|
Size: 11485
Comment:
|
Size: 11478
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 59: | Line 59: |
| * Create a new category of markers "'''blink_manual'''", using the menu Events > Add group. Select this new group. * Review the file, and mark the peaks you observe on the EOG trace, using the Ctrl+E keyboard shortcut. Do that for a few eye blinks.<<BR>><<BR>> {{attachment:markEog.gif|tsMisc.gif}} * You could repeat the same operation for all the blinks, then for all the ECG peaks and jump to the next chapter of the tutorial and compute the SSP. It would be uselessly time consuming, as there is a process that does it for you automatically. However, it is good to remember how to do it manually because you may face some cases where you don't have clean ECG/EOG, or if you want to correct for another type of artifact. |
Create a new category of markers "'''blink_manual'''", using the menu Events > Add group. Select this new group. Review the file, and mark the peaks you observe on the EOG trace, using the Ctrl+E keyboard shortcut. Do that for a few eye blinks. {{attachment:markEog.gif|tsMisc.gif}} You could repeat the same operation for all the blinks, then for all the ECG peaks and jump to the next chapter of the tutorial and compute the SSP. It would be uselessly time consuming, as there is a process that does it for you automatically. However, it is good to remember how to do it manually because you may face some cases where you don't have clean ECG/EOG, or if you want to correct for another type of artifact. |
| Line 64: | Line 66: |
| * In the event tab, select the menu: '''SSP > Detect heartbeats'''. It opens automatically the pipeline editor, with the process "Detect heartbeats" selected<<BR>><<BR>> {{attachment:detectEcg.gif|tsMisc.gif}} * '''Channel name''': Name of the channel that is used to perform the detection. Select or type "'''EEG057'''" as it is the name of the ECG channel * '''Time window''': Time range that the algorithm should scan for the selected artifact. Leave the default values to process the entire file. Ignore the small rounding errors that lead to displaying a very small negative value instead of "0". * '''Event name''': Name of the event group that is created for saving all the detected events. Leave the default "cardiac". * After the process stops, you can see a new event category "'''cardiac'''" in the events tab, with 346 occurrences. You can check a few of them, to make sure that the "cardiac" markers really indicate the ECG peaks, and that there are not too many peaks that are skipped. In the event tab, click on the "cardiac" event category, then on a time occurrence to jump to it in the MEG and Misc time series figures.<<BR>><<BR>> {{attachment:detectEcgDone.gif|tsMisc.gif}} |
In the event tab, select the menu: '''SSP > Detect heartbeats'''. It opens automatically the pipeline editor, with the process "Detect heartbeats" selected: * '''Channel name''': Name of the channel that is used to perform the detection. Select or type "'''EEG057'''" as it is the name of the ECG channel * '''Time window''': Time range that the algorithm should scan for the selected artifact. Leave the default values to process the entire file. Ignore the small rounding errors that lead to displaying a very small negative value instead of "0". * '''Event name''': Name of the event group that is created for saving all the detected events. Leave the default "cardiac". {{attachment:detectEcg.gif|tsMisc.gif}} Click on Run. After the process stops, you can see a new event category "'''cardiac'''" in the events tab, with 346 occurrences. You can check a few of them, to make sure that the "cardiac" markers really indicate the ECG peaks, and that there are not too many peaks that are skipped. In the event tab, click on the "cardiac" event category, then on a time occurrence to jump to it in the MEG and Misc time series figures. {{attachment:detectEcgDone.gif|tsMisc.gif}} |
| Line 71: | Line 79: |
| * Now do the same thing for the eye blinks. In the event tab, select the menu "SSP > Detect eye blinks". Configure the process to use the channel '''EEG058''' (name of the EOG channel), and leave the other options to the default values<<BR>><<BR>> {{attachment:detectEog.gif|tsMisc.gif}} * The result of this process is slightly different from the heartbeat detection: it created two event groups instead of one, "'''blink'''" and "'''blink2'''". Indeed, this algorithm not only detects specific events in a signal, it also classifies them by shape. If you go through all the events that were detected in the two categories, you would see that the "blink" are all round bumps, typical of the '''eye blinks''', while the "blink2" are ramps followed by sharp drops, a shape that indicates an '''eye saccade'''.<<BR>><<BR>> {{attachment:detectEogDone.gif|tsMisc.gif}} |
