits locally the shape of the head in the surroundings of each sensor. | Let's compute a more advanced forward model. The overlapping spheres method is based on the estimation of a different sphere for each sensor. Instead of using only one sphere for the whole head, it estimates a sphere that fits locally the shape of the head in the surroundings of each sensor. |
| Line 50: | Line 50: |
| 1. Right-click on ''!StimRightThumb'' condition and select ''Compute head model'' again. 1. Select the ''Overlapping spheres'' method and click on ''Run''. It doesn't ask for anything, the 151 spheres are estimated automatically. 1. At the end, the ''Check spheres'' window shows the spheres that were estimated. You check all them by following the indications written in green at the bottom of the window: use left/right arrows. At each step, the current sensor marker is displayed in red, and the sphere you see is its local estimation of the head shape. |
1. Right-click on ''Right'' condition and select ''Compute head model'' again. 1. Select the ''Overlapping spheres'' method and click on ''Run''. 1. This algorithm is supposed to use the inner skull surface from the subject, but we usually do not have this information. In this case, a pseudo-innerskull is reconstructed using a dilated version of the cortex envelope. 1. Right-click on the new head model > ''Check spheres''. This window shows the spheres that were estimated. You can check them by following the indications written in green at the bottom of the window: use left/right arrows. At each step, the current sensor marker is displayed in red, and the sphere you see is its local estimation of the head shape. |
| Line 54: | Line 55: |
| 1. Compute a head model for the Left condition (Overlapping spheres). | |
| Line 58: | Line 58: |
| We now have two head models in for our ''Subject01 / !StimRightThumb'' condition. | We now have two head models in for our ''Subject01 / Right'' condition. |
| Line 60: | Line 60: |
| * You can have several head models computed for the same dataset, but it is not really recommanded as it may not be really easy to know after that which one was used to compute the sources. * You want to keep them anyway, you have to indicate what is default one. You do that by double-clicking on of them, and it is supposed to turn green. The head model displayed in green is the one that will be used for the following computation step. |
* You can have several head models computed for the same dataset, but it is not recommended as it might be difficult afterwards to know which one was used to compute the sources. * If you want to keep them anyway, you have to indicate which one is the default one. You do that by double-clicking on one of them (or right-click > set as default head model), and it is supposed to turn green. The head model displayed in green is the one that will be used for the following computation steps. |
| Line 63: | Line 63: |
| * The only problem is that this method depends a lot on the scalp surface you use. If the surface has some irregularities, some holes or some extra blobs inside or outside the head, the fitting algorithm may crash. In that case you would have to fix your surface or use the single sphere method, as it is more robust. * For EEG, always prefer the "''3-shell sphere (BERG)''" model. |
* For EEG, always prefer the "OpenMEEG BEM" model. |
| Line 68: | Line 67: |
| You can run in two clicks the computation of overlapping spheres model for all the conditions or subjects you want in the database. | You can run in two clicks the computation of the overlapping spheres model for all the conditions or subjects you want in the database. |
| Line 70: | Line 69: |
| * The ''Compute head model'' menu is available in the many popup menus in the tree (protocol, subject, condition). It is then applied recursively to all the subjects and conditions that the node you selected include. | * The ''Compute head model'' menu is available in popup menus in the tree at all the levels (protocol, subject, condition). It is then applied recursively to all the subjects and conditions contained in the node(s) you selected. |
| Line 75: | Line 74: |
| == Additional discussions on the forum == * Sensor modeling: http://neuroimage.usc.edu/forums/showthread.php?1295 * Gain matrix: http://neuroimage.usc.edu/forums/showthread.php?918 * EEG reference: http://neuroimage.usc.edu/forums/showthread.php?1525#post6718 * EEG and default anatomy: http://neuroimage.usc.edu/forums/showthread.php?1774 |
|
| Line 76: | Line 81: |
| You are done with the forward problem. | Next step: the [[Tutorials/TutNoiseCov|estimation of the noise covariance]]. |
| Line 78: | Line 83: |
| Next step: the [[Tutorials/TutSourcesEstimation|inverse problem]]. | <<EmbedContent(http://neuroimage.usc.edu/brainstorm3_register/get_feedback.php?Tutorials/TutHeadModel)>> |
[WARNING: This page is part of the old tutorials, please refer to the new documentation]
Tutorial 6: Computing a head model
Authors: Francois Tadel, John C Mosher, Richard Leahy, Sylvain Baillet
This tutorial is still based on Sabine Meunier's somatotopy experiment, called TutorialCTF in your Brainstorm database. The recordings have already been imported and analyzed at the sensors level, they are ready for source estimation.
