Tutorial 28: Scripting
[TUTORIAL UNDER DEVELOPMENT: NOT READY FOR PUBLIC USE]
Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet
The previous tutorials explained how to use Brainstorm in an interactive way to process one subject with two acquisition runs. In the context of a typical neuroimaging study, you may have tens or hundreds of subjects to process in the same way, it is unrealistic to do everything manually. Some parts of the analysis can be processed in batches with no direct supervision, others require more attention. This tutorial introduces tools and tricks that will help you assemble an efficient analysis pipeline.
Requirements: You need a license for the Matlab environment in order to use these tools and execute custom scripts. If you are running the compiled version of Brainstorm with the MCR library, the only custom code you can run is through the menu File > Matlab console and the process "Run Matlab command".
Starting a new script
The easiest way to get started with a new Brainstorm script is to use the script generator, already introduced in the tutorial Select files and run processes . Select some files in the Process1 or Process2 tabs, select a list of processes, and use the menu Generate .m script . The example below should work on the the protocol "TutorialIntroduction" created during the introduction tutorials.
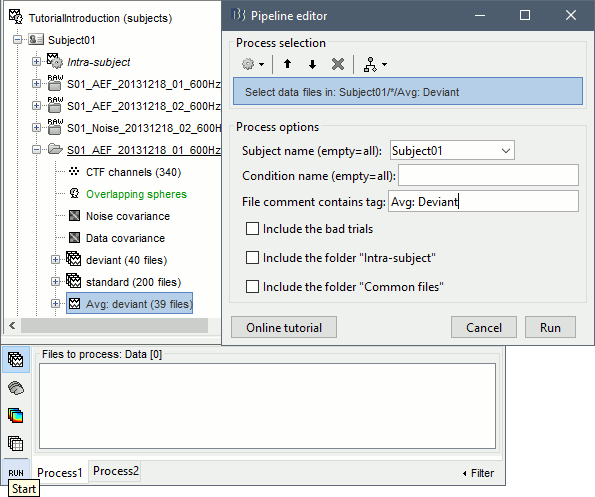
In the Process1 tab, leave the selection box empty and click on [Run]. Instead of selecting the files from the Brainstorm interface, we will select them directly from the database using a script. Select process File > Select files: Recordings (do not execute immediately)Subject01 , Condition=[Empty] , File comment=Avg: deviant (the space is important).
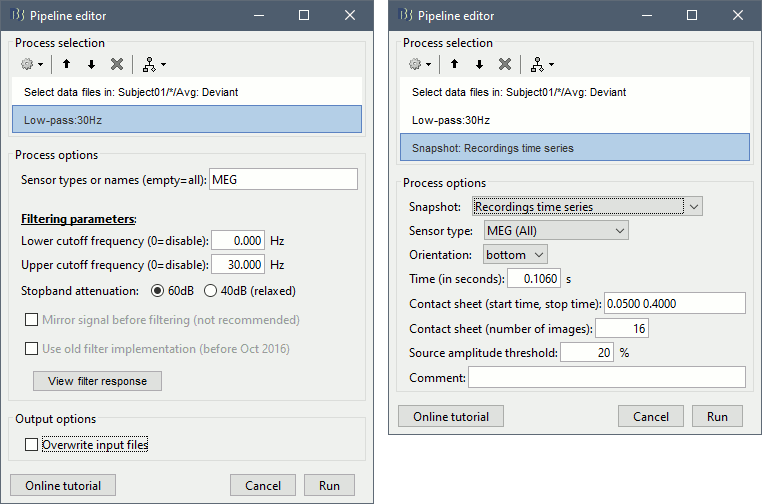
This process selects all the recordings with a comment including the string "Avg: deviant", from all the folders in Subject01 (except "Intra-subject" and "Common files"). We expect to get two files: the averages of the deviant condition for both runs. Add process Pre-process > Band-pass filter : Lower cutoff=0Hz , Upper cutoff=30Hz , Mirror. Add process File > Save snapshot : Recordings time series, Sensors=MEG.
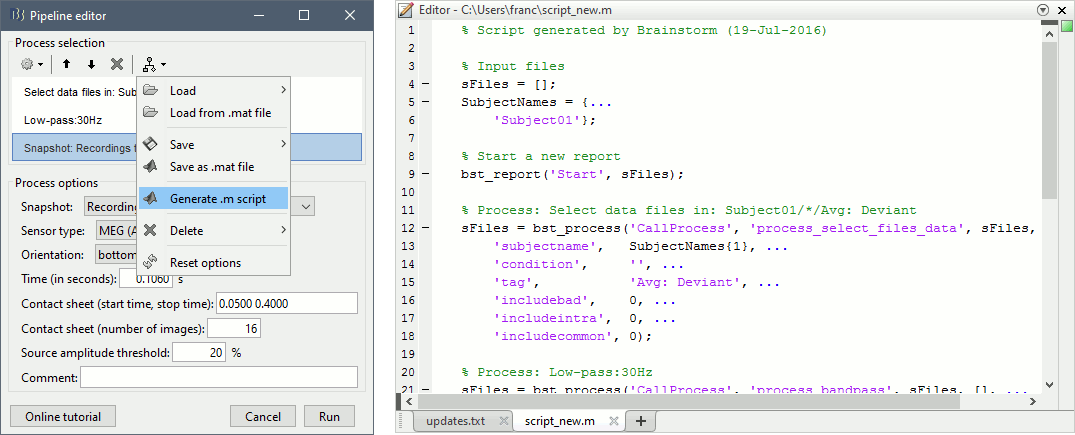
This will apply a low-pass filter at 30Hz and save a screen capture of the signals in the report. Do not run the pipeline, select the menu Generate .m script instead. It saves a new .m file and opens it in the Matlab editor. Close the pipeline editor window and look at the script.
The script you just generated can be the starting point to your own custom script. The following sections explain line by line how they work and how to edit them.
All the lines starting with a "%" are comments, they are never executed.
These lines define the script inputs:
sFiles : The list of files in input. Currently empty because we did not select anything in input in the Process1 list. If we had selected files, it would contain a cell array of strings with relative file paths.
SubjectNames : List of subject names that are used in the script. Most of the times, the generated script would contain only one entry, but it written as a cell array so that it is easier to extend it to multiple subjects with a loop (described further in this tutorial).
Start a new report of activity: Clears all the previous logs and gets ready to record new messages. The report will collect all the messages that are generated during the execution of the script by the various processes. You can explicitely add screen captures and additional messages to the current report with the function bst_report. This report will remain open until the function bst_report('Start') is called again. To display the current report, use the menu File > Report viewer .
The syntax function_name ('SubFunction ', arguments) is used a lot in Brainstorm, to call a subfunction available inside a .m file. This line above calls the function Report() in the file brainstorm3/toolbox/process/bst_report.m. This is made possible with the use of the short script "macro_methodcall" you will see at the top of many files. Many of the Brainstorm .m files are actually libraries of functions, rather than simple "scripts" or "functions".
Line by line: Body
You will find one block per process you selected in the pipeline editor. They all have the same syntax: bst_process ('CallProcess', process_name, input_files_A, input_files_B, options_list);
process_name : String indicating the function corresponding to the process to execute. To know from the pipeline editor what is the path to the process function: hover your mouse over the selected process, as illustrated in this tutorial .
input_files_A : List of input files in Process1, or FilesA in Process2. It can be a cell array of files names (full path, or relative path from the protocol folder), or an array of structures describing the files in the database (returned by a previous call to bst_process).
input_files_B : Empty for Process1, or FilesB in Process2. Cell array of strings or array of struct.
options_list : Pairs of (option_name, option_values), one for each option of the process.
output_files : Array of structures describing the files in output of the process. If the process created new files, this variable contains the new files. If the process didn't create new files or was modifying exiting files, most of the time this variable would contain the same files as the input list.
Closes the current report and saves it in the user Brainstorm report folder ($HOME/.brainstorm/reports). These reports are in .mat format and contain all the information necessary to re-run the execution exactly in the same way, but they not easy to read.
The parameter "sFiles" is optional, it indicates what are the files that are considered as the final results of the script. You can remove it without breaking your script: ReportFile = bst_report('Save');
Opens the report viewer to display what happened during the execution. This is equivalent to using the menu File > Report viewer . You can comment this line (ie. add a "%" at the beginning of the line) if you don't want to show the report at the end of the execution.
This function exports the report in readable format, as an HTML that includes all the screen captures embedded in it. It is disabled by default. If you want to use this feature: remove the "%" at the beginning of the line, and define the variable ExportDir.
ExportDir must be a string that defines where to save the HTML report. It can be wither the absolute path to a HTML file (eg. 'C:\Users\myuser\Documents\report_example.html') or just a folder (eg. 'C:\Users\myuser\Documents'). If you enter only a path to a folder, a default file name including the protocol name and a date tag is generated (report_ProtocolName_YYMMDD_HHMMSS.html).
Simplify the calls
This script you generated is like any Matlab script: you can edit it, rename the variables, add tests and loops, etc. The first important thing to understand is how to edit the options or change the inputs/outputs of a process. The script generator uses only one variable for all the file lists (sFiles ) and the output process is always the input of the following process. This is usually too restrictive to write a full analysis script: we commonly need to have multiple lists of files or to run two different operations on the same file.
Let's consider the first process call, which selects the averages for the Deviant condition in both runs.
There is no need to set the parameter sFiles, because there is no input, you can replace it with an empty matrix []. You can therefore remove the line "sFiles = [];". We can also rename the output variable "sAvgData", to be a bit more specific.
You can omit all the options that are not defined, not used, or kept to their default values:
Edit the call to the low-pass filter: Change the input to sAvgData and the output to sAvgDataLow, this way we will be able to keep track of the two files if we need to use them independently later in the script.
Edit the call to the snapshot process: Change the input to sAvgDataLow, and remove the output parameter (we are not expecting output from it).
Replace the last lines with the following, to export the report instead of opening in the report viewer (edit the file path to point at your own user folder instead).
Interaction with Matlab
Select the code for the first process in the Matlab editor, right-click > Evaluate selection (or press F9).
If you haven't executed your script yet, you will get the following error in the Matlab command window:
The variable SubjectNames is not defined yet: Execute the first two lines "SubjectNames = {'Subject01'}", then try again. You should now have a new variable in your Matlab workspace, which points at the two average files. Type "sAvgData(1)" in your command window to display the first element:
The field "sAvgData(1).FileName" contains the relative path to the to the Deviant average in the first run. This structure sAvgData contains also a lot of information that can be helpful in your script:
iStudy / iItem : Reference of the file in the database (described later in this tutorial).
FileType : 'raw' (continuous files), 'data' (recordings), 'results' (sources), 'timefreq' (time-frequency, spectrum and connectivity), or 'matrix' (any time series extracted from other files).
Comment : Comment field of the file (what is displayed in the database explorer)
Condition : Name of the condition/folder in which the file is located
SubjectFile : Relative path to the subject file (brainstormsubject.mat)
SubjectName : Name of the subject
DataFile : For types 'results' or 'timefreq', path of the parent file in the database explorer
ChannelFile : Relative path to the channel file
ChannelTypes : Cell array of channel types available for the input file
Naming conventions
To help you navigate in the Brainstorm code, here are some naming conventions:
Structures : Name starting with a "s" followed by a capital letter (eg. sFiles).
Indices : Either loop variables or array indices, name starting with a "i" (eg. iSubject, iStudy, iTime).
Counts : Number of elements in a group, name starting with a "n" (eg. nAvg, nTrials, nSubjects).
File names : Scripts and functions, only lower case, separation with "_" (eg. process_average)
Sub-functions : Inside a .m file, name starting with a capital, camel-case (eg. CallProcess, Start).
Running the script
The simplified script looks like this:
You have three ways to execute it:
Select all the lines (Ctrl+A) and evaluate it in Matlab (F9). In the Editor toolbar of the Matlab environment, click on the button [Run]. Save the file, go to this folder with Matlab (or add it to your path) and type the name of the script in the command window (without the ".m" at the end). At the end of the execution, nothing happens, because we indicated we wanted to just export the report instead of opening it. To check out the report of execution: use the menu File > Report viewer from the Brainstorm window, or open the report_test.html that was saved somewhere on your computer.
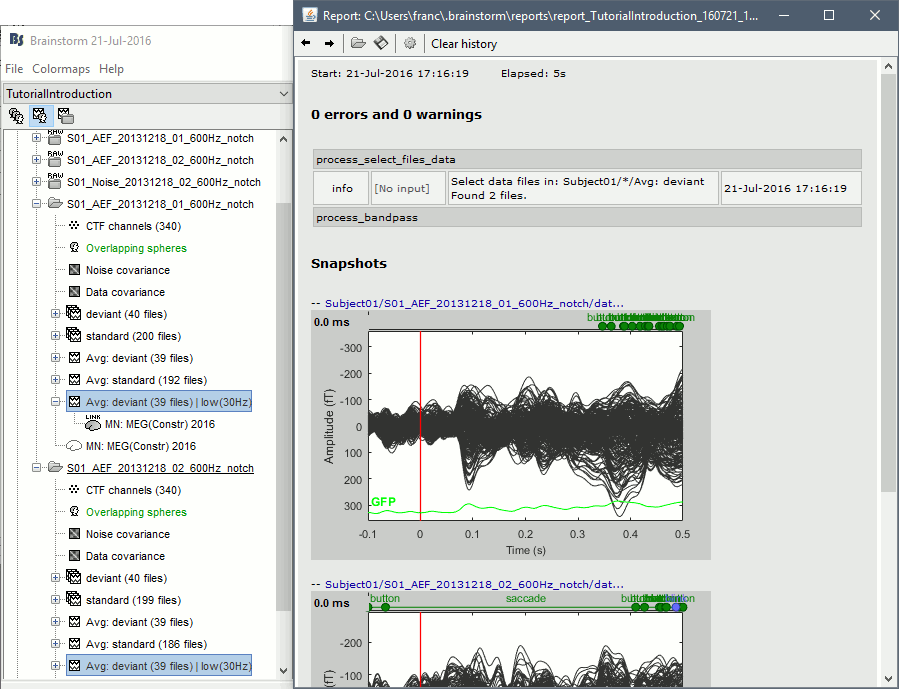
On this page, you can get review everything that happened in the script: when it was executed, how long it took, what are the processes that were executed, some produced messages (two files were selected with the first process), and at the end you have the screen captures taken by process_snapshot.
Running the script again
If you execute the script again, it will not behave as expected anymore. The selection process we used assumes that there is only one file per folder with a comment that includes "Avg: deviant". This is not the case anymore after the execution, because the low-pass filtered files also contain the same tags. The execution of the first process of the script now returns 4 files.
If you want to exclude the low-pass filtered files from this selection, you can add another process that will refine the selection. We will use the script generator again to
In Process1: Select any recordings file (at least one file)
Starting Brainstorm
- gui / nogui - selecting protocol - delete existing protocol
Selecting files
- Inputs / outputs
- Select processes
- Adding tags to help with the file selection later
Database requests
bst_get / bst_set
iStudy : Index of the study, get the structure with sStudy=bst_get('Study', sInput.iStudy)
iItem : Index of file in its category (example for recordings: sStudy.Data(sInput.iItem))
sStudy = bst_get('Study', sAvgData(1).iStudy)
sStudy.Data(sAvgData(1).iItem)
File manipulation
Modify a structure manually: Export to Matlab/Import from Matlab File manipulation: file_short, file_fullpath, in_bst_*... Documentation of all file structures: point at the appropriate tutorials Select files from the database (with bst_get and processes)
Loop over subject and runs
Creating loops is not supported yet by the script generator, but relatively easy to do from a script without having to know too much about Matlab programming.
1) Fill the cell array SubjectNames with all your subjects names, with the same dimensions as the list of input raw files (sFiles)
How to process many subjects
This section proposes a standard workflow for processing a full group study with Brainstorm. It contains the same steps of analysis as the introduction tutorials, but separating what can be done automatically from what should be done manually. This workflow can be adapted to most ERP studies (stimulus-based).
Prototype : Start by processing one or two subjects completely interactively (exactly like in the introduction tutorials). Use the few pilot subjects that you have for your study to prototype the analysis pipeline and check manually all the intermediate stages. Take notes of what you're doing along the way, so that you can later write a script that reproduces the same operations.
Anatomical fiducials : Set NAS/LPA/RPA and compute the MNI transformation for each subject.
Segmentation : Run FreeSurfer/BrainSuite to get surfaces and atlases for all the subjects.
File > Batch MRI fiducials : This menu prompts for the selection of the fiducials for all the subjects and saves a file fiducials.m in each segmentation folder. You will not have to redo this even if you have to start over your analysis from the beginning.
Script : Write a loop that calls the process "Import anatomy folder" for all the subjects.
Alternatives : Create and import the subjects one by one and set the fiducials at the import time. Or use the default anatomy for all the subjects (or use warped templates ).
Script #1 : Pre-processing: Loop on the subjects and the acquisition runs.
Create link to raw files : Link the subject and noise recordings to the database.
Event markers : Read and group triggers from digital and analog channel, fix stimulation delays
Evaluation : Power spectrum density of the recordings to evaluate their quality.
Pre-processing : Notch filter, sinusoid removal, band-pass filter.
Evaluation : Power spectrum density of the recordings to make sure the filters worked well.
Cleanup : Delete the links to the original files (the filtered ones are copied in the database).
Detect artifacts : Detect heartbeats, Detect eye blinks, Remove simultaneous.
Compute SSP : Heartbeats, Blinks (this selects the first component of each decomposition)
Compute ICA : If you have some artifacts you'd like to remove with ICA (no default selection).
Screenshots : Check the MRI/sensors registration, PSD before and after corrections, SSP.
Export the report : One report per subject, or one report for all the subjects, saved in HTML.
Manual inspection #1 :
Check the reports : Information messages (number of events, errors and warnings) and screen captures (registration problems, obvious noisy channels, incorrect SSP topographies).
Mark bad channels : Open the recordings, select the channels and mark them as bad. Or use the process "Set bad channels" to mark the same bad channels in multiple files.
Fix the SSP/ICA : For the suspicious runs: Open the file, adjust the list of blink and cardiac events, remove and recompute the SSP decompositions, manually select the components.
Detect other artifacts : Run the process on all the runs of all the subjects at once (select all the files in Process1 and run the process, or generate the equivalent script).
Mark bad segments : Review the artifacts detected in 1-7Hz and 40-240Hz, keep only the ones you really want to remove, then mark the event categories as bad. Review quickly the rest of the file and check that there are no other important artifacts.
Additional SSP : If you find one type of artifact that repeats (typically saccades and SQUID jumps), you can create additional SSP projectors, either with the process "SSP: Generic" or directly from a topography figure (right-click on the figure > Snapshot> Use as SSP projector).
Script #2 : Subject-level analysis: Epoching, averaging, sources, time-frequency.
Importing : Process "Import MEG/EEG: Events" and "Pre-process > Remove DC offset".
Averaging : Average trials by run, average runs by subject (registration problem in MEG).
Noise covariance : Compute from empty room or resting recordings, copy to other folders.
Head model : Compute for each run, or compute once and copy if the runs are co-registered.
Sources : Compute for each run, average across runs and subjects in source space for MEG.
Time-frequency : Computation with Hilbert transform or Morlet wavelets, then normalize.
Screenshots : Check the quality of all the averages (time series, topographies, sources).
Export the report : One report per subject, or one report for all the subjects, saved in HTML.
Manual inspection #2 :
Check the reports : Check the number of epochs imported and averaged in each condition, check the screen capture of the averages (all the primary responses should be clearly visible).
Regions of interest : If not using predefined regions from an atlas, define the scouts on the anatomy of each subject (or on the template and then project them to the subjects).
Script #3 : Group analysis, ROI-based analysis, etc.
Averaging : Group averages for the sensor data, the sources and the time-frequency maps.
Statistics : Contrast between conditions or groups of subjects.
Regions of interest : Any operation that involve scouts.
Final script
The following script from the Brainstorm distribution reproduces the introduction tutorials ("Get started"): brainstorm3/toolbox/script/tutorial_introduction.m
1 function tutorial_introduction(tutorial_dir, reports_dir)
2 % TUTORIAL_INTRODUCTION: Script that runs all the Brainstorm introduction tutorials.
