|
Size: 4606
Comment:
|
Size: 3021
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 9: | Line 9: |
| The interface offers a few pre-defined montages for standard acquisition systems for MEG (Elekta-Neuromag, CTF, Yokogawa, 4D) and EEG (10-20 and 10-10 caps). The montage menu is accessible from the Record tab or with the popup menu on the time series figures. | The interface offers some pre-defined montages for standard acquisition systems: MEG (Elekta-Neuromag, CTF, Yokogawa, 4D), EEG (10-20 and 10-10 caps) and NIRS (overlay of multiple wavelengths). The montage menu is accessible from the Record tab or with the popup menu on the time series figures. |
| Line 13: | Line 13: |
| {{attachment:menu_popup.gif}} | {{attachment:menu_popup.gif||height="293",width="693"}} |
| Line 15: | Line 15: |
| == Montages == A better way to review a large number of MEG/EEG signals at once is to display only a subset of them at once. This can be done using the "montage" interface. The term refers mainly to EEG, to define the referencing system used to display the values recorded on the electrodes: bipolar montages, custom reference electrodes, average reference. However, the same interface can be used in MEG to select only a subset of sensors in a given figure, using a predefined or custom set of channels. |
== Edit montages == To edit/create your own montages, use the menu "Edit montages". By default, it shows only the montages that are relevant to the type of recordings you are currently looking at (in the example below: EEG 10-20). To see all the available options, click on the button "All". |
| Line 18: | Line 18: |
| * Use the drop-down menu in the Record tab to several predefined sets (ex. CTF LF = Group of left frontal sensors). Alternatively, right-click on the figure > Montage. You may need to decrease again the gain of the channels (use SHIFT+Mouse wheel) . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutExploreRecodings?action=AttachFile&do=get&target=selectionsMenu.gif|selectionsMenu.gif|class="attachment"}} |
{{attachment:all_montages.gif}} |
| Line 21: | Line 20: |
| * Note the keyboard shortcuts: '''SHIFT+letter'''. It makes it really easy to switch from a group of sensors to another (ex: Shift+A = all sensors, Shift+B = Left-frontal, Shift+C=Right-frontal...) | To create a new montage, click on the drop-down menu in the toolbar, the select the appropriate option (detailed in the following sections). |
| Line 23: | Line 22: |
| * To edit/create your own selections, use the menu "Edit montages". You can edit the channels selection groups or the electrodes montages available for this installation of Brainstorm. By default, it shows only the ones that are relevant to the type of recordings you are currently looking at (in this case: CTF MEG recordings). To see all the available options, click on the button "All". | {{attachment:new_montage.gif}} |
| Line 25: | Line 24: |
| * Click on "New channel selection" to create a new selection Test1, and select a few channels in the list. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutExploreRecodings?action=AttachFile&do=get&target=selections.gif|selections.gif|class="attachment"}} |
== Channel selection == To create a new selection of data channels, select the menu "'''New channel selection'''", then select the list of channels you want to display in the list of available channels, holding the Shift or Ctrl/Command key of your keyboard. The channels will be displayed in the same order as they are defined in the channel file. If you want to change the order of the channels, you need to create a custom montage. |
| Line 28: | Line 27: |
| * Click on Save. Back to the time series figure. Look in the popup menu again, you can see your new selection in the menu: Test1. You can now use the time series view with these predefined selections of sensors. * To display several montages at the same time, or with different different display options, you need to open twice the same file and then set the parameters differently. You have two option to clone the figure: * Do twice: Right-click on the file in the database explorer > MEG > Display time series * Right-click on a figure > Figure > Clone figure * To have the same scale in both figures, use the buttons "..." in the figures {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutExploreRecodings?action=AttachFile&do=get&target=figClone.gif|figClone.gif|class="attachment"}} |
{{attachment:selection.gif}} |
Montage editor [Under construction]
Authors: Francois Tadel
The display of the time series figures can be configured using montages of sensors. The term montage in the Brainstorm interface can refer to a simple sub-selection of data channels, or a linear recombination of these channels (eg. average reference or other EEG re-referecing montage). The selection of channels was already introduced in the tutorials Continuous recordings and EEG and epilepsy. This page illustrates how to use the montage editor to create custom displays.