Now do the same thing for the eye blinks. In the event tab, select the menu "SSP > Detect eye blinks". Configure the process to use the channel '''EEG058''' (name of the EOG channel), and leave the other options to the default values. |
| Line 74: | Line 81: |
| {{attachment:detectEog.gif|tsMisc.gif}} | |
| Line 75: | Line 83: |
| The result of this process is slightly different from the heartbeat detection: it created two event groups instead of one, "'''blink'''" and "'''blink2'''". Indeed, this algorithm not only detects specific events in a signal, it also classifies them by shape. If you go through all the events that were detected in the two categories, you would see that the "blink" are all round bumps, typical of the '''eye blinks''', while the "blink2" are ramps followed by sharp drops, a shape that indicates an '''eye saccade'''. {{attachment:detectEogDone.gif|tsMisc.gif}} === Automatic detection: Custom === |
Artifact cleaning with SSP
It is common to have portions of recordings contaminated by events coming from the subject (eye blinks, movements, heartbeats, teeth clenching, implanted stimulators...) or from the environment (stimulation equipment, elevators, cars, trains, building vibrations...). Some of them are well defined, reproducible and can be removed efficiently using Signal Space Projections (SSP). The purpose of this tutorial is to introduce this technique to correct for the cardiac and ocular artifacts.
For this tutorial, we are going to use the protocol created in the previous tutorial ?Review continuous recordings and edit markers. If you have not followed this tutorial yet, please do it now.
Signal Space Projection
The Signal-Space Projection (SSP) is one approach to rejection of external disturbances. Here is a short description of the method by Matti Hämäläinen, from the MNE 2.7 reference manual, section 4.16.
Unlike many other noise-cancellation approaches, SSP does not require additional reference sensors to record the disturbance fields. Instead, SSP relies on the fact that the magnetic field distributions generated by the sources in the brain have spatial distributions sufficiently different from those generated by external noise sources. Furthermore, it is implicitly assumed that the linear space spanned by the significant external noise patterns has a low dimension.
Without loss of generality we can always decompose any n-channel measurement b(t) into its signal and noise components as:
b(t) = bs(t) + bn(t)
Further, if we know that bn(t) is well characterized by a few field patterns b1...bm, we can express the disturbance as
bn(t) = Ucn(t) + e(t) ,
where the columns of U constitute an orthonormal basis for b1...bm, cn(t) is an m-component column vector, and the error term e(t) is small and does not exhibit any consistent spatial distributions over time, i.e., Ce = E{eeT} = I. Subsequently, we will call the column space of U the noise subspace. The basic idea of SSP is that we can actually find a small basis set b1...bm such that the conditions described above are satisfied. We can now construct the orthogonal complement operator
P⊥ = I - UUT
and apply it to b(t) yielding
b(t) ≈ P⊥bs(t) ,
since P⊥bn(t) = P⊥Ucn(t) ≈ 0. The projection operator P⊥ is called the signal-space projection operator and generally provides considerable rejection of noise, suppressing external disturbances by a factor of 10 or more. The effectiveness of SSP depends on two factors:
The basis set b1...bm should be able to characterize the disturbance field patterns completely and
The angles between the noise subspace space spanned by b1...bm and the signal vectors bs(t) should be as close to π/2 as possible.
If the first requirement is not satisfied, some noise will leak through because P⊥bn(t) ≠ 0. If the any of the brain signal vectors bs(t) is close to the noise subspace not only the noise but also the signal will be attenuated by the application of P⊥ and, consequently, there might by little gain in signal-to-noise ratio.
Since the signal-space projection modifies the signal vectors originating in the brain, it is necessary to apply the projection to the forward solution in the course of inverse computations.