3 %
4 % INPUTS:
5 % - tutorial_dir : Directory where the sample_introduction.zip file has been unzipped
6 % - reports_dir : Directory where to save the execution report (instead of displaying it)
7
8 % @=============================================================================
9 % This function is part of the Brainstorm software:
10 % https://neuroimage.usc.edu/brainstorm
11 %
12 % Copyright (c) University of Southern California & McGill University
13 % This software is distributed under the terms of the GNU General Public License
14 % as published by the Free Software Foundation. Further details on the GPLv3
15 % license can be found at http://www.gnu.org/copyleft/gpl.html.
16 %
17 % FOR RESEARCH PURPOSES ONLY. THE SOFTWARE IS PROVIDED "AS IS," AND THE
18 % UNIVERSITY OF SOUTHERN CALIFORNIA AND ITS COLLABORATORS DO NOT MAKE ANY
19 % WARRANTY, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO WARRANTIES OF
20 % MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE, NOR DO THEY ASSUME ANY
21 % LIABILITY OR RESPONSIBILITY FOR THE USE OF THIS SOFTWARE.
22 %
23 % For more information type "brainstorm license" at command prompt.
24 % =============================================================================@
25 %
26 % Author: Francois Tadel, 2016-2017
27
28
29 % ===== FILES TO IMPORT =====
30 % Output folder for reports
31 if (nargin < 2) || isempty(reports_dir) || ~isdir(reports_dir)
32 reports_dir = [];
33 end
34 % You have to specify the folder in which the tutorial dataset is unzipped
35 if (nargin == 0) || isempty(tutorial_dir) || ~file_exist(tutorial_dir)
36 error('The first argument must be the full path to the dataset folder.');
37 end
38 % Subject name
39 SubjectName = 'Subject01';
40 % Build the path of the files to import
41 AnatDir = fullfile(tutorial_dir, 'sample_introduction', 'anatomy');
42 Run1File = fullfile(tutorial_dir, 'sample_introduction', 'data', 'S01_AEF_20131218_01_600Hz.ds');
43 Run2File = fullfile(tutorial_dir, 'sample_introduction', 'data', 'S01_AEF_20131218_02_600Hz.ds');
44 NoiseFile = fullfile(tutorial_dir, 'sample_introduction', 'data', 'S01_Noise_20131218_02_600Hz.ds');
45 % Check if the folder contains the required files
46 if ~file_exist(Run1File)
47 error(['The folder ' tutorial_dir ' does not contain the folder from the file sample_introduction.zip.']);
48 end
49 % Re-inialize random number generator
50 if (bst_get('MatlabVersion') >= 712)
51 rng('default');
52 end
53
54
55 %% ===== TUTORIAL #1: CREATE PROTOCOL ================================================
56 % ===================================================================================
57 disp([10 'DEMO> Tutorial #1: Create protocol' 10]);
58 % The protocol name has to be a valid folder name (no spaces, no weird characters...)
59 ProtocolName = 'TutorialIntroduction';
60 % Start brainstorm without the GUI
61 if ~brainstorm('status')
62 brainstorm nogui
63 end
64 % Delete existing protocol
65 gui_brainstorm('DeleteProtocol', ProtocolName);
66 % Create new protocol
67 gui_brainstorm('CreateProtocol', ProtocolName, 0, 0);
68 % Start a new report
69 bst_report('Start');
70 % Reset colormaps
71 bst_colormaps('RestoreDefaults', 'meg');
72 % Set the current display mode to 'butterfly'
73 bst_set('TSDisplayMode', 'butterfly');
74
75
76 %% ===== TUTORIAL #2: IMPORT ANATOMY =================================================
77 % ===================================================================================
78 disp([10 'DEMO> Tutorial #2: Import anatomy' 10]);
79 % Process: Import FreeSurfer folder
80 bst_process('CallProcess', 'process_import_anatomy', [], [], ...
81 'subjectname', SubjectName, ...
82 'mrifile', {AnatDir, 'FreeSurfer'}, ...
83 'nvertices', 15000, ...
84 'nas', [127, 213, 139], ...
85 'lpa', [ 52, 113, 96], ...
86 'rpa', [202, 113, 91]);
87 % This automatically calls the SPM registration procedure because the AC/PC/IH points are not defined
88
89
90
91 %% ===== TUTORIAL #3: EXPLORE ANATOMY ================================================
92 % ===================================================================================
93 disp([10 'DEMO> Tutorial #3: Explore anatomy' 10]);
94 % Get subject definition
95 sSubject = bst_get('Subject', SubjectName);
96 % Get MRI file and surface files
97 MriFile = sSubject.Anatomy(sSubject.iAnatomy).FileName;
98 CortexFile = sSubject.Surface(sSubject.iCortex).FileName;
99 HeadFile = sSubject.Surface(sSubject.iScalp).FileName;
100 % Display MRI
101 hFigMri1 = view_mri(MriFile);
102 hFigMri3 = view_mri_3d(MriFile, [], [], 'NewFigure');
103 hFigMri2 = view_mri_slices(MriFile, 'x', 20);
104 pause(0.5);
105 % Close figures
106 close([hFigMri1 hFigMri2 hFigMri3]);
107 % Display scalp and cortex
108 hFigSurf = view_surface(HeadFile);
109 hFigSurf = view_surface(CortexFile, [], [], hFigSurf);
110 hFigMriSurf = view_mri(MriFile, CortexFile);
111 % Figure configuration
112 iTess = 2;
113 panel_surface('SetShowSulci', hFigSurf, iTess, 1);
114 panel_surface('SetSurfaceColor', hFigSurf, iTess, [1 0 0]);
115 panel_surface('SetSurfaceSmooth', hFigSurf, iTess, 0.5, 0);
116 panel_surface('SetSurfaceTransparency', hFigSurf, iTess, 0.8);
117 figure_3d('SetStandardView', hFigSurf, 'left');
118 pause(0.5);
119 % Close figures
120 close([hFigSurf hFigMriSurf]);
121
122
123
124 %% ===== TUTORIAL #4: CHANNEL FILE ===================================================
125 % ===================================================================================
126 disp([10 'DEMO> Tutorial #4: Channel file' 10]);
127 % Process: Create link to raw files
128 sFilesRun1 = bst_process('CallProcess', 'process_import_data_raw', [], [], ...
129 'subjectname', SubjectName, ...
130 'datafile', {Run1File, 'CTF'}, ...
131 'channelalign', 1);
132 sFilesRun2 = bst_process('CallProcess', 'process_import_data_raw', [], [], ...
133 'subjectname', SubjectName, ...
134 'datafile', {Run2File, 'CTF'}, ...
135 'channelalign', 1);
136 sFilesNoise = bst_process('CallProcess', 'process_import_data_raw', [], [], ...
137 'subjectname', SubjectName, ...
138 'datafile', {NoiseFile, 'CTF'}, ...
139 'channelalign', 0);
140 sFilesRaw = [sFilesRun1, sFilesRun2, sFilesNoise];
141 % Process: Snapshot: Sensors/MRI registration
142 bst_process('CallProcess', 'process_snapshot', [sFilesRun1, sFilesRun2], [], ...
143 'target', 1, ... % Sensors/MRI registration
144 'modality', 1, ... % MEG (All)
145 'orient', 1, ... % left
146 'Comment', 'MEG/MRI Registration');
147
148 % View sensors
149 hFig = view_surface(HeadFile);
150 hFig = view_channels(sFilesRun1.ChannelFile, 'MEG', 1, 1, hFig);
151 % Hide sensors
152 pause(0.5);
153 hFig = view_channels(sFilesRun1.ChannelFile, 'MEG', 0, 0, hFig);
154 % View coils
155 hFig = view_channels(sFilesRun1.ChannelFile, 'CTF', 1, 1, hFig);
156 % View helmet
157 pause(0.5);
158 hFig = view_helmet(sFilesRun1.ChannelFile, hFig);
159 pause(0.5);
160 close(hFig);
161 % Edit good/bad channel for current file
162 gui_edit_channel(sFilesRun1.ChannelFile);
163 pause(0.5);
164 % Unload everything
165 bst_memory('UnloadAll', 'Forced');
166
167
168
169 %% ===== TUTORIAL #5: REVIEW RAW =====================================================
170 % ===================================================================================
171 disp([10 'DEMO> Tutorial #5: Review raw' 10]);
172 % Process: Convert to continuous (CTF): Continuous
173 bst_process('CallProcess', 'process_ctf_convert', sFilesRaw, [], ...
174 'rectype', 2); % Continuous
175
176 % View recordings
177 hFigMeg = view_timeseries(sFilesRun1.FileName, 'MEG');
178 hFigEeg = view_timeseries(sFilesRun1.FileName, 'Misc');
179 hFigSel = view_timeseries(sFilesRun1.FileName, 'MEG', {'MLT11','MLT12','MLT13'});
180 % Figure configuration
181 pause(0.5);
182 panel_record('SetTimeLength', 3);
183 panel_record('SetStartTime', 100);
184 panel_record('SetDisplayMode', hFigMeg, 'column');
185 panel_montage('SetCurrentMontage', hFigMeg, 'CTF LT');
186 % Set filters: panel_filter('SetFilters', LowPassEnabled, LowPassValue, HighPassEnabled, HighPassValue, SinRemovalEnabled, SinRemovalValue, MirrorEnabled, FullSourcesEnabled)
187 panel_filter('SetFilters', 1, 100, 1, 1, 0, [], 0, 0);
188 pause(0.5);
189 panel_record('SetDisplayMode', hFigMeg, 'butterfly');
190 panel_montage('SetCurrentMontage', hFigMeg, '');
191 % Close figures
192 close([hFigMeg hFigEeg hFigSel]);
193
194
195
196 %% ===== TUTORIAL #8: STIM DELAYS ====================================================
197 % ===================================================================================
198 disp([10 'DEMO> Tutorial #8: Stim delays' 10]);
199 % Process: Detect: standard_fix
200 bst_process('CallProcess', 'process_evt_detect_analog', [sFilesRun1, sFilesRun2], [], ...
201 'eventname', 'standard_fix', ...
202 'channelname', 'UADC001', ...
203 'timewindow', [], ...
204 'threshold', 1.2, ...
205 'blanking', 0.2, ...
206 'highpass', 0, ...
207 'lowpass', 0, ...
208 'refevent', 'standard', ...
209 'isfalling', 0, ...
210 'ispullup', 0, ...
211 'isclassify', 0);
212 % Process: Detect: deviant_fix
213 bst_process('CallProcess', 'process_evt_detect_analog', [sFilesRun1, sFilesRun2], [], ...
214 'eventname', 'deviant_fix', ...
215 'channelname', 'UADC001', ...
216 'timewindow', [], ...
217 'threshold', 1.2, ...
218 'blanking', 0.2, ...
219 'highpass', 0, ...
220 'lowpass', 0, ...
221 'refevent', 'deviant', ...
222 'isfalling', 0, ...
223 'ispullup', 0, ...
224 'isclassify', 0);
225 % Process: Read from channel
226 bst_process('CallProcess', 'process_evt_read', [sFilesRun1, sFilesRun2], [], ...
227 'stimchan', 'UDIO001', ...
228 'trackmode', 1, ... % Value: detect the changes of channel value
229 'zero', 0);
230
231 % Process: Delete events
232 bst_process('CallProcess', 'process_evt_delete', [sFilesRun1, sFilesRun2], [], ...
233 'eventname', 'standard, deviant, button');
234 % Process: Rename event (standard_fix>standard)
235 bst_process('CallProcess', 'process_evt_rename', [sFilesRun1, sFilesRun2], [], ...
236 'src', 'standard_fix', ...
237 'dest', 'standard');
238 % Process: Rename event (deviant_fix>deviant)
239 bst_process('CallProcess', 'process_evt_rename', [sFilesRun1, sFilesRun2], [], ...
240 'src', 'deviant_fix', ...
241 'dest', 'deviant');
242 % Process: Rename event (64>button)
243 bst_process('CallProcess', 'process_evt_rename', [sFilesRun1, sFilesRun2], [], ...
244 'src', '64', ...
245 'dest', 'button');
246
247
248
249 %% ===== TUTORIAL #10: FREQUENCY FILTERS =============================================
250 % ===================================================================================
251 disp([10 'DEMO> Tutorial #10: Frequency filters' 10]);
252 % Process: Sinusoid removal: 60Hz 120Hz 180Hz 300Hz
253 sFilesNotch = bst_process('CallProcess', 'process_notch', sFilesRaw, [], ...
254 'freqlist', [60, 120, 180], ...
255 'sensortypes', 'MEG', ...
256 'read_all', 0);
257 % Process: Power spectrum density (Welch)
258 sFilesPsd = bst_process('CallProcess', 'process_psd', [sFilesRaw, sFilesNotch], [], ...
259 'timewindow', [], ...
260 'win_length', 4, ...
261 'win_overlap', 50, ...
262 'clusters', {}, ...
263 'sensortypes', 'MEG', ...
264 'edit', struct(...
265 'Comment', 'Power', ...
266 'TimeBands', [], ...
267 'Freqs', [], ...
268 'ClusterFuncTime', 'none', ...
269 'Measure', 'power', ...
270 'Output', 'all', ...
271 'SaveKernel', 0));
272 % Process: Snapshot: Frequency spectrum
273 bst_process('CallProcess', 'process_snapshot', sFilesPsd, [], ...
274 'target', 10, ... % Frequency spectrum
275 'modality', 1, ... % MEG (All)
276 'Comment', 'Power spectrum density');
277 % Process: Delete folders
278 bst_process('CallProcess', 'process_delete', sFilesRaw, [], ...
279 'target', 2); % Delete folders
280 % Separate the three outputs
281 sFilesRun1 = {sFilesNotch(1).FileName};
282 sFilesRun2 = {sFilesNotch(2).FileName};
283 sFilesNoise = {sFilesNotch(3).FileName};
284
285
286
287 %% ===== TUTORIAL #11: BAD CHANNELS ==================================================
288 % ===================================================================================
289 % % Process: Set bad channels
290 % sFiles = bst_process('CallProcess', 'process_channel_setbad', sFilesRun2, [], ...
291 % 'sensortypes', 'MRT51, MLO52, MLO42, MLO43');
292
293
294
295 %% ===== TUTORIAL #12: ARTIFACTS DETECTION ===========================================
296 % ===================================================================================
297 disp([10 'DEMO> Tutorial #12: Artifacts detection' 10]);
298 % Process: Detect heartbeats
299 bst_process('CallProcess', 'process_evt_detect_ecg', [sFilesRun1, sFilesRun2], [], ...
300 'channelname', 'ECG', ...
301 'timewindow', [], ...
302 'eventname', 'cardiac');
303 % Process: Detect eye blinks
304 bst_process('CallProcess', 'process_evt_detect_eog', [sFilesRun1, sFilesRun2], [], ...
305 'channelname', 'VEOG', ...
306 'timewindow', [], ...
307 'eventname', 'blink');
308 % Process: Remove simultaneous
309 bst_process('CallProcess', 'process_evt_remove_simult', [sFilesRun1, sFilesRun2], [], ...
310 'remove', 'cardiac', ...
311 'target', 'blink', ...
312 'dt', 0.25, ...
313 'rename', 0);
314
315
316
317 %% ===== TUTORIAL #13: SSP ===========================================================
318 % ===================================================================================
319 disp([10 'DEMO> Tutorial #13: SSP' 10]);
320 % Process: SSP ECG: cardiac
321 bst_process('CallProcess', 'process_ssp_ecg', [sFilesRun1, sFilesRun2], [], ...
322 'eventname', 'cardiac', ...
323 'sensortypes', 'MEG', ...
324 'usessp', 0, ...
325 'select', 1);
326 % Process: SSP EOG: blink
327 bst_process('CallProcess', 'process_ssp_eog', sFilesRun1, [], ...
328 'eventname', 'blink', ...
329 'sensortypes', 'MEG', ...
330 'usessp', 0, ...
331 'select', [1 2]);
332 bst_process('CallProcess', 'process_ssp_eog', sFilesRun2, [], ...
333 'eventname', 'blink', ...
334 'sensortypes', 'MEG', ...
335 'usessp', 0, ...
336 'select', 1);
337
338
339 %% ===== TUTORIAL #14: BAD SEGMENTS ==================================================
340 % ===================================================================================
341 disp([10 'DEMO> Tutorial #14: Bad segments' 10]);
342 % Process: Detect other artifacts
343 bst_process('CallProcess', 'process_evt_detect_badsegment', [sFilesRun1, sFilesRun2], [], ...
344 'timewindow', [], ...
345 'sensortypes', 'MEG', ...
346 'threshold', 3, ... % 3
347 'isLowFreq', 1, ...
348 'isHighFreq', 1);
349
350 % Process: Rename event (1-7Hz > saccade) (Run02 only)
351 bst_process('CallProcess', 'process_evt_rename', sFilesRun2, [], ...
352 'src', '1-7Hz', ...
353 'dest', 'saccade');
354
355 % Manual selection of saccades (cannot be done from the pipeline editor: manual edition of the structures)
356 sMatRun2 = in_bst_data(sFilesRun2{1}, 'F');
357 iEvtSaccade = find(strcmpi({sMatRun2.F.events.label}, 'saccade'));
358 sMatRun2.F.events(iEvtSaccade).times = [30, 81.5, 104, 142.5, 167, 187.5, 246.5, 319; 31, 83, 105, 144, 168, 188.5, 248, 320];
359 sMatRun2.F.events(iEvtSaccade).epochs = ones(1, size(sMatRun2.F.events(iEvtSaccade).times, 2));
360 sMatRun2.F.events(iEvtSaccade).channels = [];
361 sMatRun2.F.events(iEvtSaccade).notes = [];
362 bst_save(file_fullpath(sFilesRun2{1}), sMatRun2, 'v6', 1);
363
364 % Process: SSP: saccade (Run02 only)
365 bst_process('CallProcess', 'process_ssp', sFilesRun2, [], ...
366 'timewindow', [], ...
367 'eventname', 'saccade', ...
368 'eventtime', [-0.2, 0.2], ...
369 'bandpass', [1.5, 7], ...
370 'sensortypes', 'MEG', ...
371 'usessp', 1, ...
372 'saveerp', 0, ...
373 'method', 1, ...
374 'select', 1);
375 % Process: Detect other artifacts (Run02 only)
376 bst_process('CallProcess', 'process_evt_detect_badsegment', sFilesRun2, [], ...
377 'timewindow', [], ...
378 'sensortypes', 'MEG', ...
379 'threshold', 3, ... % 3
380 'isLowFreq', 1, ...
381 'isHighFreq', 1);
382
383 % Process: Rename event (1-7Hz > bad_1-7Hz)
384 bst_process('CallProcess', 'process_evt_rename', [sFilesRun1, sFilesRun2], [], ...
385 'src', '1-7Hz', ...
386 'dest', 'bad_1-7Hz');
387 % Process: Rename event (40-240Hz > bad_40-240Hz)
388 bst_process('CallProcess', 'process_evt_rename', [sFilesRun1, sFilesRun2], [], ...
389 'src', '40-240Hz', ...
390 'dest', 'bad_40-240Hz');
391 % Process: Snapshot: SSP projectors
392 bst_process('CallProcess', 'process_snapshot', [sFilesRun1, sFilesRun2], [], ...
393 'target', 2, ... % SSP projectors
394 'Comment', 'SSP projectors');
395
396
397
398 %% ===== TUTORIAL #15: IMPORT EVENTS =================================================
399 % ===================================================================================
400 disp([10 'DEMO> Tutorial #15: Import events' 10]);
401 % Process: Import MEG/EEG: Events (Run01)
402 sFilesEpochs1 = bst_process('CallProcess', 'process_import_data_event', sFilesRun1, [], ...
403 'subjectname', SubjectName, ...
404 'condition', '', ...
405 'eventname', 'standard, deviant', ...
406 'timewindow', [], ...
407 'epochtime', [-0.100, 0.500], ...
408 'createcond', 0, ...
409 'ignoreshort', 1, ...
410 'usectfcomp', 1, ...
411 'usessp', 1, ...
412 'freq', [], ...