Contents
Forward problem
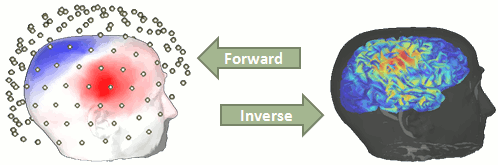
The first step consists in computing a model that explains how an electric current flowing in the brain can influence what is recorded out of the head, by the EEG or MEG sensors.
- This problem is called forward problem.
Its result is called head model in Brainstorm interface, but can also be referred as forward model or leadfield matrix.
In the Brainstorm software, we consider by default that the electric or magnetic activity which is recorded by the sensors is produced mainly by a set of electric dipoles located at the surface of the cortex.
- The grid of sources (dipoles) that is used is defined by the cortex surface we have imported in one of the previous tutorials; each vertex of this surface is considered as a dipole.
- The default surface distributed with Brainstorm have around 15,000 vertices. So we will have 15,000 dipole amplitudes to estimate. Using less vertices would just lower the resolution of the results; using more produces too much data and might lead to memory issues.
- What we expect to get at the end of this process is a matrix whose size is [Number of sensors x Number of sources]
- For computing this matrix, three methods are available for MEG recordings in Brainstorm:
Single sphere: the head is considered a homogeneous sphere
Overlapping spheres: Refining the previous model by fitting one local sphere for each sensor
OpenMEEG BEM: Symmetric Boundary Element Method from the open-source software OpenMEEG. Described in an advanced tutorial: BEM head model.
Single sphere model
Select the TutorialCTF protocol, close all the figures, and follow these steps:
Right-click on the Right condition and select Compute head model. The Head modeler window will appear.
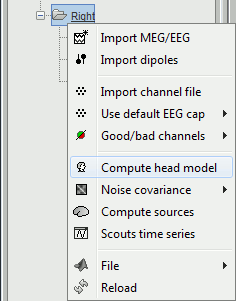 ---
--- 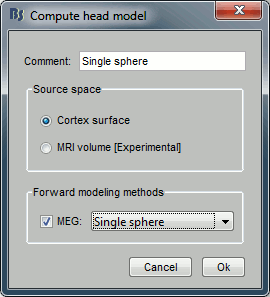
- Set the options for your head model:
Source space: Cortex surface.
The MRI volume option will be introduced in an advanced tutorial: Volume source estimation.Forward modeling method: Single sphere.
You can also edit the Comment field of the file that will be created (the string that will be representing the head model in the database explorer).
Click on Run.
Two other windows appear, to help you define the sphere. Estimating the best fitting sphere for a head is not always as easy as it looks like, because a human head is usually not spherical.

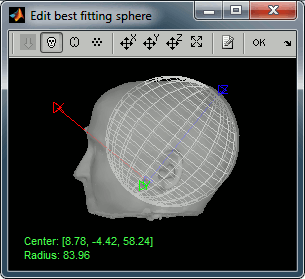
- Read and follow the instructions in the help window.
Click on the Scalp button, move and resize the sphere manually, just to see how it works.
Click again on Scalp: here we will use directly the estimation of the sphere based on the vertices of the Scalp surface (a simple least-squares fitting using all the vertices of the surface).
