If you are referring to the PCA option for computing scouts time series (ie. summarize the signals from multiple dipoles into one signal only), it computes the principal mode across all the signals in input:https://neuroimage.usc.edu/brainstorm/Tutorials/Scouts#Scout_function
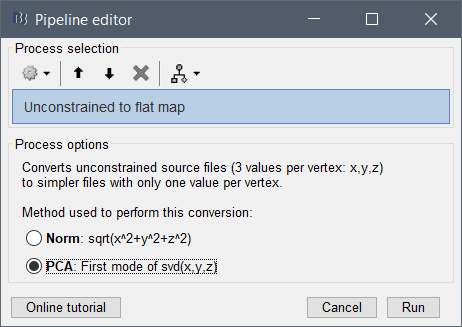
If you are referring to the PCA option of the process "Sources > Unconstrained to flat map", it groups at each spatial location the three times series (x,y,z) of unconstrained minimum norm source estimation into one signal:brainstorm3/toolbox/process/functions/process_source_flat.m at master · brainstorm-tools/brainstorm3 · GitHub
Both are computed with function bst_scout_value.m / PcaFirstMode():
[U, S] = eig((Covar + Covar')/2, 'vector'); % ensure exact symmetry for real results.
[S, iSort] = sort(S, 'descend');
PcaFirstComp = U(:, iSort(1));
explained = S(1) / sum(S);
else % use data
% This is a deprecated case, but still used for display. It's not taking into account
% baseline and data time windows like when we compute the covariance (outside this
% function). nComponents should now always be 1 here, but keep code compatible for
% non-standard calls and testing old behaviour.
explained = [0, 0];
PcaFirstComp = zeros(nRow, nComponents);
for i = 1:nComponents
% Keeping offset removal as before: whole trial.
[U, S] = svd(bsxfun(@minus, F(:,:,i), sum(F(:,:,i),2)./size(F,2)), 'econ'); % sum faster than mean
S = diag(S);
explained = explained + [S(1).^2, sum(S.^2)];
PcaFirstComp(:,i) = U(:, 1);
end
explained = explained(1) / explained(2);
end
% Remove ambiguity of arbitrary component sign for consistency across files/epochs and reproducibility.