413 'baseline', [-0.1, -0.0017]);
414 % Process: Import MEG/EEG: Events (Run02)
415 sFilesEpochs2 = bst_process('CallProcess', 'process_import_data_event', sFilesRun2, [], ...
416 'subjectname', SubjectName, ...
417 'condition', '', ...
418 'eventname', 'standard, deviant', ...
419 'timewindow', [], ...
420 'epochtime', [-0.100, 0.500], ...
421 'createcond', 0, ...
422 'ignoreshort', 1, ...
423 'usectfcomp', 1, ...
424 'usessp', 1, ...
425 'freq', [], ...
426 'baseline', [-0.1, -0.0017]);
427 % Display raster plot
428 hFigRaster = view_erpimage({sFilesEpochs1.FileName}, 'erpimage', 'MEG');
429 panel_display();
430 bst_report('Snapshot', hFigRaster, sFilesEpochs1(1).FileName, 'ERP image');
431 close(hFigRaster);
432
433
434 %% ===== TUTORIAL #16: AVERAGE =======================================================
435 % ===================================================================================
436 disp([10 'DEMO> Tutorial #16: Average' 10]);
437 % Process: Average: By trial group (folder average)
438 sFilesAvg = bst_process('CallProcess', 'process_average', [sFilesEpochs1, sFilesEpochs2], [], ...
439 'avgtype', 5, ... % By trial groups (folder average)
440 'avg_func', 1, ... % Arithmetic average: mean(x)
441 'weighted', 0, ...
442 'keepevents', 1);
443 % Process: Delete events 'cardiac'
444 bst_process('CallProcess', 'process_evt_delete', sFilesAvg, [], ...
445 'eventname', 'cardiac');
446 % Process: Snapshot: Recordings time series
447 bst_process('CallProcess', 'process_snapshot', sFilesAvg, [], ...
448 'target', 5, ... % Recordings time series
449 'modality', 1, ... % MEG (All)
450 'Comment', 'Evoked response');
451 % Set colormap: global color scale
452 bst_colormaps('SetMaxMode', 'meg', 'global');
453 % Process: Snapshot: Recordings topography (contact sheet)
454 bst_process('CallProcess', 'process_snapshot', sFilesAvg, [], ...
455 'target', 7, ... % Recordings topography (contact sheet)
456 'modality', 1, ... % MEG
457 'contact_time', [0, 0.350], ...
458 'contact_nimage', 15, ...
459 'Comment', 'Evoked response');
460
461 % Process: Average+Stderr: By trial group (subject average)
462 sFilesAvgAll = bst_process('CallProcess', 'process_average', [sFilesEpochs1, sFilesEpochs2], [], ...
463 'avgtype', 6, ... % By trial group (subject average)
464 'avg_func', 7, ... % Arithmetic average + Standard error
465 'weighted', 0, ...
466 'keepevents', 1);
467 % Process: Delete events 'cardiac'
468 bst_process('CallProcess', 'process_evt_delete', sFilesAvgAll, [], ...
469 'eventname', 'cardiac');
470 % Process: Delete events 'saccade'
471 bst_process('CallProcess', 'process_evt_delete', sFilesAvgAll, [], ...
472 'eventname', 'saccade');
473 % Process: Snapshot: Recordings time series
474 bst_process('CallProcess', 'process_snapshot', sFilesAvgAll, [], ...
475 'target', 5, ... % Recordings time series
476 'modality', 1, ... % MEG (All)
477 'Comment', 'Evoked response');
478
479
480 %% ===== TUTORIAL #17: EXPLORATION ===================================================
481 % ===================================================================================
482 disp([10 'DEMO> Tutorial #17: Bad segments' 10]);
483 % View averages
484 hFigMeg1 = view_timeseries(sFilesAvg(1).FileName, 'MEG');
485 hFigMeg2 = view_timeseries(sFilesAvg(2).FileName, 'MEG');
486 hFigEeg1 = view_timeseries(sFilesAvg(1).FileName, 'Misc');
487 hFigEeg2 = view_timeseries(sFilesAvg(2).FileName, 'Misc');
488 hFigTopo1 = view_topography(sFilesAvg(1).FileName, 'MEG', '2DSensorCap');
489 hFigTopo2 = view_topography(sFilesAvg(2).FileName, 'MEG', '2DSensorCap');
490 hFigTp2 = view_topography(sFilesAvg(3).FileName, 'MEG', '3DSensorCap');
491 hFigTp3 = view_topography(sFilesAvg(3).FileName, 'MEG', '2DDisc');
492 hFigTp4 = view_topography(sFilesAvg(3).FileName, 'MEG', '2DLayout');
493 % Set time: 90ms
494 panel_time('SetCurrentTime', 0.090);
495 % Set filters: 40Hz low-pass, no high-pass
496 panel_filter('SetFilters', 1, 40, 0, [], 0, [], 0, 0);
497 % View selected sensors
498 SelectedChannels = {'MLC31', 'MLC32'};
499 bst_figures('SetSelectedRows', SelectedChannels);
500 view_timeseries(sFilesAvg(4).FileName, [], SelectedChannels);
501 % Select time window
502 figure_timeseries('SetTimeSelectionManual', hFigMeg1, [0.070, 0.130]);
503 % Show sensors on 2DSensorCap topography
504 isMarkers = 1;
505 isLabels = 0;
506 figure_3d('ViewSensors', hFigTopo1, isMarkers, isLabels);
507 % Display time contact sheet for a figure
508 pause(0.5);
509 hContactFig = view_contactsheet( hFigTopo2, 'time', 'fig', [], 12, [0 0.120] );
510 pause(0.5);
511 close(hContactFig);
512
513
514
515 %% ===== TUTORIAL #18: COLORMAPS =====================================================
516 % ===================================================================================
517 disp([10 'DEMO> Tutorial #18: Colormaps' 10]);
518 % Set 'Meg' colormap to 'jet'
519 bst_colormaps('SetColormapName', 'meg', 'jet');
520 pause(0.5);
521 % Set 'Meg' colormap to 'rwb'
522 bst_colormaps('SetColormapName', 'meg', 'cmap_rbw');
523 % Set colormap to display absolute values
524 bst_colormaps('SetColormapAbsolute', 'meg', 1);
525 % Normalize colormap for each time frame
526 bst_colormaps('SetMaxMode', 'meg', 'local');
527 % Hide colorbar
528 bst_colormaps('SetDisplayColorbar', 'meg', 0);
529 pause(0.5);
530 % Restore colormap to default values
531 bst_colormaps('RestoreDefaults', 'meg');
532 % Edit good/bad channel for current file
533 gui_edit_channelflag(sFilesAvg(1).FileName);
534 % Close figures
535 pause(0.5);
536 bst_memory('UnloadAll', 'Forced');
537
538
539
540 %% ===== TUTORIAL #20: HEAD MODEL ====================================================
541 % ===================================================================================
542 disp([10 'DEMO> Tutorial #20: Head model' 10]);
543 % Process: Compute head model
544 bst_process('CallProcess', 'process_headmodel', sFilesAvg, [], ...
545 'comment', '', ...
546 'sourcespace', 1, ...
547 'meg', 3); % Overlapping spheres
548 % Get study structure
549 sStudy = bst_get('Study', sFilesAvg(1).iStudy);
550 % Show spheres
551 hFig = view_spheres(sStudy.HeadModel(sStudy.iHeadModel).FileName, sStudy.Channel.FileName, sSubject);
552 pause(0.5);
553 close(hFig);
554
555
556
557 %% ===== TUTORIAL #21: NOISE COVARIANCE ==============================================
558 % ===================================================================================
559 disp([10 'DEMO> Tutorial #21: Noise covariance' 10]);
560 % Process: Compute covariance (noise or data)
561 bst_process('CallProcess', 'process_noisecov', sFilesNoise, [], ...
562 'baseline', [], ...
563 'sensortypes', 'MEG, EEG, SEEG, ECOG', ...
564 'target', 1, ... % Noise covariance (covariance over baseline time window)
565 'dcoffset', 1, ... % Block by block, to avoid effects of slow shifts in data
566 'identity', 0, ...
567 'copycond', 1, ...
568 'copysubj', 0, ...
569 'replacefile', 1); % Replace
570 % Process: Snapshot: Noise covariance
571 bst_process('CallProcess', 'process_snapshot', sFilesNoise, [], ...
572 'target', 3, ... % Noise covariance
573 'Comment', 'Noise covariance');
574
575 % Process: Compute covariance (noise or data) [Run01]
576 bst_process('CallProcess', 'process_noisecov', sFilesEpochs1, [], ...
577 'baseline', [-0.1, -0.0017], ...
578 'datatimewindow', [0, 0.5], ...
579 'sensortypes', 'MEG, EEG, SEEG, ECOG', ...
580 'target', 2, ... % Data covariance (covariance over data time window)
581 'dcoffset', 1, ... % Block by block, to avoid effects of slow shifts in data
582 'identity', 0, ...
583 'copycond', 0, ...
584 'copysubj', 0, ...
585 'replacefile', 1); % Replace
586 % Process: Compute covariance (noise or data) [Run02]
587 bst_process('CallProcess', 'process_noisecov', sFilesEpochs2, [], ...
588 'baseline', [-0.1, -0.0017], ...
589 'datatimewindow', [0, 0.5], ...
590 'sensortypes', 'MEG, EEG, SEEG, ECOG', ...
591 'target', 2, ... % Data covariance (covariance over data time window)
592 'dcoffset', 1, ... % Block by block, to avoid effects of slow shifts in data
593 'identity', 0, ...
594 'copycond', 0, ...
595 'copysubj', 0, ...
596 'replacefile', 1); % Replace
597 % Process: Snapshot: Data covariance
598 bst_process('CallProcess', 'process_snapshot', [sFilesEpochs1(1), sFilesEpochs2(1)], [], ...
599 'target', 12, ... % Data covariance
600 'Comment', 'Data covariance');
601
602
603
604 %% ===== TUTORIAL #22: SOURCE ESTIMATION =============================================
605 % ===================================================================================
606 disp([10 'DEMO> Tutorial #22: Source estimation' 10]);
607 % === GET DEVIANT AVERAGE RUN01 ===
608 % Process: Select recordings in: Subject01/S01_AEF_20131218_01_600Hz_notch
609 sFiles01 = bst_process('CallProcess', 'process_select_files_data', [], [], ...
610 'subjectname', SubjectName, ...
611 'condition', 'S01_AEF_20131218_01_600Hz_notch', ...
612 'includebad', 0);
613 % Process: Select file comments with tag: deviant
614 sFilesAvgDeviant01 = bst_process('CallProcess', 'process_select_tag', sFiles01, [], ...
615 'tag', 'Avg: deviant', ...
616 'search', 2, ... % Search the file comments
617 'select', 1); % Select only the files with the tag
618
619 % === CONSTRAINED EXAMPLE ===
620 % Minimum norm options
621 InverseOptions = struct(...
622 'Comment', 'MN: MEG', ...
623 'InverseMethod', 'minnorm', ...
624 'InverseMeasure', 'amplitude', ...
625 'SourceOrient', {{'fixed'}}, ...
626 'Loose', 0.2, ...
627 'UseDepth', 1, ...
628 'WeightExp', 0.5, ...
629 'WeightLimit', 10, ...
630 'NoiseMethod', 'reg', ...
631 'NoiseReg', 0.1, ...
632 'SnrMethod', 'fixed', ...
633 'SnrRms', 0.001, ...
634 'SnrFixed', 3, ...
635 'ComputeKernel', 1, ...
636 'DataTypes', {{'MEG'}});
637 % Process: Compute sources [2018]
638 sFilesSrcDeviant01 = bst_process('CallProcess', 'process_inverse_2018', sFilesAvgDeviant01, [], ...
639 'output', 2, ... % Kernel only: one per file
640 'inverse', InverseOptions);
641
642 % === DISPLAY SOURCES MANUALLY ===
643 % View time series
644 hFigSrc1 = view_timeseries(sFilesAvgDeviant01(1).FileName, 'MEG');
645 % View on the cortex surface
646 hFigSrc2 = script_view_sources(sFilesSrcDeviant01.FileName, 'cortex');
647 % Set current time to 90ms
648 panel_time('SetCurrentTime', 0.090);
649 % Set orientation
650 figure_3d('SetStandardView', hFigSrc2, 'left');
651 % Set surface threshold
652 iSurf = 1;
653 thresh = .30;
654 panel_surface('SetDataThreshold', hFigSrc2, iSurf, thresh);
655 % Set surface smoothing
656 panel_surface('SetSurfaceSmooth', hFigSrc2, iSurf, .6, 0);
657 % Show sulci
658 panel_surface('SetShowSulci', hFigSrc2, iSurf, 1);
659
660 % View sources on MRI (3D orthogonal slices)
661 hFigSrc3 = script_view_sources(sFilesSrcDeviant01.FileName, 'mri3d');
662 panel_surface('SetDataThreshold', hFigSrc3, iSurf, thresh);
663 % Set the position of the cuts in the 3D figure
664 cutsPosVox = [74 93 159];
665 panel_surface('PlotMri', hFigSrc3, cutsPosVox);
666
667 % View sources with MRI Viewer
668 hFigSrc4 = script_view_sources(sFilesSrcDeviant01.FileName, 'mriviewer');
669 panel_surface('SetDataThreshold', hFigSrc4, iSurf, thresh);
670 % Set the position of the cuts in the MRI Viewer (values in millimeters)
671 figure_mri('SetLocation', 'voxel', hFigSrc4, [], cutsPosVox);
672 % Close figures
673 close([hFigSrc1 hFigSrc2 hFigSrc3 hFigSrc4]);
674
675 % === UNCONSTRAINED EXAMPLE ===
676 % Unconstrained minnorm
677 InverseOptions.Comment = 'MN: MEG';
678 InverseOptions.InverseMeasure = 'amplitude';
679 InverseOptions.SourceOrient = {'free'};
680 % Process: Compute sources [2018]
681 sFilesSrcUnconst = bst_process('CallProcess', 'process_inverse_2018', sFilesAvgDeviant01, [], ...
682 'output', 2, ... % Kernel only: one per file
683 'inverse', InverseOptions);
684
685
686 % === NORMALIZED SOURCES ===
687 % dSPM
688 InverseOptions.Comment = 'dSPM: MEG';
689 InverseOptions.InverseMeasure = 'dspm2018';
690 InverseOptions.SourceOrient = {'fixed'};
691 sFilesSrcDspm = bst_process('CallProcess', 'process_inverse_2018', sFilesAvgDeviant01, [], ...
692 'output', 2, ... % Kernel only: one per file
693 'inverse', InverseOptions);
694 % sLORETA (old function)
695 sFilesSrcSloreta = bst_process('CallProcess', 'process_inverse', sFilesAvgDeviant01, [], ...
696 'comment', '', ...
697 'method', 3, ... % sLORETA
698 'wmne', struct(...
699 'SourceOrient', {{'fixed'}}, ...
700 'loose', 0.2, ...
701 'SNR', 3, ...
702 'pca', 1, ...
703 'diagnoise', 0, ...
704 'regnoise', 1, ...
705 'magreg', 0.1, ...
706 'gradreg', 0.1, ...
707 'depth', 1, ...
708 'weightexp', 0.5, ...
709 'weightlimit', 10), ...
710 'sensortypes', 'MEG, MEG MAG, MEG GRAD, EEG', ...
711 'output', 2); % Kernel only: one per file
712 % Process: Z-score normalization: [-100ms,-2ms]
713 sFilesSrcZscore = bst_process('CallProcess', 'process_baseline_norm', sFilesSrcDeviant01, [], ...
714 'baseline', [-0.100, -0.002], ...
715 'source_abs', 0, ...
716 'method', 'zscore'); % Z-score transformation: x_std = (x - μ) / σ
717
718 % Process: Snapshot: Sources (one time)
719 bst_process('CallProcess', 'process_snapshot', sFilesSrcDeviant01, [], ...
720 'target', 8, ... % Sources (one time)
721 'orient', 1, ... % left
722 'time', 0.09, ...
723 'threshold', 30, ...
724 'Comment', 'Current density map (Constrained)');
725 bst_process('CallProcess', 'process_snapshot', sFilesSrcDspm, [], ...
726 'target', 8, ... % Sources (one time)
727 'orient', 1, ... % left
728 'time', 0.09, ...
729 'threshold', 60, ...
730 'Comment', 'dSPM');
731 bst_process('CallProcess', 'process_snapshot', sFilesSrcSloreta, [], ...
732 'target', 8, ... % Sources (one time)
733 'orient', 1, ... % left
734 'time', 0.09, ...
735 'threshold', 60, ...
736 'Comment', 'sLORETA');
737 bst_process('CallProcess', 'process_snapshot', sFilesSrcZscore, [], ...
738 'target', 8, ... % Sources (one time)
739 'orient', 1, ... % left
740 'time', 0.09, ...
741 'threshold', 60, ...
742 'Comment', 'Z-score');
743 bst_process('CallProcess', 'process_snapshot', sFilesSrcUnconst, [], ...
744 'target', 8, ... % Sources (one time)
745 'orient', 1, ... % left
746 'time', 0.0917, ...
747 'threshold', 0, ...
748 'Comment', 'Current density map (Unconstrained)');
749
750 % === DELETE EXPERIMENTS ===
751 % Process: Delete constrained example
752 bst_process('CallProcess', 'process_delete', [sFilesSrcDeviant01, sFilesSrcDspm, sFilesSrcSloreta, sFilesSrcZscore, sFilesSrcUnconst], [], ...
753 'target', 1); % Delete selected files
754
755
756 % === SHARED KERNEL ===
757 % Constrained minnorm
758 InverseOptions.Comment = 'MN: MEG';
759 InverseOptions.InverseMeasure = 'amplitude';
760 InverseOptions.SourceOrient = {'fixed'};
761 % Process: Compute sources [2018]
762 sFilesAvgSrc = bst_process('CallProcess', 'process_inverse_2018', sFilesAvg, [], ...
763 'output', 1, ... % Kernel only: shared
764 'inverse', InverseOptions);
765
766
767 % === AVERAGE SOURCES ACROSS RUNS ===
768 % Process: Average: By trial group (subject average)
769 sFilesIntraSrc = bst_process('CallProcess', 'process_average', sFilesAvgSrc, [], ...
770 'avgtype', 6, ... % By trial group (subject average)
771 'avg_func', 1, ... % Arithmetic average: mean(x)
772 'weighted', 1, ...
773 'scalenormalized', 0);
774 % Process: Low-pass:40Hz
775 sFilesIntraSrcLow = bst_process('CallProcess', 'process_bandpass', sFilesIntraSrc, [], ...
776 'highpass', 0, ...
777 'lowpass', 40, ...
778 'attenuation', 'strict', ... % 60dB
779 'mirror', 0, ...
780 'overwrite', 0);
781 % Process: Z-score normalization: [-100ms,-2ms]
782 sFilesIntraZscore = bst_process('CallProcess', 'process_baseline_norm', sFilesIntraSrcLow, [], ...
783 'baseline', [-0.100, -0.0017], ...
784 'source_abs', 0, ...
785 'method', 'zscore'); % Z-score transformation: x_std = (x - μ) / σ
786
787 % Process: Delete intermediate results
788 bst_process('CallProcess', 'process_delete', sFilesIntraSrcLow, [], ...
789 'target', 1); % Delete selected files
790 % Screen captures
791 bst_process('CallProcess', 'process_snapshot', sFilesIntraZscore, [], ...
792 'target', 8, ... % Sources (one time)
793 'orient', 1, ... % left
794 'time', 0.09, ...
795 'threshold', 40, ...
796 'Comment', 'Average across runs (left)');
797 bst_process('CallProcess', 'process_snapshot', sFilesIntraZscore, [], ...
798 'target', 8, ... % Sources (one time)
799 'orient', 2, ... % right
800 'time', 0.09, ...
801 'threshold', 40, ...
802 'Comment', 'Average across runs (right)');
803 bst_process('CallProcess', 'process_snapshot', sFilesIntraZscore(1), [], ...
804 'target', 9, ... % Sources (contact sheet)
805 'orient', 1, ... % left
806 'contact_time', [0, 0.35], ...
807 'contact_nimage', 15, ...
808 'threshold', 20, ...