For more information on the SSP method, please consult the following publications:
- C. D. Tesche, M. A. Uusitalo, R. J. Ilmoniemi, M. Huotilainen, M. Kajola, and O. Salonen, "Signal-space projections of MEG data characterize both distributed and well-localized neuronal sources," Electroencephalogr Clin Neurophysiol, vol. 95, pp. 189-200, 1995.
- M. A. Uusitalo and R. J. Ilmoniemi, "Signal-space projection method for separating MEG or EEG into components," Med Biol Eng Comput, vol. 35, pp. 135-40, 1997.
Identifying the artifact
The first step is to identify several repetitions of the artifact (the vectors b1...bm). We need to set markers in the recordings that indicate when the events that we want to correct for occur. To help with this task, it is recommended to always record with bipolar electrodes the activity of the eyes (electro-oculogram or EOG), the heart (electro-cardiogram or ECG), and possibly other sources of muscular contaminations (electromyogramor EMG). In this example, we are going to use the ECG and EOG traces to mark the cardiac activity and the eye blinks. Two methods can be used, manual or automatic.
EOG/ECG channels
Select the protocol TutorialRaw created in the previous tutorial, and select the "Functional data" view (second button in the toolbar on top of the database explorer).
- Double-click on the continuous recordings ("Link to raw file") to open the MEG recordings.
Set the length of the reviewed time window to 3 seconds (in the Event tab, text box "Duration")
Right-click on the continuous recordings again > Misc > Display time series.
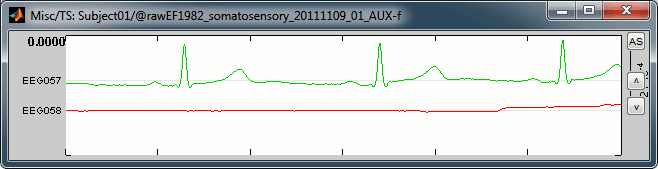
In this file the channel type "Misc" groups the two channels EEG057 (ECG, in green) and EEG058 (EOG, in red). This configuration depends on the acqusition setup, and can be redefined afterwards in Brainstorm (right-click on the channel file > Edit channel file, and then change manually the string in the column Type for any channel).
- Use the shortcuts introduced in the previous tutorial to adjust the vertical scale of this display: Shift+mouse wheel, +/- keys, or the buttons on the right side of the figure.
Then go further in time to see what is happening on those channels over the time: use the ">>>" buttons, or the associated shortcuts (read the tooltips of the button)
ECG: On the green trace, you can recognize the very typical shape of the electric activity of the heart (P, QRS and T waves). This is a very good example, the signal is not always that clean.
EOG: On the red trace, there is not much happening for most of the recordings except for a few bumps, typical of eye blinks, eg. at 33.590s. This subject is sitting very still and is not blinking much. We can expect MEG recordings of a very good quality.
You can observe the contamination from a blink on the left-frontal sensors: move to 33.590s (you can use the text boxes in the time panel, the event tab, or the scrollbar in the figure), and select a subset of sensors (Shift+B, or right-click on the figure > Display setup > CTF LF)
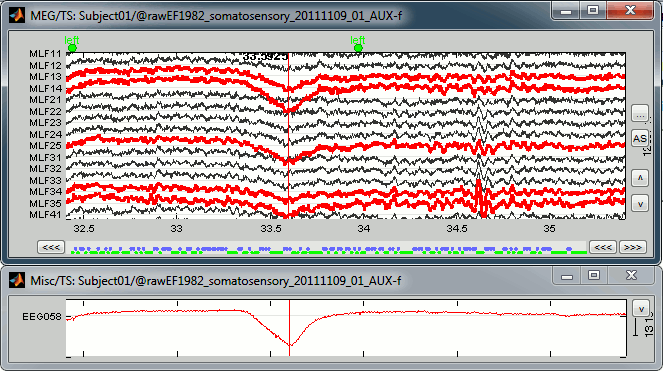
Manual marking
Create a new category of markers "blink_manual", using the menu Events > Add group. Select this new group. Review the file, and mark the peaks you observe on the EOG trace, using the Ctrl+E keyboard shortcut. Do that for a few eye blinks.
