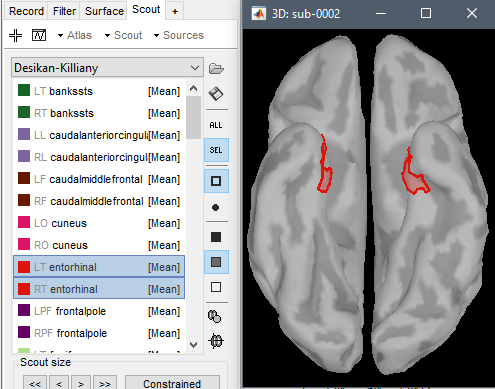
For example, the vertices that correspond to the left entorhinal all in one ROI?
Use the cortical (or volume) parcellations generated by FreeSurfer or CAT.
- https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Cortical_parcellations
- https://neuroimage.usc.edu/brainstorm/Tutorials/SegCAT12#Cortical_parcellations
Data exported as "SurfaceMat"
>> iAtlas = find(strcmpi({SurfaceMat.Atlas.Name}, 'Desikan-Killiany'))
iAtlas =
3
>> iScout = find(strcmpi({SurfaceMat.Atlas(iAtlas).Scouts.Label}, 'entorhinal L'))
iScout =
9
>> SurfaceMat.Atlas(iAtlas).Scouts(iScout)
ans =
struct with fields:
Vertices: [33×1 double]
Seed: 4929
Color: [0.8627 0.0784 0.0392]
Label: 'entorhinal L'
Function: 'Mean'
Region: 'LT'
Handles: []