809 'Comment', 'Average across runs');
810
811
812 %% ===== TUTORIAL #23: SCOUTS ========================================================
813 % ===================================================================================
814 disp([10 'DEMO> Tutorial #23: Scouts' 10]);
815 % Load surface file
816 sCortex = in_tess_bst(CortexFile);
817 % Add new scouts in first atlas
818 sCortex.iAtlas = find(strcmpi({sCortex.Atlas.Name}, 'Destrieux'));
819 % Save file
820 bst_save(file_fullpath(CortexFile), sCortex, 'v7');
821 % Unload everything
822 bst_memory('UnloadAll', 'Forced');
823 % Find scouts indices to display: {'A1L', 'A1R', 'IFGL', 'M1L'}
824 [tmp,iScouts,tmp] = intersect({sCortex.Atlas(sCortex.iAtlas).Scouts.Label}, {'G_temp_sup-G_T_transv L', 'G_temp_sup-G_T_transv R', 'G_front_inf-Opercular L', 'G_precentral L'});
825
826 % View cortex
827 hFigSurf1 = view_surface(CortexFile, [], [], 'NewFigure');
828 hFigSurf2 = view_surface(CortexFile, [], [], 'NewFigure');
829 figure_3d('SetStandardView', hFigSurf1, 'left');
830 figure_3d('SetStandardView', hFigSurf2, 'right');
831 panel_surface('SetSurfaceSmooth', hFigSurf1, 1, .6, 0);
832 panel_surface('SetSurfaceSmooth', hFigSurf2, 1, .6, 0);
833 panel_surface('SetShowSulci', hFigSurf1, 1, 1);
834 panel_surface('SetShowSulci', hFigSurf2, 1, 1);
835 % Configure scouts display
836 panel_scout('SetScoutsOptions', 0, 1, 1, 'all', 0.7, 1, 1, 0);
837 % View scouts
838 hFigScouts = view_scouts({sFilesIntraZscore.FileName}, iScouts);
839 hLegend = findobj(hFigScouts, 'Type', 'legend');
840 if ~isempty(hLegend) && ishandle(hLegend(1))
841 set(hLegend(1), 'Units', 'pixels');
842 pos = get(hLegend(1), 'Position');
843 set(hLegend(1), 'Position', [1, 1, pos(3), pos(4)]);
844 end
845 % Save figures
846 bst_report('Snapshot', hFigScouts, sFilesIntraZscore(1).FileName, 'Scouts', [100 100 670 250]);
847 % Close all
848 pause(1);
849 bst_memory('UnloadAll', 'Forced');
850
851
852
853 %% ===== TUTORIAL #24: TIME-FREQUENCY ================================================
854 % ===================================================================================
855 disp([10 'DEMO> Tutorial #24: Time-frequency' 10]);
856 % Process: Simulate generic signals
857 sSim = bst_process('CallProcess', 'process_simulate_matrix', [], [], ...
858 'subjectname', 'Test', ...
859 'condition', 'Simulation', ...
860 'samples', 6000, ...
861 'srate', 1000, ...
862 'matlab', ['f1 = 2; f2 = 20; f3 = 50;' 10 'i =2000:6000;' 10 'Data(1,i) = sin(f1*2*pi*t(i)) + 0.4 * cos(f2*2*pi*t(i));' 10 'Data = Data + 0.2 * sin(f3*2*pi*t) + 0.4 * rand(1,6000);']);
863
864 % Time-frequency options
865 TfOptions = struct(...
866 'Comment', 'Power,1-60Hz', ...
867 'TimeBands', [], ...
868 'Freqs', [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60], ...
869 'MorletFc', 1, ...
870 'MorletFwhmTc', 3, ...
871 'ClusterFuncTime', 'none', ...
872 'Measure', 'power', ...
873 'Output', 'average', ...
874 'SaveKernel', 0);
875 % Process: Time-frequency (Morlet wavelets)
876 sSimTf1 = bst_process('CallProcess', 'process_timefreq', sSim, [], ...
877 'edit', TfOptions, ...
878 'normalize', 'multiply'); % 1/f compensation: Multiply output values by frequency
879
880 % === NORMALIZATION ===
881 % Process: Time-frequency (Morlet wavelets)
882 sSimTf2 = bst_process('CallProcess', 'process_timefreq', sSim, [], ...
883 'edit', TfOptions, ...
884 'normalize', 'none'); % None: Save non-standardized time-frequency maps
885 % Process: Spectral flattening
886 sSimTf2Multi = bst_process('CallProcess', 'process_tf_norm', sSimTf2, [], ...
887 'normalize', 'multiply', ... % 1/f compensation (multiple by frequency)
888 'overwrite', 0);
889 % Process: Event-related perturbation (ERS/ERD): [750ms,1250ms]
890 sSimTf2Ersd = bst_process('CallProcess', 'process_baseline_norm', sSimTf2, [], ...
891 'baseline', [0.75, 1.25], ...
892 'method', 'ersd', ... % Event-related perturbation (ERS/ERD): x_std = (x - μ) / μ * 100
893 'overwrite', 0);
894
895 % Process: Snapshot: Time-frequency maps
896 bst_process('CallProcess', 'process_snapshot', sSimTf2, [], ...
897 'target', 14, ... % Time-frequency maps
898 'Comment', 'Not normalized');
899 % Process: Snapshot: Time-frequency maps
900 bst_process('CallProcess', 'process_snapshot', sSimTf2Multi, [], ...
901 'target', 14, ... % Time-frequency maps
902 'Comment', 'Spectral flattening: 1/f compensation');
903 % Process: Snapshot: Time-frequency maps
904 bst_process('CallProcess', 'process_snapshot', sSimTf2Ersd, [], ...
905 'target', 14, ... % Time-frequency maps
906 'Comment', 'ERS/ERD');
907 % Spectrum/Time series
908 hFigTf1 = view_spectrum(sSimTf2.FileName, 'Spectrum');
909 hFigTf2 = view_spectrum(sSimTf2.FileName, 'TimeSeries');
910 panel_time('SetCurrentTime', 0.5);
911 panel_freq('SetCurrentFreq', 2, 0);
912 bst_report('Snapshot', [hFigTf1 hFigTf2], sSimTf2.FileName, 'Not normalized: 2s/20Hz', [200, 200, 400, 250]);
913 panel_time('SetCurrentTime', 2.02);
914 panel_freq('SetCurrentFreq', 20, 0);
915 bst_report('Snapshot', [hFigTf1 hFigTf2], sSimTf2.FileName, 'Not normalized: 2s/20Hz', [200, 200, 400, 250]);
916 bst_memory('UnloadAll', 'Forced');
917
918
919 % === HILBERT TRANSFORM ===
920 % Process: Hilbert transform
921 sSimHilbert = bst_process('CallProcess', 'process_hilbert', sSim, [], ...
922 'edit', struct(...
923 'Comment', 'Power', ...
924 'TimeBands', [], ...
925 'Freqs', {{'delta', '2, 4', 'mean'; 'theta', '5, 7', 'mean'; 'alpha', '8, 12', 'mean'; 'beta', '15, 29', 'mean'; 'gamma1', '30, 59', 'mean'; 'gamma2', '60, 90', 'mean'}}, ...
926 'ClusterFuncTime', 'none', ...
927 'Measure', 'power', ...
928 'Output', 'all', ...
929 'SaveKernel', 0), ...
930 'normalize', 'none', ... % None: Save non-standardized time-frequency maps
931 'mirror', 0);
932 % Process: Spectral flattening
933 sSimHilbertMulti = bst_process('CallProcess', 'process_tf_norm', sSimHilbert, [], ...
934 'normalize', 'multiply', ... % 1/f compensation (multiple by frequency)
935 'overwrite', 0);
936 % Process: Event-related perturbation (ERS/ERD): [750ms,1250ms]
937 sSimHilbertErsd = bst_process('CallProcess', 'process_baseline_norm', sSimHilbert, [], ...
938 'baseline', [0.75, 1.25], ...
939 'method', 'ersd', ... % Event-related perturbation (ERS/ERD): x_std = (x - μ) / μ * 100
940 'overwrite', 0);
941
942 % Process: Snapshot: Time-frequency maps
943 bst_process('CallProcess', 'process_snapshot', sSimHilbert, [], ...
944 'target', 14, ... % Time-frequency maps
945 'Comment', 'Not normalized');
946 % Process: Snapshot: Time-frequency maps
947 bst_process('CallProcess', 'process_snapshot', sSimHilbertMulti, [], ...
948 'target', 14, ... % Time-frequency maps
949 'Comment', 'Spectral flattening: 1/f compensation');
950 % Process: Snapshot: Time-frequency maps
951 bst_process('CallProcess', 'process_snapshot', sSimHilbertErsd, [], ...
952 'target', 14, ... % Time-frequency maps
953 'Comment', 'ERS/ERD');
954
955 % === SINGLE TRIALS ===
956 TfOptions.Comment = 'Avg,Power,1-150Hz';
957 TfOptions.Freqs = [1, 2, 3.1, 4.2, 5.4, 6.7, 8, 9.5, 11, 12.6, 14.3, 16.1, 18.1, 20.1, 22.3, 24.6, 27, 29.6, 32.4, 35.3, 38.4, 41.6, 45.1, 48.8, 52.7, 56.9, 61.3, 66, 70.9, 76.2, 81.8, 87.7, 94, 100.6, 107.7, 115.2, 123.1, 131.6, 140.5, 150];
958 % Process: Time-frequency (Morlet wavelets)
959 sEpochsAvgTf = bst_process('CallProcess', 'process_timefreq', sFilesEpochs1, [], ...
960 'sensortypes', 'MEG, EEG', ...
961 'edit', TfOptions, ...
962 'normalize', 'none'); % None: Save non-standardized time-frequency maps
963 % Process: Event-related perturbation (ERS/ERD): [-75ms,0ms]
964 sEpochsAvgTfErsd = bst_process('CallProcess', 'process_baseline_norm', sEpochsAvgTf, [], ...
965 'baseline', [-0.075, 0], ...
966 'method', 'ersd', ... % Event-related perturbation (ERS/ERD): x_std = (x - μ) / μ * 100
967 'overwrite', 0);
968
969 % === DISPLAY ===
970 % View time-frequency file
971 hFigTf1 = view_timefreq(sEpochsAvgTfErsd.FileName, 'SingleSensor');
972 % Configure display
973 sOptions = panel_display('GetDisplayOptions');
974 sOptions.HideEdgeEffects = 1;
975 sOptions.HighResolution = 1;
976 panel_display('SetDisplayOptions', sOptions);
977 % Other display modes
978 hFigTf2 = view_timefreq(sEpochsAvgTfErsd.FileName, 'AllSensors');
979 hFigTf3 = view_timefreq(sEpochsAvgTfErsd.FileName, '2DLayout');
980 hFigTf4 = view_timefreq(sEpochsAvgTfErsd.FileName, '2DLayoutOpt');
981 bst_colormaps('SetColormapName', 'stat2', 'jet');
982 bst_colormaps('SetColormapAbsolute', 'stat2', 1);
983 bst_report('Snapshot', hFigTf1, sEpochsAvgTfErsd.FileName, 'Time-frequency', [200, 200, 400, 250]);
984 bst_report('Snapshot', [hFigTf2 hFigTf3 hFigTf4], sEpochsAvgTfErsd.FileName, 'Time-frequency', [200, 200, 750, 400]);
985 close([hFigTf1 hFigTf2 hFigTf3 hFigTf4]);
986 % Image [channel x time]
987 hFigTf5 = view_erpimage(sEpochsAvgTfErsd.FileName, 'trialimage');
988 % Topographies
989 hFigTf6 = view_topography(sEpochsAvgTfErsd.FileName, 'MEG', '3DSensorCap');
990 hFigTf7 = view_topography(sEpochsAvgTfErsd.FileName, 'MEG', '2DDisc');
991 hFigTf8 = view_topography(sEpochsAvgTfErsd.FileName, 'MEG', '2DSensorCap');
992 hFigTf9 = view_topography(sEpochsAvgTfErsd.FileName, 'MEG', '2DLayout');
993 panel_time('SetCurrentTime', 0.175);
994 panel_freq('SetCurrentFreq', 8, 0);
995 bst_report('Snapshot', [hFigTf5 hFigTf6 hFigTf7 hFigTf8 hFigTf9], sEpochsAvgTfErsd.FileName, 'Time-frequency: 8Hz', [200, 200, 400, 250]);
996 close([hFigTf5 hFigTf6 hFigTf7 hFigTf8 hFigTf9]);
997
998
999 % === AVERAGE FOR SCOUTS ===
1000 % Select all sources for the single deviant epochs
1001 sFilesEpochDeviantSrc = bst_process('CallProcess', 'process_select_files_results', [], [], ...
1002 'subjectname', SubjectName, ...
1003 'condition', '', ...
1004 'includebad', 0);
1005 sFilesEpochDeviantSrc = bst_process('CallProcess', 'process_select_tag', sFilesEpochDeviantSrc, [], ...
1006 'tag', 'deviant', ...
1007 'search', 1, ... % Search the file names
1008 'select', 1); % Select only the files with the tag
1009 sFilesEpochDeviantSrc = bst_process('CallProcess', 'process_select_tag', sFilesEpochDeviantSrc, [], ...
1010 'tag', 'average', ...
1011 'search', 1, ... % Search the file names
1012 'select', 2); % Exclude the files with the tag
1013
1014 % Process: Time-frequency (Morlet wavelets)
1015 sFilesTfScout = bst_process('CallProcess', 'process_timefreq', sFilesEpochDeviantSrc, [], ...
1016 'clusters', {'Destrieux', {'G_temp_sup-G_T_transv L', 'G_temp_sup-G_T_transv R', 'G_front_inf-Opercular L', 'G_precentral L'}}, ...
1017 'scoutfunc', 1, ... % Mean
1018 'edit', struct(...
1019 'Comment', 'Deviant: Scouts,Avg,Power,1-150Hz', ...
1020 'TimeBands', [], ...
1021 'Freqs', [1, 2, 3.1, 4.2, 5.4, 6.7, 8, 9.5, 11, 12.6, 14.3, 16.1, 18.1, 20.1, 22.3, 24.6, 27, 29.6, 32.4, 35.3, 38.4, 41.6, 45.1, 48.8, 52.7, 56.9, 61.3, 66, 70.9, 76.2, 81.8, 87.7, 94, 100.6, 107.7, 115.2, 123.1, 131.6, 140.5, 150], ...
1022 'MorletFc', 1, ...
1023 'MorletFwhmTc', 3, ...
1024 'ClusterFuncTime', 'after', ...
1025 'Measure', 'power', ...
1026 'Output', 'average', ...
1027 'SaveKernel', 0), ...
1028 'normalize', 'none'); % None: Save non-standardized time-frequency maps)
1029 % Process: Event-related perturbation (ERS/ERD): [-75ms,0ms]
1030 sFilesTfScoutErsd = bst_process('CallProcess', 'process_baseline_norm', sFilesTfScout, [], ...
1031 'baseline', [-0.075, 0], ...
1032 'method', 'ersd', ... % Event-related perturbation (ERS/ERD): x_std = (x - μ) / μ * 100
1033 'overwrite', 0);
1034 % Process: Snapshot: Time-frequency maps
1035 bst_process('CallProcess', 'process_snapshot', sFilesTfScoutErsd, [], ...
1036 'target', 14, ... % Time-frequency maps
1037 'Comment', 'ERS/ERD');
1038
1039
1040 % === FULL CORTEX / HILBERT ===
1041 % Process: Hilbert transform
1042 sFilesHilbertCortex = bst_process('CallProcess', 'process_hilbert', sFilesEpochDeviantSrc, [], ...
1043 'clusters', {}, ...
1044 'scoutfunc', 1, ... % Mean
1045 'edit', struct(...
1046 'Comment', 'Deviant: Avg,Magnitude', ...
1047 'TimeBands', [], ...
1048 'Freqs', {{'delta', '2, 4', 'mean'; 'theta', '5, 7', 'mean'; 'alpha', '8, 12', 'mean'; 'beta', '15, 29', 'mean'; 'gamma1', '30, 59', 'mean'; 'gamma2', '60, 90', 'mean'}}, ...
1049 'ClusterFuncTime', 'none', ...
1050 'Measure', 'power', ...
1051 'Output', 'average', ...
1052 'RemoveEvoked', 0, ...
1053 'SaveKernel', 0), ...
1054 'normalize', 'none', ... % None: Save non-standardized time-frequency maps
1055 'mirror', 0);
1056 % Process: Event-related perturbation (ERS/ERD): [-75ms,0ms]
1057 sFilesHilbertCortexErsd = bst_process('CallProcess', 'process_baseline_norm', sFilesHilbertCortex, [], ...
1058 'baseline', [-0.075, 0], ...
1059 'method', 'ersd', ... % Event-related perturbation (ERS/ERD): x_std = (x - μ) / μ * 100
1060 'overwrite', 0);
1061
1062 % View results
1063 hFigTf1 = view_surface_data([], sFilesHilbertCortexErsd.FileName);
1064 hFigTf2 = view_timefreq(sFilesHilbertCortexErsd.FileName, 'SingleSensor', 362);
1065 figure_3d('SetStandardView', hFigTf1, 'left');
1066 panel_surface('SetDataThreshold', hFigTf1, 1, 0.5);
1067 panel_time('SetCurrentTime', 0.175);
1068 panel_freq('SetCurrentFreq', 3);
1069 bst_colormaps('RestoreDefaults', 'stat2');
1070 bst_report('Snapshot', [hFigTf1 hFigTf2], sFilesHilbertCortexErsd.FileName, 'Hilbert transform: Alpha band', [200, 200, 400, 250]);
1071 bst_memory('UnloadAll', 'Forced');
1072
1073
1074 %% ===== TUTORIAL #25: DIFFRERENCE ===================================================
1075 % ===================================================================================
1076 disp([10 'DEMO> Tutorial #25: Difference' 10]);
1077
1078 % ===== SELECT TRIALS (DATA) =====
1079 % Process: Select recordings in: Subject01/*
1080 sFilesAll = bst_process('CallProcess', 'process_select_files_data', [], [], ...
1081 'subjectname', SubjectName, ...
1082 'condition', '', ...
1083 'includebad', 0);
1084 % Process: Select file names with tag: deviant_trial
1085 sEpochDeviant = bst_process('CallProcess', 'process_select_tag', sFilesAll, [], ...
1086 'tag', 'deviant_trial', ...
1087 'search', 1, ... % Search the file names
1088 'select', 1); % Select only the files with the tag
1089 % Process: Select file names with tag: standard_trial
1090 sEpochStandard = bst_process('CallProcess', 'process_select_tag', sFilesAll, [], ...
1091 'tag', 'standard_trial', ...
1092 'search', 1, ... % Search the file names
1093 'select', 1); % Select only the files with the tag
1094
1095 % ===== SELECT TRIALS (SOURCE) =====
1096 % Process: Select recordings in: Subject01/*
1097 sFilesAllSrc = bst_process('CallProcess', 'process_select_files_results', [], [], ...
1098 'subjectname', SubjectName, ...
1099 'condition', '', ...
1100 'includebad', 0);
1101 % Process: Select file names with tag: deviant_trial
1102 sEpochDeviantSrc = bst_process('CallProcess', 'process_select_tag', sFilesAllSrc, [], ...
1103 'tag', 'deviant_trial', ...
1104 'search', 1, ... % Search the file names
1105 'select', 1); % Select only the files with the tag
1106 % Process: Select file names with tag: standard_trial
1107 sEpochStandardSrc = bst_process('CallProcess', 'process_select_tag', sFilesAllSrc, [], ...
1108 'tag', 'standard_trial', ...
1109 'search', 1, ... % Search the file names
1110 'select', 1); % Select only the files with the tag
1111
1112 % ===== ABSOLUTE DIFFERENCE ======
1113 % Process: Difference: A-B, abs
1114 sDiffSrc = bst_process('CallProcess', 'process_diff_ab', sFilesIntraSrc(1).FileName, sFilesIntraSrc(2).FileName, ...
1115 'source_abs', 1);
1116 % Process: Set comment: deviant|abs - standard|abs
1117 sDiffSrc = bst_process('CallProcess', 'process_set_comment', sDiffSrc, [], ...
1118 'tag', 'deviant|abs - standard|abs', ...
