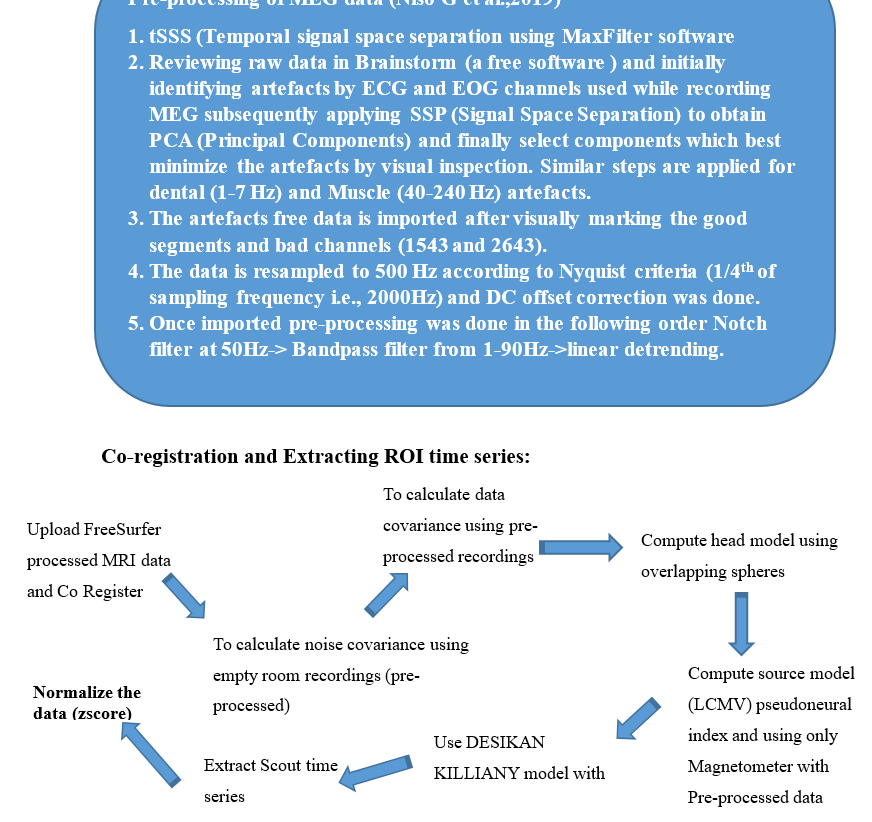
Can someone tell me, if this is the right pre-processing pipeline for continuous MEG data from Elekta Neuromag?. We were trying to calculate non linear measures after importing pre-processed data into MATLAB? We are getting some unexpected results.
Thanks in advance
Some comments:
- The frequency filters (band-pass and notch) should be applied on continuous files. We recommend running these before any other pre-processing step:
- See the order of the introduction tutorials: https://neuroimage.usc.edu/brainstorm/Tutorials
- Advanced tutorial on Elekta recordings: https://neuroimage.usc.edu/brainstorm/Tutorials/VisualSingle
- Summary of the recommended workflows: https://neuroimage.usc.edu/brainstorm/Tutorials/Workflows#Common_pre-processing_pipeline
- If you apply a band-pass filter 1-90Hz: any baseline offset or linear trend is already removed (they correspond to frequencies < 1Hz) => Do not apply any DC offset correction or detrending
- Empty-room: make sure the pre-processing procedure is the same as for the recordings of interest
Thank you very much.
Another small query, after we identify the bad segments, while analysing do the bad segments are taken into account for the analysis??
Because what I have seen is, when we import the bad segments are remain in the imported data.
We want 20 seconds of artefact free data. Have continuous file up to 5min. Do you suggest to split the data into epochs while importing?? Or we can just analyse the entire data at once. As we are not doing any task related or event related analysis. It’s just the resting state analysis.
Another small query, after we identify the bad segments, while analysing do the bad segments are taken into account for the analysis??
Bad segments in continuous files: it depends for what.
For PSD computations: the windows overlapping with a bad segment are ignore.
For other processes, like filters: the bad segments are ignored.
When epoching the data, epochs overlapping with bad segments are marked as bad, which are ignore from the analysis:
- https://neuroimage.usc.edu/brainstorm/Tutorials/Epoching#Import_in_database
- https://neuroimage.usc.edu/brainstorm/Tutorials/Averaging#Averaging
Bad segment in imported files are usually ignored (except by a few processes, like the PSD computation).
We want 20 seconds of artefact free data. Have continuous file up to 5min. Do you suggest to split the data into epochs while importing?? Or we can just analyse the entire data at once.
Split the data in a way that is convenient for you.
If an imported file is marked as bad and you need to be able to process it, mark it as good manually:
https://neuroimage.usc.edu/brainstorm/Tutorials/Epoching#Review_the_individual_trials