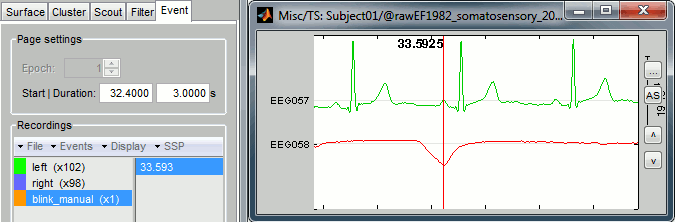
You could repeat the same operation for all the blinks, then for all the ECG peaks and jump to the next chapter of the tutorial and compute the SSP. It would be uselessly time consuming, as there is a process that does it for you automatically. However, it is good to remember how to do it manually because you may face some cases where you don't have clean ECG/EOG, or if you want to correct for another type of artifact.
Automatic detection: ECG
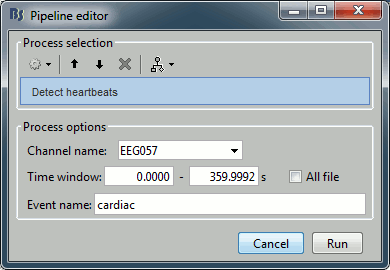
In the event tab, select the menu: SSP > Detect heartbeats. It opens automatically the pipeline editor, with the process "Detect heartbeats" selected:
Channel name: Name of the channel that is used to perform the detection. Select or type "EEG057" as it is the name of the ECG channel
Time window: Time range that the algorithm should scan for the selected artifact. Leave the default values to process the entire file. Ignore the small rounding errors that lead to displaying a very small negative value instead of "0".
Event name: Name of the event group that is created for saving all the detected events. Leave the default "cardiac".
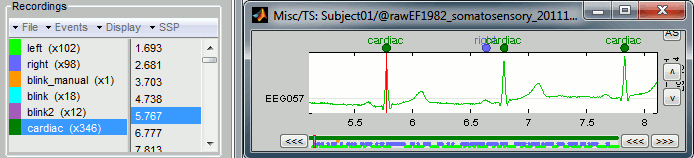
Click on Run. After the process stops, you can see a new event category "cardiac" in the events tab, with 346 occurrences. You can check a few of them, to make sure that the "cardiac" markers really indicate the ECG peaks, and that there are not too many peaks that are skipped. In the event tab, click on the "cardiac" event category, then on a time occurrence to jump to it in the MEG and Misc time series figures.
Automatic detection: EOG
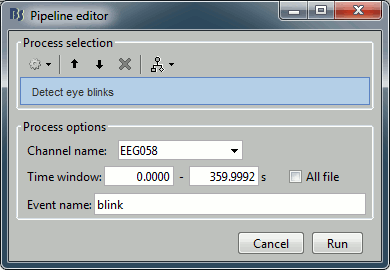
Now do the same thing for the eye blinks. In the event tab, select the menu "SSP > Detect eye blinks". Configure the process to use the channel EEG058 (name of the EOG channel), and leave the other options to the default values.
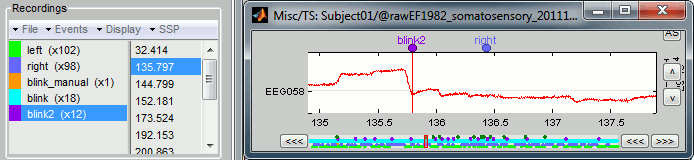
The result of this process is slightly different from the heartbeat detection: it created two event groups instead of one, "blink" and "blink2". Indeed, this algorithm not only detects specific events in a signal, it also classifies them by shape. If you go through all the events that were detected in the two categories, you would see that the "blink" are all round bumps, typical of the eye blinks, while the "blink2" are ramps followed by sharp drops, a shape that indicates an eye saccade.
Automatic detection: Custom
Calculating the projector
Then we concatenante those examples of the artifact, and we estimate an orthonormal basis U of this artifact subspace, from which when can finally calculate the projector that we will apply to the recordings.
Correction seperately EEG, MEG, and of Neuromag Vectorview systems, seperately MEG and EEG.