1119 'isindex', 1);
1120 % Process: Low-pass:40Hz
1121 sDiffSrcZscore = bst_process('CallProcess', 'process_bandpass', sDiffSrc, [], ...
1122 'highpass', 0, ...
1123 'lowpass', 40, ...
1124 'attenuation', 'strict', ... % 60dB
1125 'mirror', 0, ...
1126 'overwrite', 1);
1127 % Process: Z-score transformation: [-100ms,-2ms]
1128 sDiffSrcZscore = bst_process('CallProcess', 'process_baseline_norm', sDiffSrcZscore, [], ...
1129 'baseline', [-0.1, -0.002], ...
1130 'source_abs', 0, ...
1131 'method', 'zscore', ... % Z-score transformation: x_std = (x - μ) / σ
1132 'overwrite', 0);
1133 % Process: Snapshot: Sources (contact sheet)
1134 bst_process('CallProcess', 'process_snapshot', sDiffSrcZscore, [], ...
1135 'target', 9, ... % Sources (contact sheet)
1136 'modality', 1, ... % MEG (All)
1137 'orient', 1, ... % left
1138 'contact_time', [0, 0.35], ...
1139 'contact_nimage', 15, ...
1140 'threshold', 30, ...
1141 'Comment', 'Difference deviant - standard (absolute)');
1142
1143 % ===== RELATIVE DIFFERENCE =====
1144 % Process: Difference: A-B
1145 sDiffSrcRel = bst_process('CallProcess', 'process_diff_ab', sFilesIntraSrc(1).FileName, sFilesIntraSrc(2).FileName, ...
1146 'source_abs', 0);
1147 % Process: Set comment: deviant - standard
1148 sDiffSrcRel = bst_process('CallProcess', 'process_set_comment', sDiffSrcRel, [], ...
1149 'tag', 'deviant - standard', ...
1150 'isindex', 1);
1151 % Process: Low-pass:40Hz
1152 sDiffSrcRelZscore = bst_process('CallProcess', 'process_bandpass', sDiffSrcRel, [], ...
1153 'highpass', 0, ...
1154 'lowpass', 40, ...
1155 'attenuation', 'strict', ... % 60dB
1156 'mirror', 0, ...
1157 'overwrite', 1);
1158 % Process: Z-score transformation: [-100ms,-2ms]
1159 sDiffSrcRelZscore = bst_process('CallProcess', 'process_baseline_norm', sDiffSrcRelZscore, [], ...
1160 'baseline', [-0.1, -0.002], ...
1161 'source_abs', 0, ...
1162 'method', 'zscore', ... % Z-score transformation: x_std = (x - μ) / σ
1163 'overwrite', 0);
1164 % Configure colormap: hot/absolute
1165 bst_colormaps('SetColormapName', 'stat2', 'hot');
1166 bst_colormaps('SetColormapAbsolute', 'stat2', 1);
1167 % Process: Snapshot: Sources (contact sheet)
1168 bst_process('CallProcess', 'process_snapshot', sDiffSrcRelZscore, [], ...
1169 'target', 9, ... % Sources (contact sheet)
1170 'modality', 1, ... % MEG (All)
1171 'orient', 1, ... % left
1172 'contact_time', [0, 0.35], ...
1173 'contact_nimage', 15, ...
1174 'threshold', 30, ...
1175 'Comment', 'Difference deviant - standard (relative)');
1176 % Restore colormap: rwb/relative
1177 bst_colormaps('SetColormapName', 'stat2', 'cmap_rbw');
1178 bst_colormaps('SetColormapAbsolute', 'stat2', 0);
1179
1180 % ===== DIFFERENCE OF MEANS =====
1181 % Process: Select uniform number of files [uniform]
1182 [sEpochDeviantUni, sEpochStandardUni] = bst_process('CallProcess', 'process_select_uniform2', sEpochDeviant, sEpochStandard, ...
1183 'nfiles', 0, ...
1184 'method', 4); % Uniformly distributed
1185 % Process: Difference of means [mean]
1186 sDiffMean = bst_process('CallProcess', 'process_diff_mean', sEpochDeviantUni, sEpochStandardUni, ...
1187 'avg_func', 1, ... % Arithmetic average mean(A) - mean(B)
1188 'weighted', 0);
1189 % Process: Snapshot: Recordings time series
1190 bst_process('CallProcess', 'process_snapshot', sDiffMean, [], ...
1191 'target', 5, ... % Recordings time series
1192 'modality', 1, ... % MEG (All)
1193 'Comment', 'Difference of means');
1194
1195
1196
1197 %% ===== TUTORIAL #26: STATISTICS ====================================================
1198 % ===================================================================================
1199 disp([10 'DEMO> Tutorial #26: Statistics' 10]);
1200 % ===== HISTOGRAMS =====
1201 % Process: Extract values: [160ms] MLP57
1202 sHistoDeviant = bst_process('CallProcess', 'process_extract_values', sEpochDeviant, [], ...
1203 'timewindow', [0.16, 0.16], ...
1204 'sensortypes', 'MLP57', ...
1205 'isabs', 0, ...
1206 'avgtime', 0, ...
1207 'avgrow', 0, ...
1208 'dim', 2, ... % Concatenate time (dimension 2)
1209 'Comment', '');
1210 % Process: Extract values: [160ms] MLP57
1211 sHistoStandard = bst_process('CallProcess', 'process_extract_values', sEpochStandard, [], ...
1212 'timewindow', [0.16, 0.16], ...
1213 'sensortypes', 'MLP57', ...
1214 'isabs', 0, ...
1215 'avgtime', 0, ...
1216 'avgrow', 0, ...
1217 'dim', 2, ... % Concatenate time (dimension 2)
1218 'Comment', '');
1219 % Display histograms
1220 hFigHisto = view_histogram({sHistoDeviant.FileName, sHistoStandard.FileName});
1221 bst_report('Snapshot', hFigHisto, sHistoDeviant.FileName, 'Histograms for MLP57/160ms');
1222 close(hFigHisto);
1223
1224 % ===== EXEMPLE #1: PARAMETRIC/DATA =====
1225 % Process: t-test [equal] [-100ms,500ms] H0:(A-B = 0)
1226 sTestParamData = bst_process('CallProcess', 'process_test_parametric2', sEpochDeviant, sEpochStandard, ...
1227 'timewindow', [-0.1, 0.5], ...
1228 'sensortypes', '', ...
1229 'isabs', 0, ...
1230 'avgtime', 0, ...
1231 'avgrow', 0, ...
1232 'Comment', '', ...
1233 'test_type', 'ttest_equal', ... % Student's t-test (equal variance) t = (mean(A)-mean(B)) / (Sx * sqrt(1/nA + 1/nB))Sx = sqrt(((nA-1)*var(A) + (nB-1)*var(B)) / (nA+nB-2)) df = nA + nB - 2
1234 'tail', 'two'); % Two-tailed
1235 % Set display properties
1236 StatThreshOptions = bst_get('StatThreshOptions');
1237 StatThreshOptions.pThreshold = 0.05;
1238 StatThreshOptions.Correction = 'fdr';
1239 StatThreshOptions.Control = [1 2 3];
1240 bst_set('StatThreshOptions', StatThreshOptions);
1241 % Process: Snapshot: Recordings time series
1242 bst_process('CallProcess', 'process_snapshot', sTestParamData, [], ...
1243 'target', 5, ... % Recordings time series
1244 'modality', 1, ... % MEG (All)
1245 'time', 0.16, ...
1246 'Comment', 'Parametric t-test (p<0.05, FDR)');
1247 % Process: Snapshot: Recordings topography (one time)
1248 bst_process('CallProcess', 'process_snapshot', sTestParamData, [], ...
1249 'target', 6, ... % Recordings topography (one time)
1250 'modality', 1, ... % MEG (All)
1251 'time', 0.16, ...
1252 'Comment', 'Parametric t-test (p<0.05, FDR)');
1253
1254 % ===== EXEMPLE #2: NON-PARAMETRIC/DATA =====
1255 % Process: Perm t-test equal [-100ms,500ms MEG] H0:(A=B), H1:(A<>B)
1256 sTestPermData = bst_process('CallProcess', 'process_test_permutation2', sEpochDeviant, sEpochStandard, ...
1257 'timewindow', [-0.1, 0.5], ...
1258 'sensortypes', 'MEG', ...
1259 'isabs', 0, ...
1260 'avgtime', 0, ...
1261 'avgrow', 0, ...
1262 'iszerobad', 1, ...
1263 'Comment', '', ...
1264 'test_type', 'ttest_equal', ... % Student's t-test (equal variance) t = (mean(A)-mean(B)) / (Sx * sqrt(1/nA + 1/nB))Sx = sqrt(((nA-1)*var(A) + (nB-1)*var(B)) / (nA+nB-2))
1265 'randomizations', 1000, ...
1266 'tail', 'two'); % Two-tailed
1267 % Process: Snapshot: Recordings time series
1268 bst_process('CallProcess', 'process_snapshot', sTestPermData, [], ...
1269 'target', 5, ... % Recordings time series
1270 'modality', 1, ... % MEG (All)
1271 'time', 0.16, ...
1272 'Comment', 'Non-parametric t-test (p<0.05, FDR)');
1273 % Process: Snapshot: Recordings topography (one time)
1274 bst_process('CallProcess', 'process_snapshot', sTestPermData, [], ...
1275 'target', 6, ... % Recordings topography (one time)
1276 'modality', 1, ... % MEG (All)
1277 'time', 0.16, ...
1278 'Comment', 'Non-parametric t-test (p<0.05, FDR)');
1279
1280 % ===== EXEMPLE #3: CLUSTER/DATA =====
1281 % Process: FT t-test unequal cluster [-100ms,500ms MEG] H0:(A=B), H1:(A<>B)
1282 sTestClustData = bst_process('CallProcess', 'process_ft_timelockstatistics', sEpochDeviant, sEpochStandard, ...
1283 'sensortypes', 'MEG', ...
1284 'timewindow', [-0.1, 0.5], ...
1285 'isabs', 0, ...
1286 'avgtime', 0, ...
1287 'avgchan', 0, ...
1288 'randomizations', 1000, ...
1289 'statistictype', 1, ... % Independent t-test
1290 'tail', 'two', ... % Two-tailed
1291 'correctiontype', 2, ... % cluster
1292 'minnbchan', 0, ...
1293 'clusteralpha', 0.05);
1294 % Process: Snapshot: Recordings time series
1295 bst_process('CallProcess', 'process_snapshot', sTestClustData, [], ...
1296 'target', 5, ... % Recordings time series
1297 'time', 0.16, ...
1298 'modality', 1, ... % MEG (All)
1299 'Comment', 'Cluster-based permutation test');
1300 % Process: Snapshot: Recordings topography (one time)
1301 bst_process('CallProcess', 'process_snapshot', sTestClustData, [], ...
1302 'target', 6, ... % Recordings topography (one time)
1303 'modality', 1, ... % MEG (All)
1304 'time', 0.16, ...
1305 'Comment', 'Cluster-based permutation test');
1306
1307 % ===== EXAMPLE #4: PARAMETRIC/SOURCES =====
1308 % Process: t-test [equal] [-100ms,500ms] H0:(A-B = 0)
1309 sTestParamSrc = bst_process('CallProcess', 'process_test_parametric2', sEpochDeviantSrc, sEpochStandardSrc, ...
1310 'timewindow', [-0.1, 0.5], ...
1311 'scoutsel', {}, ...
1312 'scoutfunc', 1, ... % Mean
1313 'isnorm', 0, ...
1314 'avgtime', 0, ...
1315 'Comment', '', ...
1316 'test_type', 'ttest_equal', ... % Student's t-test (equal variance) t = (mean(A)-mean(B)) / (Sx * sqrt(1/nA + 1/nB))Sx = sqrt(((nA-1)*var(A) + (nB-1)*var(B)) / (nA+nB-2)) df = nA + nB - 2
1317 'tail', 'two'); % Two-tailed
1318 % Process: Difference of means [abs(mean)]
1319 sDiffMeanSrc = bst_process('CallProcess', 'process_diff_mean', sEpochDeviantSrc, sEpochStandardSrc, ...
1320 'avg_func', 2, ... % Absolute value of average abs(mean(A)) - abs(mean(B))
1321 'weighted', 0);
1322 % Process: Apply statistic threshold: p<0.05 (FDR:1,2,3)
1323 sDiffMeanSrcThresh = bst_process('CallProcess', 'process_extract_pthresh2', sTestParamSrc, sDiffMeanSrc, ...
1324 'pthresh', 0.05, ...
1325 'correction', 3, ... % False discovery rate (FDR)
1326 'control1', 1, ...
1327 'control2', 1, ...
1328 'control3', 1);
1329 % Process: Snapshot: Sources (one time)
1330 bst_process('CallProcess', 'process_snapshot', sTestParamSrc, [], ...
1331 'target', 8, ... % Sources (one time)
1332 'orient', 1, ... % left
1333 'time', 0.148, ...
1334 'threshold', 40, ...
1335 'Comment', 'Parametric t-test (p<0.05, FDR)');
1336 % Process: Snapshot: Sources (one time)
1337 bst_process('CallProcess', 'process_snapshot', sDiffMeanSrc, [], ...
1338 'target', 8, ... % Sources (one time)
1339 'orient', 1, ... % left
1340 'time', 0.148, ...
1341 'threshold', 40, ...
1342 'Comment', 'abs(average(deviant)) - abs(average(standard))');
1343 % Process: Snapshot: Sources (one time)
1344 bst_process('CallProcess', 'process_snapshot', sDiffMeanSrcThresh, [], ...
1345 'target', 8, ... % Sources (one time)
1346 'orient', 1, ... % left
1347 'time', 0.148, ...
1348 'threshold', 0, ...
1349 'Comment', 'Different of mean thresholded with t-test results');
1350
1351 % ===== EXAMPLE #5: PARAMETRIC/SCOUTS =====
1352 % Process: t-test equal [-100ms,500ms] H0:(A=B), H1:(A<>B)
1353 sTestParamScout = bst_process('CallProcess', 'process_test_parametric2', sEpochDeviantSrc, sEpochStandardSrc, ...
1354 'timewindow', [-0.1, 0.5], ...
1355 'scoutsel', {'Destrieux', {'G_front_inf-Opercular L', 'G_precentral L', 'G_temp_sup-G_T_transv L'}}, ...
1356 'scoutfunc', 1, ... % Mean
1357 'isnorm', 0, ...
1358 'avgtime', 0, ...
1359 'Comment', '', ...
1360 'test_type', 'ttest_equal', ... % Student's t-test (equal variance) t = (mean(A)-mean(B)) / (Sx * sqrt(1/nA + 1/nB))Sx = sqrt(((nA-1)*var(A) + (nB-1)*var(B)) / (nA+nB-2)) df = nA + nB - 2
1361 'tail', 'two'); % Two-tailed
1362 % Process: Apply statistic threshold: p<0.05 (FDR:1,2,3)
1363 sTestParamScoutThresh = bst_process('CallProcess', 'process_extract_pthresh', sTestParamScout, [], ...
1364 'pthresh', 0.05, ...
1365 'correction', 3, ... % False discovery rate (FDR)
1366 'control1', 1, ...
1367 'control2', 1, ...
1368 'control3', 1);
1369 % Process: Compute head model
1370 bst_process('CallProcess', 'process_headmodel', sTestParamScoutThresh, [], ...
1371 'sourcespace', 1, ... % Cortex surface
1372 'meg', 3); % Overlapping spheres
1373 % Process: Simulate recordings from scouts
1374 sSimulData = bst_process('CallProcess', 'process_simulate_recordings', sTestParamScoutThresh, [], ...
1375 'scouts', {'Destrieux', {'G_front_inf-Opercular L', 'G_precentral L', 'G_temp_sup-G_T_transv L'}}, ...
1376 'savesources', 1);
1377
1378 % Get corresponding source file
1379 [sStudy,iStudy,iRes] = bst_get('ResultsForDataFile', sSimulData.FileName);
1380 sSimulSrc = sStudy.Result(iRes).FileName;
1381 % Reset visualization filters
1382 panel_filter('SetFilters', 0, [], 0, [], 0, [], 0, 0);
1383 % Process: Snapshot: Recordings time series
1384 bst_process('CallProcess', 'process_snapshot', sTestParamScout, [], ...
1385 'target', 5, ... % Recordings time series
1386 'time', 0.148, ...
1387 'Comment', 'Parametric t-test (p<0.05, FDR)');
1388 % Process: Snapshot: Sources (one time)
1389 bst_process('CallProcess', 'process_snapshot', sSimulSrc, [], ...
1390 'target', 8, ... % Sources (one time)
1391 'orient', 1, ... % left
1392 'time', 0.148, ...
1393 'threshold', 0, ...
1394 'Comment', 'Simulated sources');
1395 % Process: Snapshot: Recordings time series
1396 bst_process('CallProcess', 'process_snapshot', sSimulData, [], ...
1397 'target', 5, ... % Recordings time series
1398 'modality', 1, ... % MEG (All)
1399 'time', 0.148, ...
1400 'Comment', 'Simulated MEG recordings');
1401
1402
1403 % ===== EXAMPLE #6: NON-PARAMETRIC/TIMEFREQ =====
1404 TfOptions.Output = 'all';
1405 % Process: Time-frequency (Morlet wavelets) / DEVIANT
1406 sEpochDeviantTf = bst_process('CallProcess', 'process_timefreq', sEpochDeviant, [], ...
1407 'sensortypes', 'MLP57', ...
1408 'edit', TfOptions, ...
1409 'normalize', 'none'); % None: Save non-standardized time-frequency maps
1410 % Process: Time-frequency (Morlet wavelets) / STANDARD
1411 sEpochStandardTf = bst_process('CallProcess', 'process_timefreq', sEpochStandard, [], ...
1412 'sensortypes', 'MLP57', ...
1413 'edit', TfOptions, ...
1414 'normalize', 'none'); % None: Save non-standardized time-frequency maps
1415 % Process: Perm t-test equal [-100ms,500ms 1-150Hz] H0:(A=B), H1:(A<>B)
1416 sTestTf = bst_process('CallProcess', 'process_test_permutation2', sEpochDeviantTf, sEpochStandardTf, ...
1417 'timewindow', [-0.1, 0.5], ...
1418 'freqrange', [1, 150], ...
1419 'rows', '', ...
1420 'isabs', 0, ...
1421 'avgtime', 0, ...
1422 'avgrow', 0, ...
1423 'avgfreq', 0, ...
1424 'matchrows', 0, ...
1425 'iszerobad', 1, ...
1426 'Comment', '', ...
1427 'test_type', 'ttest_equal', ... % Student's t-test (equal variance) t = (mean(A)-mean(B)) / (Sx * sqrt(1/nA + 1/nB))Sx = sqrt(((nA-1)*var(A) + (nB-1)*var(B)) / (nA+nB-2))
1428 'randomizations', 1000, ...
1429 'tail', 'two'); % Two-tailed
1430 % Set stat threshold
1431 StatThreshOptions.pThreshold = 0.05;
1432 StatThreshOptions.Correction = 'none';
1433 bst_set('StatThreshOptions', StatThreshOptions);
1434 % Process: Snapshot: Time-frequency maps
1435 bst_process('CallProcess', 'process_snapshot', sTestTf, [], ...
1436 'target', 14, ... % Time-frequency maps
1437 'Comment', 'Non-parametric t-test (p<0.05, Uncorrected)');
1438 % Process: Delete intermediate results
1439 bst_process('CallProcess', 'process_delete', [sEpochDeviantTf, sEpochStandardTf], [], ...
1440 'target', 1); % Delete selected files
1441
1442
1443
1444 %% ===== SAVE REPORT =====
1445 % Save and display report
1446 ReportFile = bst_report('Save', []);
1447 if ~isempty(reports_dir) && ~isempty(ReportFile)
1448 bst_report('Export', ReportFile, reports_dir);
1449 else
1450 bst_report('Open', ReportFile);
1451 end
1452
1453 disp([10 'DEMO> Done.' 10]);
MEG visual: single subject . brainstorm3/toolbox/script/tutorial_visual_single.m
1 function tutorial_visual_single(bids_dir, reports_dir)
2 % TUTORIAL_VISUAL_SINGLE: Runs the Brainstorm/SPM group analysis pipeline (single subject, BIDS version).