Pre-defined montages
The interface offers some pre-defined montages for standard acquisition systems: MEG (Elekta-Neuromag, CTF, Yokogawa, 4D), EEG (10-20 and 10-10 caps) and NIRS (overlay of multiple wavelengths). The montage menu is accessible from the Record tab or with the popup menu on the time series figures.
The keyboard shortcuts (Shift+A, Shift+B, etc), allow to jump quickly to a different montage. To display a second montage on the same recordings, open again the same file (right-click > Display time) and change the montage of the second window, as illustrated in the tutorial EEG and epilepsy.
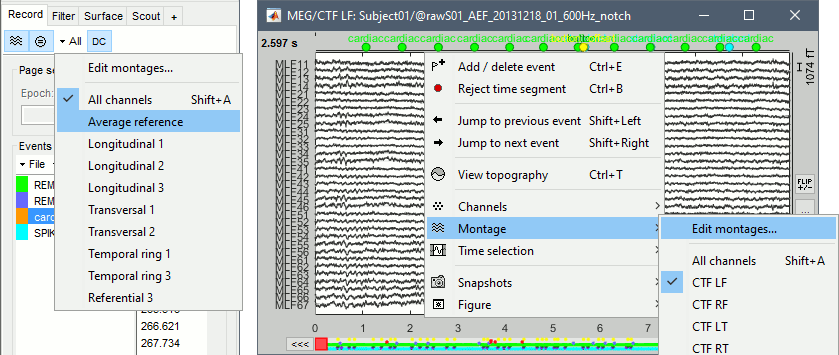
Edit montages
To edit/create your own montages, use the menu "Edit montages". By default, it shows only the montages that are relevant to the type of recordings you are currently looking at (in the example below: EEG 10-20). To see all the available options, click on the button "All".
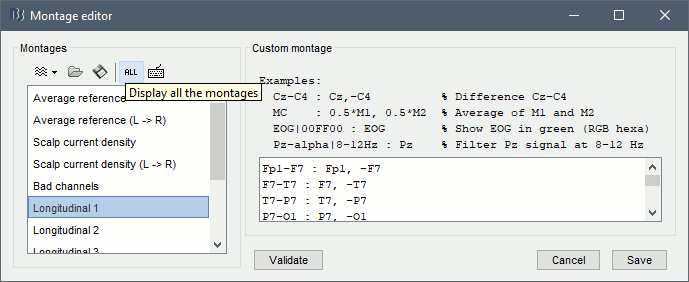
To create a new montage, click on the drop-down menu in the toolbar, the select the appropriate option (detailed in the following sections).
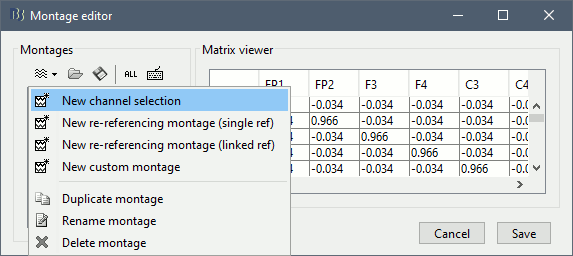
Channel selection
To create a new selection of data channels, select the menu "New channel selection", then select the list of channels you want to display in the list of available channels, holding the Shift or Ctrl/Command key of your keyboard. The channels will be displayed in the same order as they are defined in the channel file. If you want to change the order of the channels, you need to create a custom montage.
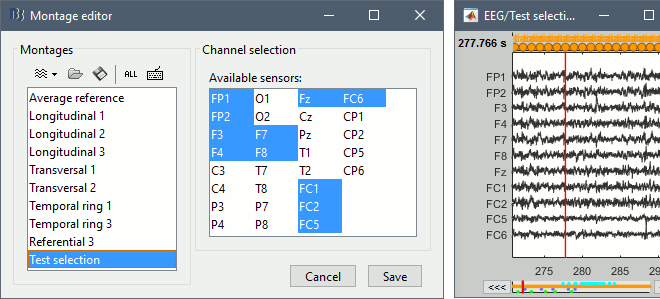
Apply a montage to the recordings
The montage selection is only a visualization option, it never modifies the recordings. If you want to apply a specific montage to the recordings, you need to call the process Standardize > Re-reference EEG (see EEG and Epilepsy) or the process Standardize > Apply montage (only available for imported epochs).