For EEG 3-shell spheres models, you just estimate and manipulate the largest sphere (scalp), and then use the Edit properties... button in the toolbar to define the relative radii of the 2 other spheres, and their respective conductivities. This will be described in another tutorial.
Click on Ok, and wait for a few seconds.
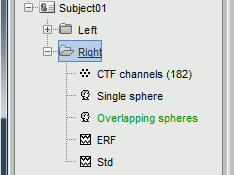
A new file appeared just below the channel file, it represents the head model.
- There is not much you can do with this file, as it is only a matrix that converts the cortical sources into MEG/EEG recordings, and we do not have any sources information yet.
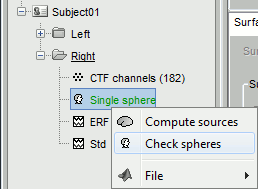
- You may just check the sphere(s) that were used to compute the head model.
Overlapping spheres model
Let's compute a more advanced forward model. The overlapping spheres method is based on the estimation of a different sphere for each sensor. Instead of using only one sphere for the whole head, it estimates a sphere that fits locally the shape of the head in the surroundings of each sensor.
Right-click on Right condition and select Compute head model again.
Select the Overlapping spheres method and click on Run.
- This algorithm is supposed to use the inner skull surface from the subject, but we usually do not have this information. In this case, a pseudo-innerskull is reconstructed using a dilated version of the cortex envelope.
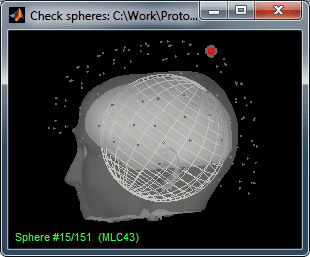
Right-click on the new head model > Check spheres. This window shows the spheres that were estimated. You can check them by following the indications written in green at the bottom of the window: use left/right arrows. At each step, the current sensor marker is displayed in red, and the sphere you see is its local estimation of the head shape.
Close this window when you reviewed them all.
- Compute a head model for the Left condition (Overlapping spheres).
Selection of a head model
We now have two head models in for our Subject01 / Right condition.
- You can have several head models computed for the same dataset, but it is not recommended as it might be difficult afterwards to know which one was used to compute the sources.
If you want to keep them anyway, you have to indicate which one is the default one. You do that by double-clicking on one of them (or right-click > set as default head model), and it is supposed to turn green. The head model displayed in green is the one that will be used for the following computation steps.
- For MEG, when it works properly, the overlapping spheres model usually gives better results than the single sphere one. In this particular case, it produces more focal results, so we are going to use it for the next steps.
- For EEG, always prefer the "OpenMEEG BEM" model.
Now to make things clearer: delete the Single sphere head model, and keep the Overlapping spheres.
Batching head model computation
You can run in two clicks the computation of the overlapping spheres model for all the conditions or subjects you want in the database.
The Compute head model menu is available in popup menus in the tree at all the levels (protocol, subject, condition). It is then applied recursively to all the subjects and conditions contained in the node(s) you selected.
Example: If you want to compute it on all the subjects and all the conditions, select the Compute head model menu from the protocol node TutorialCtf. For all the conditions of Subject01, run it from the Subject01 popup menu. Etc.
If you only want to compute it on some subjects of the protocol, select them at once holding the Ctrl key, right-click on one, and select the Compute head model menu.
To process all the subjects for one condition, switch to the Functional data (sorted by conditions) view of the database.
Additional discussions on the forum
Sensor modeling: http://neuroimage.usc.edu/forums/showthread.php?1295
Gain matrix: http://neuroimage.usc.edu/forums/showthread.php?918
EEG reference: http://neuroimage.usc.edu/forums/showthread.php?1525#post6718
EEG and default anatomy: http://neuroimage.usc.edu/forums/showthread.php?1774
Next
Next step: the ?estimation of the noise covariance.