3 %
4 % ONLINE TUTORIALS: https://neuroimage.usc.edu/brainstorm/Tutorials/VisualSingle
5 %
6 % INPUTS:
7 % - bids_dir: Path to folder ds000117 (https://openneuro.org/datasets/ds000117)
8 % |- derivatives/freesurfer/sub-XX : Segmentation folders generated with FreeSurfer
9 % |- derivatives/meg_derivatives/sub-XX/ses-meg/meg/*.fif : MEG+EEG recordings (processed with MaxFilter's SSS)
10 % |- derivatives/meg_derivatives/sub-emptyroom/ses-meg/meg/*.fif : Empty room measurements
11 % - reports_dir: If defined, exports all the reports as HTML to this folder
12
13 % @=============================================================================
14 % This function is part of the Brainstorm software:
15 % https://neuroimage.usc.edu/brainstorm
16 %
17 % Copyright (c) University of Southern California & McGill University
18 % This software is distributed under the terms of the GNU General Public License
19 % as published by the Free Software Foundation. Further details on the GPLv3
20 % license can be found at http://www.gnu.org/copyleft/gpl.html.
21 %
22 % FOR RESEARCH PURPOSES ONLY. THE SOFTWARE IS PROVIDED "AS IS," AND THE
23 % UNIVERSITY OF SOUTHERN CALIFORNIA AND ITS COLLABORATORS DO NOT MAKE ANY
24 % WARRANTY, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO WARRANTIES OF
25 % MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE, NOR DO THEY ASSUME ANY
26 % LIABILITY OR RESPONSIBILITY FOR THE USE OF THIS SOFTWARE.
27 %
28 % For more information type "brainstorm license" at command prompt.
29 % =============================================================================@
30 %
31 % Author: Francois Tadel, Elizabeth Bock, 2016-2018
32
33
34 %% ===== SCRIPT VARIABLES =====
35 % Full list of subjects to process
36 SubjectNames = {'sub-01', 'sub-02', 'sub-03', 'sub-04', 'sub-05', 'sub-06', 'sub-07', 'sub-08', ...
37 'sub-09', 'sub-10', 'sub-11', 'sub-12', 'sub-13', 'sub-14', 'sub-15', 'sub-16'};
38 % Empty-room dates for each subject (so that we can match automatically recordings with empty-room)
39 EmptyRoomSubj = 'sub-emptyroom';
40 AcquisitionDates = {'09-Apr-2009', '06-May-2009', '11-May-2009', '18-May-2009', '15-May-2009', '15-May-2009', '15-May-2009', '15-May-2009', ...
41 '15-May-2009', '15-May-2009', '01-Jun-2009', '01-Jun-2009', '01-Jun-2009', '26-Nov-2009', '08-Dec-2009', '08-Dec-2009'};
42 % Bad channels {iSubj} = {Run01, Run02, Run03, Run04, Run05, Run06}
43 BadChannels{1} = {'EEG016', 'EEG070', 'EEG050',{'EEG008','EEG050'}, [], []};
44 BadChannels{2} = {{'EEG027', 'EEG030', 'EEG038'}, 'EEG010', 'EEG010', 'EEG010', 'EEG010', 'EEG010'};
45 BadChannels{3} = {{'EEG008','EEG017'}, {'EEG008','EEG017'}, {'EEG008','EEG017'}, {'EEG008','EEG017'}, {'EEG008','EEG017','EEG001'}, {'EEG008','EEG017','EEG020'}};
46 BadChannels{4} = {{'EEG038'}, {'EEG038','EEG001','EEG016'}, {'EEG038','EEG001','EEG016'}, {'EEG038','EEG001'}, {'EEG038','EEG001','EEG016'}, {'EEG038','EEG001','EEG016'}};
47 BadChannels{5} = {'EEG001', 'EEG001', [], [], [], []};
48 BadChannels{6} = {'EEG068', [], 'EEG004', [], [], []};
49 BadChannels{7} = {[], [], {'EEG004','EEG008'}, {'EEG004','EEG008'},{'EEG004','EEG008','EEG043','EEG045','EEG047'}, {'EEG004','EEG008'}};
50 BadChannels{8} = {[], [], [], [], [], []};
51 BadChannels{9} = {[], 'EEG004', 'EEG004', [], 'EEG004', 'EEG004'};
52 BadChannels{10} = {[], [], [], [], [], []};
53 BadChannels{11} = {{'EEG010','EEG050'}, 'EEG050', 'EEG050', 'EEG050', 'EEG050', 'EEG050'};
54 BadChannels{12} = {{'EEG024','EEG057'}, {'EEG024','EEG057'}, {'EEG024','EEG057'}, {'EEG024','EEG057','EEG070'}, {'EEG024','EEG057'}, {'EEG024','EEG057','EEG070'}};
55 BadChannels{13} = {'EEG009', 'EEG009', {'EEG009','EEG057','EEG69'}, 'EEG009', {'EEG009','EEG044'}, {'EEG009','EEG044'}};
56 BadChannels{14} = {'EEG029', 'EEG029', 'EEG029', {'EEG004','EEG008','EEG016','EEG029'}, {'EEG004','EEG008','EEG016','EEG029'}, {'EEG004','EEG008','EEG016','EEG029'}};
57 BadChannels{15} = {'EEG038', 'EEG038', 'EEG038', 'EEG038', {'EEG054','EEG038'}, 'EEG038'};
58 BadChannels{16} = {'EEG008', 'EEG008', 'EEG008', 'EEG008', 'EEG008', 'EEG008'};
59 % SSP components to remove {iSubj} = {sRun01, sRun02, sRun03, sRun03, sRun04, sRun05, sRun06}, sRun0X={ECG_GRAD,ECG_MAG}
60 SspSelect{1} = {{1,1}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
61 SspSelect{2} = {{1,1}, {1,1}, {1,1}, {1,1}, {3,1}, {1,1}};
62 SspSelect{3} = {{[],1}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
63 SspSelect{4} = {{[],1}, {[],1}, {[],1}, {[],1}, {[],1}, {[],1}};
64 SspSelect{5} = {{2,1}, {1,1}, {1,1}, {[],1}, {1,1}, {1,1}};
65 SspSelect{6} = {{2,1}, {2,1}, {1,1}, {2,1}, {1,1}, {2,1}};
66 SspSelect{7} = {{1,1}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
67 SspSelect{8} = {{1,1}, {1,1}, {[],1}, {2,1}, {1,1}, {2,1}};
68 SspSelect{9} = {{1,1}, {1,1}, {[],1}, {[],1}, {1,1}, {1,1}};
69 SspSelect{10} = {{1,1}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
70 SspSelect{11} = {{[],1}, {[],1}, {[],1}, {[],1}, {[],[]}, {[],[]}};
71 SspSelect{12} = {{[1,2],[1,2]}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
72 SspSelect{13} = {{[],[]}, {[],[]}, {[],[]}, {[],[]}, {[],[]}, {[],[]}};
73 SspSelect{14} = {{1,1}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
74 SspSelect{15} = {{1,1}, {1,1}, {1,1}, {1,1}, {1,1}, {1,1}};
75 SspSelect{16} = {{1,1}, {1,1}, {1,1}, {2,1}, {1,1}, {1,1}};
76
77
78 %% ===== CREATE PROTOCOL =====
79 % Start brainstorm without the GUI
80 if ~brainstorm('status')
81 brainstorm nogui
82 end
83 % Output folder for reports
84 if (nargin < 2) || isempty(reports_dir) || ~isdir(reports_dir)
85 reports_dir = [];
86 end
87 % You have to specify the folder in which the tutorial dataset is unzipped
88 if (nargin < 1) || isempty(bids_dir) || ~file_exist(bids_dir) || ~file_exist(bst_fullfile(bids_dir, 'derivatives')) || ~file_exist(bst_fullfile(bids_dir, 'dataset_description.json'))
89 error('The first argument must be the full path to the tutorial folder.');
90 end
91 % The protocol name has to be a valid folder name (no spaces, no weird characters...)
92 ProtocolName = 'TutorialVisual';
93 % Delete existing protocol
94 gui_brainstorm('DeleteProtocol', ProtocolName);
95 % Create new protocol
96 gui_brainstorm('CreateProtocol', ProtocolName, 0, 0);
97 % Set visualization filters: 40Hz low-pass, no high-pass
98 panel_filter('SetFilters', 1, 40, 0, [], 0, [], 0, 0);
99 % Set colormap: local color scale
100 bst_colormaps('SetMaxMode', 'meg', 'local');
101 bst_colormaps('SetMaxMode', 'eeg', 'local');
102
103
104 %% ===== PRE-PROCESS AND IMPORT =====
105 for iSubj = 1:16
106 % Start a new report (one report per subject)
107 bst_report('Start');
108 disp(sprintf('\n===== IMPORT: SUBJECT #%d =====\n', iSubj));
109
110 % If subject already exists: delete it
111 [sSubject, iSubject] = bst_get('Subject', SubjectNames{iSubj});
112 if ~isempty(sSubject)
113 db_delete_subjects(iSubject);
114 end
115
116 % ===== FILES TO IMPORT =====
117 % Build the path of the files to import
118 AnatDir = fullfile(bids_dir, 'derivatives', 'freesurfer', SubjectNames{iSubj}, 'ses-mri', 'anat');
119 DataDir = fullfile(bids_dir, 'derivatives', 'meg_derivatives', SubjectNames{iSubj}, 'ses-meg', 'meg');
120 % Check if the folder contains the required files
121 if ~file_exist(AnatDir)
122 error(['The folder "' AnatDir '" does not exist.']);
123 end
124 if ~file_exist(DataDir)
125 error(['The folder "' DataDir '" does not exist.']);
126 end
127
128 % ===== ANATOMY =====
129 % Process: Import anatomy folder
130 bst_process('CallProcess', 'process_import_anatomy', [], [], ...
131 'subjectname', SubjectNames{iSubj}, ...
132 'mrifile', {AnatDir, 'FreeSurfer'}, ...
133 'nvertices', 15000);
134
135 % ===== PROCESS EACH RUN =====
136 for iRun = 1:6
137 % Files to import
138 FifFile = bst_fullfile(DataDir, sprintf('%s_ses-meg_task-facerecognition_run-%02d_proc-sss_meg.fif', SubjectNames{iSubj}, iRun));
139
140 % ===== LINK CONTINUOUS FILE =====
141 % Process: Create link to raw file
142 sFileRaw = bst_process('CallProcess', 'process_import_data_raw', [], [], ...
143 'subjectname', SubjectNames{iSubj}, ...
144 'datafile', {FifFile, 'FIF'}, ...
145 'channelreplace', 1, ...
146 'channelalign', 0);
147 % Set acquisition date
148 panel_record('SetAcquisitionDate', sFileRaw.iStudy, AcquisitionDates{iSubj});
149
150 % ===== PREPARE CHANNEL FILE =====
151 % Process: Set channels type
152 bst_process('CallProcess', 'process_channel_settype', sFileRaw, [], ...
153 'sensortypes', 'EEG061, EEG064', ...
154 'newtype', 'NOSIG');
155 bst_process('CallProcess', 'process_channel_settype', sFileRaw, [], ...
156 'sensortypes', 'EEG062', ...
157 'newtype', 'EOG');
158 bst_process('CallProcess', 'process_channel_settype', sFileRaw, [], ...
159 'sensortypes', 'EEG063', ...
160 'newtype', 'ECG');
161
162 % Process: Remove head points
163 sFileRaw = bst_process('CallProcess', 'process_headpoints_remove', sFileRaw, [], ...
164 'zlimit', 0);
165 % Process: Refine registration
166 sFileRaw = bst_process('CallProcess', 'process_headpoints_refine', sFileRaw, []);
167 % Process: Project electrodes on scalp
168 sFileRaw = bst_process('CallProcess', 'process_channel_project', sFileRaw, []);
169
170 % Process: Snapshot: Sensors/MRI registration
171 bst_process('CallProcess', 'process_snapshot', sFileRaw, [], ...
172 'target', 1, ... % Sensors/MRI registration
173 'modality', 1, ... % MEG (All)
174 'orient', 1, ... % left
175 'Comment', sprintf('MEG/MRI Registration: Subject #%d, Run #%d', iSubj, iRun));
176 bst_process('CallProcess', 'process_snapshot', sFileRaw, [], ...
177 'target', 1, ... % Sensors/MRI registration
178 'modality', 4, ... % EEG
179 'orient', 1, ... % left
180 'Comment', sprintf('EEG/MRI Registration: Subject #%d, Run #%d', iSubj, iRun));
181
182 % ===== IMPORT TRIGGERS =====
183 % Process: Read from channel
184 bst_process('CallProcess', 'process_evt_read', sFileRaw, [], ...
185 'stimchan', 'STI101', ...
186 'trackmode', 2, ... % Bit: detect the changes for each bit independently
187 'zero', 0);
188 % Process: Group by name
189 bst_process('CallProcess', 'process_evt_groupname', sFileRaw, [], ...
190 'combine', 'Unfamiliar=3,4', ...
191 'dt', 0, ...
192 'delete', 1);
193 % Process: Rename event
194 bst_process('CallProcess', 'process_evt_rename', sFileRaw, [], ...
195 'src', '3', ...
196 'dest', 'Famous');
197 % Process: Rename event
198 bst_process('CallProcess', 'process_evt_rename', sFileRaw, [], ...
199 'src', '5', ...
200 'dest', 'Scrambled');
201 % Process: Add time offset
202 bst_process('CallProcess', 'process_evt_timeoffset', sFileRaw, [], ...
203 'info', [], ...
204 'eventname', 'Famous, Unfamiliar, Scrambled', ...
205 'offset', 0.0345);
206 % Process: Delete events
207 bst_process('CallProcess', 'process_evt_delete', sFileRaw, [], ...
208 'eventname', '1,2,6,7,8,9,10,11,12,13,14,15,16');
209 % Process: Detect cHPI activity (Elekta):STI201
210 bst_process('CallProcess', 'process_evt_detect_chpi', sFileRaw, [], ...
211 'eventname', 'chpi_bad', ...
212 'channelname', 'STI201', ...
213 'method', 'off'); % Mark as bad when the HPI coils are OFF
214
215 % ===== FREQUENCY FILTERS =====
216 % Process: Notch filter: 50Hz 100Hz 150Hz 200Hz
217 sFileClean = bst_process('CallProcess', 'process_notch', sFileRaw, [], ...
218 'freqlist', [50, 100, 150, 200], ...
219 'sensortypes', 'MEG, EEG', ...
220 'read_all', 0);
221 % Process: Power spectrum density (Welch)
222 sFilesPsd = bst_process('CallProcess', 'process_psd', [sFileRaw, sFileClean], [], ...
223 'timewindow', [], ...
224 'win_length', 4, ...
225 'win_overlap', 50, ...
226 'sensortypes', 'MEG, EEG', ...
227 'edit', struct(...
228 'Comment', 'Power', ...
229 'TimeBands', [], ...
230 'Freqs', [], ...
231 'ClusterFuncTime', 'none', ...
232 'Measure', 'power', ...
233 'Output', 'all', ...
234 'SaveKernel', 0));
235 % Process: Snapshot: Frequency spectrum
236 bst_process('CallProcess', 'process_snapshot', sFilesPsd, [], ...
237 'target', 10, ... % Frequency spectrum
238 'Comment', sprintf('Power spctrum: Subject #%d, Run #%d', iSubj, iRun));
239
240 % ===== BAD CHANNELS =====
241 if ~isempty(BadChannels{iSubj}{iRun})
242 % Process: Set bad channels
243 bst_process('CallProcess', 'process_channel_setbad', sFileClean, [], ...
244 'sensortypes', BadChannels{iSubj}{iRun});
245 end
246
247 % ===== EEG REFERENCE =====
248 % Process: Re-reference EEG
249 bst_process('CallProcess', 'process_eegref', sFileClean, [], ...
250 'eegref', 'AVERAGE', ...
251 'sensortypes', 'EEG');
252
253 % ===== DETECT ARTIFACTS ======
254 % Process: Detect heartbeats
255 bst_process('CallProcess', 'process_evt_detect_ecg', sFileClean, [], ...
256 'channelname', 'EEG063', ...
257 'timewindow', [], ...
258 'eventname', 'cardiac');
259 % Different amplitude thresholds for different subjects
260 if strcmpi(SubjectNames{iSubj}, 'sub-05')
261 thresholdMAX = 50;
262 else
263 thresholdMAX = 100;
264 end
265 % Process: Detect: blink_BAD - Detects all events where the amplitude exceeds 100uV
266 bst_process('CallProcess', 'process_evt_detect_threshold', sFileClean, [], ...
267 'eventname', 'blink_BAD', ...
268 'channelname', 'EEG062', ...
269 'timewindow', [], ...
270 'thresholdMAX', thresholdMAX, ...
271 'units', 3, ... % uV (10^-6)
272 'bandpass', [0.3, 20], ...
273 'isAbsolute', 1, ...
274 'isDCremove', 0);
275
276 % ===== SSP COMPUTATION =====
277 % Process: SSP ECG: cardiac
278 bst_process('CallProcess', 'process_ssp_ecg', sFileClean, [], ...
279 'eventname', 'cardiac', ...
280 'sensortypes', 'MEG GRAD', ...
281 'usessp', 1, ...
282 'select', SspSelect{iSubj}{iRun}{1});
283 bst_process('CallProcess', 'process_ssp_ecg', sFileClean, [], ...
284 'eventname', 'cardiac', ...
285 'sensortypes', 'MEG MAG', ...
286 'usessp', 1, ...
287 'select', SspSelect{iSubj}{iRun}{2});
288 % Process: Snapshot: SSP projectors
289 bst_process('CallProcess', 'process_snapshot', sFileClean, [], ...
290 'target', 2, ...
291 'Comment', sprintf('Subject #%d, Run #%d', iSubj, iRun)); % SSP projectors
292
293 % ===== IMPORT BAD EVENTS =====
294 % Get bad segments: this is typically done manually, not from a script
295 BadSegments = GetBadSegments(iSubj, iRun);
296 % Process: Import from file
297 bst_process('CallProcess', 'process_evt_import', sFileClean, [], ...
298 'evtfile', {BadSegments, 'ARRAY-TIMES'}, ...
299 'evtname', 'BAD');
300
301 % ===== IMPORT TRIALS =====
302 % Process: Import MEG/EEG: Events
303 sFilesEpochs = bst_process('CallProcess', 'process_import_data_event', sFileClean, [], ...
304 'subjectname', SubjectNames{iSubj}, ...
305 'condition', '', ...
306 'eventname', 'Famous, Scrambled, Unfamiliar', ...
307 'timewindow', [], ...
308 'epochtime', [-0.5, 1.2], ...
309 'createcond', 0, ...
310 'ignoreshort', 1, ...
311 'usectfcomp', 1, ...
312 'usessp', 1, ...
313 'freq', [], ...
314 'baseline', [-0.5, -0.0009]);
315
316 % ===== AVERAGE: RUN =====
317 % Process: Average: By trial group (folder average)
318 sFilesAvg = bst_process('CallProcess', 'process_average', sFilesEpochs, [], ...
319 'avgtype', 5, ... % By trial group (folder average)
320 'avg_func', 1, ... % Arithmetic average: mean(x)
321 'weighted', 0, ...
322 'keepevents', 0);
323 % Process: Snapshot: Recordings time series
324 bst_process('CallProcess', 'process_snapshot', sFilesAvg, [], ...
325 'target', 5, ... % Recordings time series
326 'modality', 4, ... % EEG
327 'time', 0.11, ...
328 'Comment', sprintf('Subject #%d, Run #%d', iSubj, iRun));
329 % Process: Snapshot: Recordings topography
330 bst_process('CallProcess', 'process_snapshot', sFilesAvg, [], ...
331 'target', 6, ... % Recordings topography (one time)
332 'modality', 4, ... % EEG
333 'time', 0.11, ...
334 'Comment', sprintf('Subject #%d, Run #%d', iSubj, iRun));
335
336 % ===== COMPUTE NOISECOV: EEG =====
337 % Process: Compute covariance (noise or data)
338 bst_process('CallProcess', 'process_noisecov', sFilesEpochs, [], ...
339 'baseline', [-0.5, -0.0009], ...
340 'sensortypes', 'EEG', ...
341 'target', 1, ... % Noise covariance (covariance over baseline time window)
342 'dcoffset', 1, ... % Block by block, to avoid effects of slow shifts in data
343 'identity', 0, ...
344 'copycond', 0, ...
345 'copysubj', 0, ...
346 'replacefile', 1); % Replace
347 end
348
349 % Save report
350 ReportFile = bst_report('Save', []);
351 if ~isempty(reports_dir) && ~isempty(ReportFile)
352 bst_report('Export', ReportFile, bst_fullfile(reports_dir, ['report_' ProtocolName '_' SubjectNames{iSubj} '.html']));
353 end
354 end
355
356
357 %% ===== EMPTY ROOM RECORDINGS =====
358 disp(sprintf('\n===== IMPORT: EMPTY-ROOM =====\n'));
359 % Loop on all the noise sessions
360 NoiseFiles = {};
361 for ses = {'20090409', '20090506', '20090511', '20090515', '20090518', '20090601', '20091126', '20091208'}
362 NoiseFiles{end+1} = fullfile(bids_dir, 'derivatives', 'meg_derivatives', EmptyRoomSubj, ['ses-' ses{1}], 'meg', ['sub-emptyroom_ses-' ses{1} '_task-noise_proc-sss_meg.fif']);
363 end
364 % Process: Create link to raw file
365 sFilesNoise = bst_process('CallProcess', 'process_import_data_raw', [], [], ...
366 'subjectname', EmptyRoomSubj, ...
367 'datafile', {NoiseFiles, 'FIF'}, ...
368 'channelreplace', 1, ...
369 'channelalign', 0);
370 % Process: Notch filter: 50Hz 100Hz 150Hz 200Hz
371 sFileNoiseClean = bst_process('CallProcess', 'process_notch', sFilesNoise, [], ...
372 'freqlist', [50, 100, 150, 200], ...
373 'sensortypes', 'MEG, EEG', ...
374 'read_all', 0);
375 % Process: Compute noise covariance
376 bst_process('CallProcess', 'process_noisecov', sFileNoiseClean, [], ...
377 'baseline', [], ...
378 'sensortypes', 'MEG', ...
379 'target', 1, ... % Noise covariance (covariance over baseline time window)
380 'dcoffset', 1, ... % Block by block, to avoid effects of slow shifts in data
381 'identity', 0, ...
382 'copycond', 1, ...
383 'copysubj', 1, ...
384 'copymatch', 1, ...
385 'replacefile', 2); % Merge
386
387
388 %% ===== SOURCE ESTIMATION =====
389 % Start a new report (one report for the source estimation of all the subjects)
390 bst_report('Start');
391 % Loop on the subjects: This loop is separated from the previous one, because we should
392 % compute the BEM surfaces after importing all the runs, so that the registration is done
393 % using the high resolution head surface, instead of the smooth scalp BEM layer.
394 for iSubj = 1:length(SubjectNames)
395 disp(sprintf('\n===== SOURCES: SUBJECT #%d =====\n', iSubj));
396
397 % ===== BEM SURFACES =====
398 % Process: Generate BEM surfaces
399 bst_process('CallProcess', 'process_generate_bem', [], [], ...
400 'subjectname', SubjectNames{iSubj}, ...
401 'nscalp', 1082, ...
402 'nouter', 642, ...
403 'ninner', 642, ...
404 'thickness', 4, ...
405 'method', 'brainstorm');
406
407 % ===== SELECT ALL AVERAGES =====
408 % Process: Select data files in: */*
409 sFilesAvg = bst_process('CallProcess', 'process_select_files_data', [], [], ...
410 'subjectname', SubjectNames{iSubj});
411 % Process: Select file comments with tag: Avg
412 sFilesAvg = bst_process('CallProcess', 'process_select_tag', sFilesAvg, [], ...
413 'tag', 'Avg'); % Select only the files with the tag
414
415 % ===== COMPUTE HEAD MODELS =====
416 % Process: Compute head model (only for the first run of the subject)
417 bst_process('CallProcess', 'process_headmodel', sFilesAvg(1), [], ...
418 'sourcespace', 1, ... % Cortex surface
419 'meg', 3, ... % Overlapping spheres
420 'eeg', 3, ... % OpenMEEG BEM
421 'ecog', 1, ... %
422 'seeg', 1, ... %
423 'openmeeg', struct(...
424 'BemSelect', [1, 1, 1], ...
425 'BemCond', [1, 0.0125, 1], ...
426 'BemNames', {{'Scalp', 'Skull', 'Brain'}}, ...
427 'BemFiles', {{}}, ...
428 'isAdjoint', 0, ...
429 'isAdaptative', 1, ...
430 'isSplit', 0, ...
431 'SplitLength', 4000));
432 % Get all the runs for this subject (ie the list of the study indices)
433 iStudyOther = setdiff(unique([sFilesAvg.iStudy]), sFilesAvg(1).iStudy);
434 % Copy the forward model file to the other runs
435 sHeadmodel = bst_get('HeadModelForStudy', sFilesAvg(1).iStudy);
436 for iStudy = iStudyOther
437 db_add(iStudy, sHeadmodel.FileName);
438 end
439
440 % ===== COMPUTE SOURCES: MEG =====
441 % Process: Compute sources [2018]
442 sAvgSrcMeg = bst_process('CallProcess', 'process_inverse_2018', sFilesAvg, [], ...
443 'output', 1, ... % Kernel only: shared
444 'inverse', struct(...
445 'Comment', 'MN: MEG ALL', ...
446 'InverseMethod', 'minnorm', ...
447 'InverseMeasure', 'amplitude', ...
448 'SourceOrient', {{'fixed'}}, ...
449 'Loose', 0.2, ...
450 'UseDepth', 1, ...
451 'WeightExp', 0.5, ...
452 'WeightLimit', 10, ...
453 'NoiseMethod', 'reg', ...
454 'NoiseReg', 0.1, ...
455 'SnrMethod', 'fixed', ...
456 'SnrRms', 1e-06, ...
457 'SnrFixed', 3, ...
458 'ComputeKernel', 1, ...
459 'DataTypes', {{'MEG GRAD', 'MEG MAG'}}));
460 % Process: Snapshot: Sources (one time) - Loop only to get a correct comment for the report
461 for i = 1:length(sAvgSrcMeg)
462 bst_process('CallProcess', 'process_snapshot', sAvgSrcMeg(i), [], ...
463 'target', 8, ... % Sources (one time)
464 'orient', 4, ... % bottom
465 'time', 0.11, ...
466 'threshold', 20, ...
467 'Comment', ['MEG sources: ' sFilesAvg(i).FileName]);
468 end
469
470 % ===== COMPUTE SOURCES: EEG =====
471 % Process: Compute sources [2018]
472 sAvgSrcEeg = bst_process('CallProcess', 'process_inverse_2018', sFilesAvg, [], ...
473 'output', 1, ... % Kernel only: shared
474 'inverse', struct(...
475 'Comment', 'MN: EEG', ...
476 'InverseMethod', 'minnorm', ...
477 'InverseMeasure', 'amplitude', ...
478 'SourceOrient', {{'fixed'}}, ...
479 'Loose', 0.2, ...
480 'UseDepth', 1, ...
481 'WeightExp', 0.5, ...
482 'WeightLimit', 10, ...
483 'NoiseMethod', 'reg', ...
484 'NoiseReg', 0.1, ...
485 'SnrMethod', 'fixed', ...
486 'SnrRms', 1e-06, ...
487 'SnrFixed', 3, ...
488 'ComputeKernel', 1, ...
489 'DataTypes', {{'EEG'}}));
490 % Process: Snapshot: Sources (one time) - Loop only to get a correct comment for the report
491 for i = 1:length(sAvgSrcEeg)
492 bst_process('CallProcess', 'process_snapshot', sAvgSrcEeg(i), [], ...
493 'target', 8, ... % Sources (one time)
494 'orient', 4, ... % bottom
495 'time', 0.11, ...
496 'threshold', 10, ...
497 'Comment', ['EEG sources: ' sFilesAvg(i).FileName]);
498 end
499 end
500 % Save report
501 ReportFile = bst_report('Save', []);
502 if ~isempty(reports_dir) && ~isempty(ReportFile)
503 bst_report('Export', ReportFile, bst_fullfile(reports_dir, ['report_' ProtocolName '_sources.html']));
504 end
505
506
507 %% ===== TIME-FREQUENCY =====
508 % Start a new report (one report for the time-frequency of all the subjects)
509 bst_report('Start');
510 % List of conditions to process separately
511 AllConditions = {'Famous', 'Scrambled', 'Unfamiliar'};
512 % Channels to display in the screen capture, by order of preference (if the first channel is bad, use the following)
513 SelChannel = {'EEG070','EEG060','EEG065','EEG050','EEG003'};
514 % Compute one separate time-frequency average for each subject/run/condition
515 for iSubj = 1:length(SubjectNames)
516 disp(sprintf('\n===== TIME-FREQUENCY: SUBJECT #%d =====\n', iSubj));
517 for iRun = 1:6
518 % Process: Select data files in: Subject/Run
519 sTrialsAll = bst_process('CallProcess', 'process_select_files_data', [], [], ...
520 'subjectname', SubjectNames{iSubj}, ...
521 'condition', sprintf('sub-%02d_ses-meg_task-facerecognition_run-%02d_proc-sss_meg_notch', iSubj, iRun));
522 % Loop on the conditions
523 for iCond = 1:length(AllConditions)
524 % Comment describing this average
525 strComment = [SubjectNames{iSubj}, ' / ', sprintf('run_%02d', iRun), ' / ', AllConditions{iCond}];
526 disp(['BST> ' strComment]);
527 % Find the first good channel in the display list
528 if isempty(BadChannels{iSubj}{iRun})
529 iSel = 1;
530 else
531 iSel = find(~ismember(SelChannel,BadChannels{iSubj}{iRun}), 1);
532 end
533 % Process: Select file comments with tag: Avg
534 sTrialsCond = bst_process('CallProcess', 'process_select_tag', sTrialsAll, [], ...
535 'tag', [AllConditions{iCond}, '_trial'], ...
536 'search', 1, ... % Search the file names
537 'select', 1); % Select only the files with the tag
538 % Process: Time-frequency (Morlet wavelets), averaged across trials
539 sTimefreq = bst_process('CallProcess', 'process_timefreq', sTrialsCond, [], ...
540 'sensortypes', 'MEG MAG, EEG', ...
541 'edit', struct(...
542 'Comment', ['Avg: ' AllConditions{iCond} ', Power, 6-60Hz'], ...
543 'TimeBands', [], ...
544 'Freqs', [6, 6.8, 7.6, 8.6, 9.7, 11, 12.4, 14, 15.8, 17.9, 20.2, 22.8, 25.7, 29, 32.7, 37, 41.7, 47.1, 53.2, 60], ...
545 'MorletFc', 1, ...
546 'MorletFwhmTc', 3, ...
547 'ClusterFuncTime', 'none', ...
548 'Measure', 'power', ...
549 'Output', 'average', ...
550 'RemoveEvoked', 0, ...
551 'SaveKernel', 0), ...
552 'normalize', 'none'); % None: Save non-standardized time-frequency maps
553 % Process: Extract time: [-200ms,900ms]
554 sTimefreq = bst_process('CallProcess', 'process_extract_time', sTimefreq, [], ...
555 'timewindow', [-0.2, 0.9], ...
556 'overwrite', 1);
557 % Screen capture of one sensor
558 hFigTf = view_timefreq(sTimefreq.FileName, 'SingleSensor', SelChannel{iSel});
559 bst_report('Snapshot', hFigTf, strComment, 'Time-frequency', [200, 200, 400, 250]);
560 close(hFigTf);
561 end
562 end
563 end
564 % Save report
565 ReportFile = bst_report('Save', []);
566 if ~isempty(reports_dir) && ~isempty(ReportFile)
567 bst_report('Export', ReportFile, bst_fullfile(reports_dir, ['report_' ProtocolName '_timefreq.html']));
568 end
569 end
570
571
572
573
574 %% ===== SUPPORT FUNCTIONS =====
575 function BadSeg = GetBadSegments(iSubj, iRun)
576 BadSegments{1} = {...
577 [247.867 248.185; 598.999 598.999; 598.999 598.999; 611.999 611.999; 612.999 612.999; 613.999 613.999; 616.999 616.999; 617.999 617.999; 623.999 623.999; 715.209 715.467], ...
578 [84.791 85.166], ...
579 [79.183 80.167], ...
580 [64.309 65.185], ...
581 [90.958 91.167; 178.005 178.355; 293.282 295.919; 312.298 316.479; 353.835 357.716], ...
582 [60.292 66.802; 69.975 71.210; 105.233 107.586; 108.822 109.506; 376.225 376.325]};
583 BadSegments{2} = {...
584 [223.806 224.199; 279.772 279.895; 453.241 455.108; 692.423 692.593], ...
585 [65.298 66.194; 304.727 306.178; 399.165 400.732], ...
586 [203.141 205.085; 281.579 287.883; 420.395 421.128], ...
587 [387.118 388.229; 440.318 441.900; 554.825 558.744], ...
588 [71.000 80.999; 82.750 87.367; 149.528 149.667; 264.747 267.995; 368.415 371.973; 376.263 378.763; 398.334 401.551; 537.410 541.645], ...
589 [38.000 47.999; 47.825 50.046; 61.298 61.384; 249.653 253.379; 282.917 283.820; 286.135 287.616; 298.167 300.196; 328.254 329.511; 335.957 337.817; 478.277 480.707]};
590 BadSegments{3} = {...
591 [406.312 407.207; 727.055 728.714], ...
592 [84.894 85.156; 152.028 152.946; 297.835 298.915; 418.272 421.845; 554.084 554.794], ...
593 [73.758 74.159; 378.212 378.536; 406.065 407.099; 470.541 471.698; 488.900 491.168; 529.596 530.453], ...
594 [94.874 95.152; 317.385 321.374; 325.696 327.055; 439.220 439.829; 454.473 455.175; 486.196 486.829; 518.660 522.015; 524.400 525.249; 562.417 570.325], ...
595 [96.208 97.181; 98.942 99.096; 135.005 135.754; 143.990 144.599; 250.139 250.247; 300.459 300.559; 338.265 339.322; 545.913 546.067], ...
596 [91.415 92.156; 284.843 286.525; 297.886 298.404; 317.046 317.163; 332.698 332.791; 358.946 359.402; 428.405 428.775; 478.374 478.690; 549.866 550.128]};
597 BadSegments{4} = {...
598 [22.967 22.967; 50.036 50.098; 52.058 52.058; 156.653 156.653; 171.565 173.386; 239.544 242.105; 268.162 270.175; 268.992 268.992; 316.032 316.032; 338.283 339.000; 357.959 361.909; 370.871 370.871; 381.579 383.677; 437.731 437.731; 463.482 468.505; 476.135 479.838; 486.652 488.272; 504.860 508.999], ...
599 [309.493 311.707; 342.681 344.525; 354.019 357.321; 390.023 391.225; 393.926 395.855; 404.221 405.069; 432.522 435.932; 459.048 460.715; 471.763 478.529; 549.387 551.999; 591.087 594.143; 608.541 611.079; 624.847 626.615; 649.648 651.570], ...
600 [57.411 58.198; 88.346 88.955; 200.761 202.335; 227.016 227.688; 257.726 258.054; 356.798 359.005; 404.260 411.003], ...
601 [46.000 54.823; 61.000 70.332; 203.005 207.125; 275.875 278.121; 313.500 314.824; 337.973 338.636; 422.505 426.239], ...
602 [58.000 62.479; 78.250 85.166; 89.955 91.360; 116.322 117.888; 130.013 131.987; 149.509 150.489; 174.650 175.823; 182.030 183.334; 196.758 197.384; 204.458 204.697; 205.236 208.663; 311.028 316.383; 320.700 327.181; 332.437 335.354; 344.205 346.133; 374.208 374.865; 385.519 386.214; 441.942 444.241; 453.957 456.997; 486.039 487.004; 501.238 504.185; 512.962 514.675; 553.398 556.215], ...
603 [41.406 45.743; 58.681 59.144; 108.086 108.896; 140.633 143.750; 196.110 199.474; 210.778 210.971; 234.649 235.143; 258.081 259.632; 339.101 340.805; 390.277 390.609; 438.935 442.122; 528.221 534.031]};
604 BadSegments{5} = {...
605 [265.539 265.778; 266.334 266.495; 268.479 268.965; 367.428 367.636; 439.655 442.779; 453.497 453.853; 504.997 505.329; 519.513 519.683; 595.674 595.982; 602.000 602.463], ...
606 [121.113 121.499; 124.971 126.213; 253.735 254.075; 272.232 272.464; 272.895 273.104; 346.368 346.645; 368.812 369.052; 406.382 406.605; 452.920 453.113; 454.903 455.112; 507.655 507.840; 508.766 509.013; 584.853 585.030; 594.831 595.656; 597.261 602.249], ...
607 [37.251 37.497; 38.825 39.056; 40.615 41.849; 43.624 44.758; 53.333 53.641; 54.698 55.076; 57.668 59.196; 79.129 79.360; 81.475 81.714; 122.658 123.375; 284.787 285.296; 288.754 288.993; 345.790 346.022; 421.212 421.459; 481.428 482.207; 503.408 503.685; 504.272 504.449; 524.451 524.714; 526.913 531.499], ...
608 [87.322 88.178; 91.085 91.325; 95.121 95.491; 114.174 114.397; 129.874 130.113; 151.220 151.544; 281.689 281.959; 532.966 533.345], ...
609 [59.176 60.218; 74.854 75.317; 308.180 309.877; 380.705 381.059], ...
610 [182.382 183.245; 196.220 196.736; 276.018 276.327; 292.490 294.086; 370.755 370.847; 435.644 436.624; 467.535 468.460; 522.838 525.847]};
611 BadSegments{6} = {...
612 [141.690 142.424; 157.070 157.417; 355.138 356.025; 423.999 423.999; 424.999 424.999; 426.999 426.999; 427.999 427.999; 486.430 488.151; 493.999 493.999; 501.511 501.619; 501.549 501.549; 501.585 501.585; 502.999 502.999; 503.999 503.999; 540.999 540.999; 541.999 541.999; 555.999 555.999; 556.999 556.999; 561.999 561.999; 563.999 563.999; 564.999 564.999; 565.999 565.999; 567.999 567.999], ...
613 [64.898 65.161; 71.700 72.718; 226.185 226.740; 324.124 324.425; 329.062 329.301; 486.143 486.975], ...
614 [62.300 63.148; 266.254 266.639; 409.920 410.221], ...
615 [54.048 55.214; 330.893 331.255], ...
616 [185.616 186.495; 331.411 331.796; 386.843 387.028; 387.999 387.999; 389.999 389.999; 434.575 434.875; 519.802 519.995], ...
617 [44.720 45.167; 211.446 211.964; 368.955 369.172]};
618 BadSegments{7} = {...
619 [154.966 155.144; 551.639 551.855], ...
620 [88.774 89.167; 107.999 107.999; 109.999 109.999; 110.999 110.999; 112.999 112.999; 113.999 113.999; 114.999 114.999; 119.999 119.999; 121.999 121.999; 124.223 125.465; 137.011 138.871], ...
621 [81.627 82.136; 377.953 381.046], ...
622 [241.136 242.171; 543.849 544.196; 596.639 598.553; 600.227 601.075; 603.999 603.999; 605.999 605.999; 609.999 609.999; 611.305 612.809; 614.999 614.999; 615.803 616.937; 623.694 625.877; 653.999 653.999; 655.055 655.756; 663.999 663.999], ...
623 [68.852 69.481; 74.034 75.200; 78.426 78.600; 104.497 105.963; 253.951 254.034; 256.964 257.038; 257.915 258.048; 323.254 324.156; 365.880 368.131; 369.952 370.060; 371.728 372.999; 430.931 431.965; 535.521 542.999], ...
624 [70.271 71.205; 94.441 96.445; 98.613 99.126; 112.318 112.749; 131.686 132.265; 148.935 150.615; 161.120 161.600; 205.325 208.214; 215.035 215.863; 217.403 218.818; 286.178 287.171; 399.075 404.853]};
625 BadSegments{8} = {...
626 [238.505 238.546; 256.354 257.224; 316.116 316.167; 341.519 341.558; 356.493 356.566; 380.095 380.170; 391.906 392.048; 448.457 448.562; 469.931 470.028; 530.488 530.575; 555.180 558.136; 562.152 562.245; 588.403 588.502; 625.205 625.662; 638.019 638.438; 649.982 650.008; 650.691 651.357; 651.925 652.061; 665.445 665.472; 695.923 696.015; 706.528 706.720; 729.706 732.534], ...
627 [99.546 106.114; 113.245 114.199; 150.885 154.989; 277.486 278.075; 333.645 335.574; 339.358 340.646; 371.860 375.394; 497.459 499.156], ...
628 [49.008 50.205; 231.446 233.406; 329.590 329.659; 355.101 356.019; 360.733 360.973; 372.955 374.891; 389.283 392.068; 453.610 455.431; 464.632 465.265; 489.209 489.996; 514.777 515.295], ...
629 [178.368 179.232; 293.865 294.521; 418.252 418.314; 450.124 450.209; 480.236 480.445; 492.725 493.975; 495.170 497.624; 500.382 500.459; 504.247 504.402; 628.017 628.079; 628.827 628.905; 630.209 630.332], ...
630 [100.616 101.172; 107.691 108.539; 188.814 188.875; 193.119 193.281; 207.964 208.033; 432.254 432.385; 489.834 489.911; 517.960 518.037; 518.955 519.040; 520.938 521.062; 521.945 522.037; 523.959 524.052; 525.942 526.057; 526.945 527.015; 528.958 529.059; 531.138 531.192; 531.979 532.048; 532.951 533.028; 533.962 534.024], ...
631 [135.997 137.232; 383.565 383.657; 418.763 418.955]};
632 BadSegments{9} = {...
633 [215.107 216.187; 262.388 262.388; 287.519 287.635; 289.895 290.135; 311.999 311.999; 350.161 351.179; 526.154 527.033; 564.999 564.999; 584.935 585.059; 587.999 587.999; 601.999 601.999; 603.999 603.999; 608.999 608.999; 612.999 612.999], ...
634 [47.667 47.775; 49.172 54.681; 67.195 70.805; 78.988 80.477; 138.701 138.948; 138.727 138.727; 138.728 138.728; 138.728 138.728; 138.734 138.734; 138.734 138.734; 138.881 138.881; 138.881 138.881; 138.907 138.907; 140.993 141.093; 141.037 141.037; 141.045 141.045; 155.795 155.864; 155.822 155.822; 155.849 155.849; 168.999 168.999; 206.999 206.999; 219.999 219.999; 225.999 225.999; 226.999 226.999; 228.999 228.999; 236.999 236.999; 242.463 242.463; 242.463 242.463; 242.487 242.487; 247.999 247.999; 251.999 251.999; 252.483 252.483; 253.315 254.163; 265.999 265.999; 267.999 267.999; 272.358 272.358; 272.410 272.410; 298.999 298.999; 300.999 300.999; 314.037 314.037; 314.047 314.047; 321.813 321.813; 321.813 321.813; 321.833 321.833; 329.117 329.117; 329.117 329.117; 329.156 329.156; 346.999 346.999; 347.999 347.999; 349.320 349.320; 349.329 349.329; 352.528 355.869; 364.040 364.040; 396.278 397.420; 404.865 404.865; 407.905 407.905; 407.905 407.905; 407.954 407.954; 418.454 418.454; 418.454 418.454; 418.486 418.486; 441.999 441.999; 444.999 444.999; 447.999 447.999; 453.550 453.650; 454.931 455.055; 457.999 457.999; 479.964 480.079; 481.999 481.999; 482.949 483.073; 488.948 489.064; 511.999 511.999; 520.999 520.999; 523.999 523.999; 526.954 527.069; 533.999 533.999; 537.999 537.999], ...
635 [82.889 83.198; 206.143 207.424; 403.873 404.096; 406.790 407.485; 413.936 414.075; 415.912 416.112; 536.621 537.532], ...
636 [182.999 182.999; 182.999 182.999; 183.999 183.999; 195.999 195.999; 208.999 208.999; 209.999 209.999; 278.601 278.955; 413.913 414.067; 415.250 417.792; 419.940 420.087; 420.999 420.999; 512.999 512.999; 514.363 516.254; 521.917 522.134; 522.999 522.999; 523.915 524.101; 533.999 533.999; 538.999 538.999; 539.892 540.062; 543.999 543.999; 548.874 549.089; 549.900 550.062; 552.755 554.075; 585.999 585.999; 587.957 588.073; 588.999 588.999; 594.999 594.999; 603.971 604.056; 604.928 605.097; 615.986 618.495; 623.999 623.999], ...
637 [53.215 54.203; 164.227 168.965; 201.433 202.775; 292.973 295.889; 303.961 304.817; 309.354 311.236; 313.123 313.639; 322.547 323.017; 331.141 334.852; 356.637 356.985; 367.855 368.079; 369.995 373.459; 377.849 378.142; 406.072 407.306; 436.164 436.665; 458.033 458.905; 516.889 518.656; 517.812 518.684], ...
638 [113.945 115.225; 198.570 198.863; 264.162 264.795; 383.705 385.641; 396.477 397.064; 399.706 406.457; 452.261 453.179; 486.338 487.102; 498.869 499.579; 507.968 508.925; 546.266 547.285; 558.825 560.128]};
639 BadSegments{10} = {...
640 [235.104 236.184; 324.272 325.028; 330.799 331.401; 541.062 541.826; 564.747 565.072], ...
641 [100.581 101.229; 285.644 285.644; 285.644 285.644; 297.979 298.033; 298.999 298.999; 300.949 301.026; 301.999 301.999; 303.999 303.999; 304.999 304.999; 306.999 306.999; 307.999 307.999; 310.999 310.999; 311.999 311.999; 313.971 314.025; 314.999 314.999; 316.995 317.035; 317.999 317.999; 319.999 319.999; 320.923 321.046; 326.971 327.048; 327.999 327.999; 329.999 329.999; 330.999 330.999; 332.974 333.028; 333.999 333.999; 335.971 336.025; 336.999 336.999; 338.973 339.035; 339.999 339.999; 341.951 342.044; 343.999 343.999; 344.964 345.033; 346.999 346.999; 347.999 347.999; 452.015 453.411; 458.735 459.776; 467.974 468.089], ...
642 [41.266 41.675; 53.905 55.240; 159.159 159.345; 220.255 220.394], ...
643 [66.751 67.190; 201.674 201.812; 294.119 295.168; 303.188 303.528; 316.992 317.037; 317.999 317.999; 317.999 317.999; 319.999 319.999; 320.980 321.042; 322.999 322.999; 325.999 325.999; 326.955 327.048; 328.999 328.999; 333.999 333.999; 334.999 334.999; 342.999 342.999; 343.999 343.999; 351.915 352.038; 352.999 352.999; 407.979 408.056; 409.999 409.999; 411.999 411.999; 415.405 416.154; 435.999 435.999; 436.962 437.046; 468.885 469.100; 469.999 469.999; 470.999 470.999; 516.210 516.357], ...
644 [105.369 106.172; 141.468 142.178; 199.008 199.818; 201.269 201.716; 352.851 353.083; 449.865 452.041; 459.508 459.802], ...
645 [229.948 230.033; 230.999 230.999; 230.999 230.999; 259.588 259.990; 343.970 345.798; 361.999 361.999; 362.970 363.046; 364.999 364.999; 366.999 366.999; 367.962 368.031; 372.521 374.179; 405.999 405.999; 409.999 409.999; 411.999 411.999; 412.999 412.999; 434.999 434.999; 435.970 436.031; 516.999 516.999; 519.999 519.999]};
646 BadSegments{11} = {...
647 [179.045 180.225; 323.999 323.999; 323.999 323.999; 324.999 324.999; 327.978 328.025; 328.966 329.028; 330.999 330.999; 365.999 365.999; 367.999 367.999; 370.999 370.999; 371.999 371.999; 373.999 373.999; 375.965 376.042; 377.999 377.999], ...
648 [52.107 53.156; 521.077 521.155], ...
649 [65.331 66.156], ...
650 [81.579 82.143; 108.565 108.565; 108.566 108.566; 112.176 112.176; 122.377 122.377; 122.565 122.565; 213.882 213.882; 215.305 215.305; 224.851 224.851; 224.919 224.919; 255.815 255.815; 257.893 257.893; 359.952 359.952; 361.630 361.754; 370.285 370.285; 376.853 376.853; 511.600 511.600; 513.631 513.631; 513.988 513.988; 518.215 518.215], ...
651 [63.205 64.154; 170.951 171.036; 176.841 176.841; 176.842 176.842; 177.282 177.282; 223.886 223.971; 259.963 259.963; 261.547 261.547; 341.185 341.302; 368.999 368.999; 370.999 370.999; 374.999 374.999; 382.971 383.033; 383.999 383.999; 386.999 386.999; 388.958 389.035; 390.999 390.999; 391.955 392.031; 394.955 395.040; 396.999 396.999; 398.999 398.999; 400.999 400.999; 402.999 402.999; 404.963 405.033; 406.999 406.999; 429.829 429.829; 430.346 430.346; 465.184 465.184; 470.290 470.290], ...
652 [50.195 51.145; 158.850 159.012]};
653 BadSegments{12} = {...
654 [193.435 194.175; 500.000 501.000; 08.988 509.868; 528.999 528.999; 528.999 528.999; 529.999 529.999; 531.999 531.999; 547.999 547.999], ...
655 [133.484 134.185; 134.911 135.065; 553.999 553.999; 553.999 553.999; 554.999 554.999; 557.999 557.999; 558.999 558.999; 564.999 564.999; 565.977 566.046; 568.999 568.999; 569.951 570.028; 571.999 571.999; 579.959 580.028; 580.977 581.055; 583.971 584.033; 583.999 583.999; 585.999 585.999; 586.999 586.999; 588.980 589.026; 590.999 590.999; 591.958 592.044; 595.999 595.999], ...
656 [46.028 47.216; 129.557 130.236; 200.999 200.999; 200.999 200.999; 201.975 202.075; 203.999 203.999; 204.967 205.053; 213.627 214.515; 218.958 219.044; 222.959 228.252; 309.999 309.999; 311.897 317.537; 311.999 311.999; 351.968 353.234; 399.999 399.999; 446.508 451.454; 487.999 487.999; 512.278 512.926], ...
657 [82.535 84.062; 102.247 102.355; 134.999 134.999; 134.999 134.999; 136.999 136.999; 138.973 139.042; 147.999 147.999; 148.999 148.999; 149.999 149.999; 193.999 193.999; 194.999 194.999; 224.984 226.095; 241.954 242.077; 243.948 244.087; 280.999 280.999; 281.999 281.999; 305.000 305.451; 310.945 311.068; 328.999 328.999; 352.999 352.999; 353.999 353.999; 397.025 397.411; 415.944 418.096; 415.999 415.999; 470.999 470.999; 477.971 478.041; 483.997 484.737; 492.958 493.089; 493.999 493.999; 494.999 494.999; 522.999 522.999; 523.999 523.999; 524.955 525.039], ...
658 [175.183 176.155; 246.112 246.991; 412.005 413.340; 483.915 484.061; 485.959 486.028; 497.360 498.880; 594.457 594.944; 596.641 598.006; 615.277 618.710], ...
659 [66.900 72.232; 75.510 78.688; 543.495 545.999]};
660 BadSegments{13} = {...
661 [307.246 308.179; 627.881 628.089; 629.941 630.049], ...
662 [171.506 172.301; 172.999 172.999; 430.999 430.999; 432.999 432.999; 434.999 434.999; 448.999 448.999; 479.999 479.999; 489.999 489.999; 490.999 490.999; 491.999 491.999], ...
663 [92.059 93.209], ...
664 [52.791 53.231; 54.999 54.999; 65.985 68.138; 91.240 91.386; 92.137 92.207; 105.346 105.493; 121.120 121.398; 146.546 146.955; 194.025 197.652; 242.985 243.571; 247.101 247.556; 269.889 270.083; 270.946 271.070; 274.408 275.866; 294.565 295.267; 319.370 319.879; 365.999 365.999; 375.913 376.044; 376.869 377.055; 377.999 377.999; 390.130 393.225; 403.965 404.035; 404.953 405.075; 419.265 419.565; 427.015 427.154; 427.879 428.118; 427.999 427.999; 428.913 429.067; 480.945 482.095; 484.285 484.571; 516.929 517.099; 517.963 518.079], ...
665 [115.921 116.060; 117.999 117.999; 120.999 120.999; 126.896 127.058; 130.955 131.039; 131.957 132.043; 132.961 133.015; 134.999 134.999; 135.950 136.043; 139.999 139.999; 143.999 143.999; 144.850 145.066; 145.999 145.999; 146.999 146.999; 160.929 161.045; 161.999 161.999; 189.086 189.310; 193.939 194.063; 195.975 196.045; 199.961 200.045; 201.952 202.059; 202.939 205.084; 210.999 210.999; 211.738 213.629; 217.954 218.061; 218.987 219.049; 219.975 220.028; 258.953 259.037; 259.952 260.045; 260.963 261.032; 261.958 262.043; 262.954 263.054; 262.999 262.999; 288.999 288.999; 295.477 295.655; 298.976 299.054; 299.979 300.018; 300.975 301.005; 336.952 337.044; 337.977 338.032; 338.988 339.027; 338.999 338.999; 339.991 340.045; 340.975 341.029; 341.978 342.016; 343.976 344.054; 361.961 362.045; 365.942 366.050; 370.999 370.999; 384.961 385.045; 409.977 410.047; 415.984 416.030; 426.944 427.059; 430.999 430.999; 436.999 436.999; 448.999 448.999; 449.500 452.131; 481.962 482.055; 482.973 483.027; 484.975 485.060; 503.999 503.999; 504.999 504.999; 507.967 508.028; 508.955 509.032; 510.999 510.999; 512.999 512.999; 518.865 520.292; 522.958 523.043; 523.999 523.999; 524.948 525.034; 525.959 526.044; 526.946 527.055; 527.942 528.027; 528.961 529.053; 529.945 530.022; 535.999 535.999; 536.957 537.058; 538.967 539.036; 539.963 540.040; 541.976 542.045; 543.999 543.999; 551.956 552.034; 552.990 553.028; 553.977 554.024; 555.968 556.022; 556.204 557.454; 557.963 558.032; 559.969 560.023; 563.973 564.035; 563.999 563.999], ...
666 [87.308 87.639; 107.039 108.189; 427.943 428.035; 437.965 438.027; 439.973 440.026; 491.275 492.571]};
667 BadSegments{14} = {...
668 [365.982 367.194; 368.999 368.999; 368.999 368.999; 369.999 369.999; 371.999 371.999; 373.999 373.999; 375.942 376.050; 376.999 376.999; 378.999 378.999; 380.999 380.999; 382.999 382.999; 383.999 383.999; 386.999 386.999; 387.999 387.999; 418.999 418.999; 444.999 444.999; 690.178 693.218], ...
669 [101.796 102.183; 218.999 218.999; 219.999 219.999; 221.999 221.999; 222.999 222.999; 227.999 227.999; 229.999 229.999; 230.999 230.999; 232.999 232.999; 233.999 233.999; 235.999 235.999; 237.999 237.999; 238.985 239.031; 240.999 240.999; 242.999 242.999; 243.999 243.999; 246.999 246.999; 247.999 247.999; 253.999 253.999; 258.999 258.999; 259.999 259.999; 260.227 261.123; 264.999 264.999; 265.999 265.999; 268.999 268.999; 269.999 269.999; 274.999 274.999; 276.999 276.999; 277.999 277.999; 280.999 280.999; 282.966 283.021; 284.999 284.999; 285.999 285.999; 360.999 360.999; 362.973 363.026; 364.999 364.999; 365.999 365.999; 367.999 367.999; 368.985 369.024; 370.999 370.999; 371.999 371.999; 378.999 378.999; 379.999 379.999; 381.999 381.999; 383.999 383.999; 385.999 385.999; 387.999 387.999; 388.999 388.999; 398.999 398.999; 415.999 415.999; 417.999 417.999; 418.999 418.999; 421.999 421.999; 422.999 422.999], ...
670 [89.285 90.187], ...
671 [85.991 87.179; 88.999 88.999; 89.999 89.999; 92.966 93.027; 93.999 93.999; 107.999 107.999; 110.999 110.999; 111.971 112.025; 118.999 118.999; 121.999 121.999; 122.999 122.999; 166.999 166.999; 168.957 169.035; 173.999 173.999; 174.999 174.999; 175.957 176.035; 198.999 198.999; 199.999 199.999; 202.999 202.999; 203.999 203.999; 205.999 205.999; 207.999 207.999; 208.999 208.999; 210.974 211.044; 211.999 211.999; 212.999 212.999; 232.984 233.023; 234.999 234.999; 235.999 235.999; 236.999 236.999; 239.958 240.044; 240.999 240.999], ...
672 [71.132 72.227], ...
673 [98.134 98.827; 110.497 110.814; 114.000 117.272; 279.999 279.999; 279.999 279.999; 280.999 280.999; 282.999 282.999; 283.999 283.999; 284.999 284.999; 286.999 286.999; 287.999 287.999; 288.999 288.999; 289.999 289.999; 291.966 292.044; 292.999 292.999; 346.999 346.999; 437.608 437.646; 518.988 519.104; 522.969 523.015; 534.999 534.999; 536.967 537.029; 562.725 563.426]};
674 BadSegments{15} = {...
675 [66.755 67.172; 257.988 258.042; 258.714 259.385; 260.958 261.044; 265.931 266.062; 267.968 268.022; 268.978 269.033; 270.992 271.031; 272.964 273.017; 273.968 274.030; 275.975 276.059; 276.970 277.046; 288.976 289.038; 289.984 290.015; 295.994 296.025; 296.982 297.013; 300.966 301.152; 364.730 364.800], ...
676 [46.531 46.555; 65.156 66.148; 87.535 87.642; 183.896 183.934; 294.342 294.375; 330.004 330.528; 333.785 333.815; 345.177 345.203; 399.544 399.573; 480.964 480.980], ...
677 [53.557 54.175; 148.851 149.136], ...
678 [43.681 44.214; 361.255 361.587; 409.462 411.021], ...
679 [90.868 91.130; 410.935 411.044], ...
680 [64.786 65.141; 132.385 132.786; 177.735 178.252; 278.747 278.902; 313.959 314.028; 314.970 315.031]};
681 BadSegments{16} = {...
682 [38.958 44.335; 305.280 312.826; 324.778 326.067; 393.183 398.367; 403.410 422.854], ...
683 [49.726 50.251; 58.675 65.465; 263.918 264.088; 321.795 322.195; 522.881 523.081], ...
684 [61.245 62.325; 65.628 65.905; 315.686 317.275; 335.184 336.449; 387.870 388.117; 389.892 390.062], ...
685 [56.020 57.255; 60.711 61.096; 64.353 64.924; 73.202 73.757; 76.134 76.844; 95.625 96.859; 171.436 172.625; 178.359 178.715; 212.828 213.415; 352.425 352.617; 367.755 369.097; 488.400 489.426], ...
686 [71.665 72.513; 76.347 77.181; 127.134 128.862], ...
687 [46.190 47.209; 53.461 55.752; 194.317 194.510; 213.101 213.240; 216.356 216.442; 225.962 226.286; 245.163 245.464; 252.041 253.422; 254.648 254.856; 292.966 294.340; 340.928 341.067; 346.298 346.452]};
688 BadSeg = BadSegments{iSubj}{iRun}';
689 end
690
Report viewer
Click on Run to start the script.
As this process is taking screen captures, do not use your computer for something else at the same time: if another window covers the Brainstorm figures, it will not capture the right images.
At the end, the report viewer is opened to show the status of all the processes, the information messages, the list of input and output files, and the screen captures. The report is saved in your home folder ($home/.brainstorm/reports). If you close this window, you can get it back with the menu File > Report viewer.