|
Size: 4057
Comment:
|
Size: 15975
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Review continuous files = ''Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet'' |
= Tutorial 5: Review continuous recordings = ''Authors: Francois Tadel, Elizabeth Bock, John C Mosher, Sylvain Baillet'' |
| Line 6: | Line 6: |
| == Access the raw file == The basic tutorials you read before explain how to import recordings in the database: this operation creates a copy of all the data in Matlab .mat files in the Brainstorm database folders. You could process continuous recordings in the same way, but the .mat format has this limitation that the entire file has to be read even when you want to access just a portion of it. Long recordings usually cannot fit in memory and have to be split in small blocks of a few seconds, which makes it very difficult to review and process. |
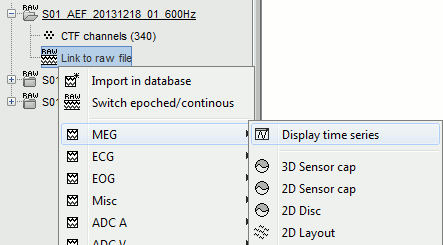
== Open the recordings == Let's look at the first file in the list: '''AEF#01'''.<<BR>>Right-click on the Link to raw file. Below the first to menus, you have the list of channel types: |
| Line 9: | Line 9: |
| Brainstorm offers the possibility to visualize continuous MEG/EEG recordings in any of the supported file formats without having to fully "import" them. A link to the native file is created in the database, which can be then manipulated almost like the "imported" recording blocks. Only the description of the file is saved in the database, and when displaying it the values are read directly from the native file. | * '''MEG''': 274 axial gradiometers * '''ECG''': 1 electrocadiogram, bipolar electrode across the chest * '''EOG''': 2 electrooculograms (vertical and horizontal) * '''Misc''': EEG electrodes Cz and Pz * '''ADC A''': Unused * '''ADC V''': Auditory signal sent to the subject * '''DAC''': Unused * '''FitErr''': Fitting error when trying to localize the three head localization coils (NAS, LPA, RPA) * '''HLU''': Head Localizing Unit, displacements in the three directions (x,y,z) for the three coils * '''MEG REF''': 26 reference sensors used for removing the environmental noise * '''Other''': Unused * '''Stim''': Stimulation channel, records the stim triggers generated by the Psychophysics toolbox and other input channels, such as button presses generated by the subject * '''SysClock''': System clock, unused |
| Line 11: | Line 23: |
| In addition, an interface allows to edit the time markers that are saved in the file. Those markers can then be used to import the recordings in the database (ie. to do the segmentation of the continuous recordings in epochs/trials). Then the imported epochs/trials (hard copies in .mat format) can be pre-processed and averaged. | Select > MEG > Display time series (or double-click on the file). |
| Line 13: | Line 25: |
| * Select the exploration mode: "Functional data (sorted by subject)"<<BR>><<BR>>{{attachment:view_functional.gif}} * Right-click on the subject node, and select: "Review raw file". Select the "MEG: CTF" file type, and pick the ds folder in "/sample_raw/Data". |
. {{attachment:link_menu.gif}} |
| Line 16: | Line 27: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawViewer?action=AttachFile&do=get&target=menuReview.gif|menuReview.gif|class="attachment"}} * Then you're asked if you want to "Refine the registration with the head points". This operation improves the initial MRI/MEG registration by fitting the head points digitized before the MEG acquisition on the scalp surface with an ICP algorithm. Answer yes. Even if the result is not perfect, it usually improves the positioning of the head in the MEG helmet. The grey surface represents the head extracted from the MRI, the yellow surface represents the inside of the MEG helmet, and the green dots are the head shape points digitized with the Polhemus device; the goal is to align the green points on the grey surface. |
It will open a new figure and enable many controls in the Brainstorm window. |
| Line 19: | Line 29: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawViewer?action=AttachFile&do=get&target=refine.gif|refine.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawViewer?action=AttachFile&do=get&target=refineBefore.gif|refineBefore.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawViewer?action=AttachFile&do=get&target=refineAfter.gif|refineAfter.gif|class="attachment"}} * Two new files appeared in the database explorer: |
{{attachment:review_epoch.gif}} |
| Line 23: | Line 31: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawViewer?action=AttachFile&do=get&target=linkInTree.gif|linkInTree.gif|class="attachment"}} * The channel file contains the definition of the sensors, exactly as when importing the files in the database with the "Import MEG/EEG" menu. It is saved in the folder ''(Common files)'', because the subject was created using the option "Yes, use one channel file per subject". Therefore, the same channel file will be used for all the folders of Subject01. * The node named "Link to raw file" contains all the information that was read from the continuous file (file format, time vector, sampling frequency, events, bad channels, path to the original file, etc.), but no recordings. The MEG and EEG values recorded will be read directly from the native file. |
== Navigate in time == The files we have imported here are shown the way they have been saved by the CTF MEG system: as contiguous epochs of 1 second each. These epochs are not related with the stimulus triggers or the subject's responses, they are just a way of saving the files. We will first explore the recordings in this epoched mode before switching to the continuous mode. |
| Line 27: | Line 34: |
| <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/ExploreAnatomy&next=Tutorials/EventMarkers")>> | ==== From the time series figure ==== * '''Click''': Click on the white or grey parts of figure to move the time cursor (red vertical line).<<BR>>If you click on the signals, it selects the corresponding channels. Click again to unselect. * '''Shortcuts''': See the tooltips in the time panel for important keyboard shortcuts: <<BR>>Left arrow, right arrow, page up, page down, F3, Shift+F3, etc... * '''Bottom bar''': The red square in the bottom bar represents the portion of the file that is currently displayed from the current file or epoch. Right now we show all the epoch #1. This will be more useful in the continuous mode. * '''Zoom''': Scroll to zoom horizontally around the time cursor (mouse wheel or two-finger up/down). * '''[<<<]''' and '''[>>>]''': Previous/next epoch or page ==== From the time panel ==== * '''Time''': [0, 998]ms is the time segment over which the first epoch is defined. * '''Sampling''': We downsampled these files to 600Hz for easier processing in the tutorials. * '''Text box''': Current time, can be edited manually. * '''[<]''' and '''[>]''': Previous/next time sample - Read the tooltip for details and shortcuts * '''[<<]''' and '''[>>]''': Previous/next time sample (x10) - Read the tooltip for details and shortcuts * '''[<<<]''' and '''[>>>]''': Previous/next epoch or page - Read the tooltip for details and shortcuts ==== From the page settings ==== * '''Epoch''': Selects the current time block that is displayed in the time series figure. * '''Start''': Starting point of the time segment displayed in the figure. Useful in continuous mode only. * '''Duration''': Length of this time segment. Useful in continuous mode only. ==== Time selection ==== * In the time series figure, click and drag your mouse for selecting a time segment. * At the bottom of the figure, you will see the duration of the selected block, and min/max values. * Useful for quickly estimating the latencies between two events, or the period of an oscillation. * Click anywhere on the figure to cancel this time selection. <<BR>><<BR>> {{attachment:review_timesel.gif}} == Epoched vs. continuous == * The CTF MEG system can save two types of files: epoched (.ds) or continuous (_AUX.ds). * Here we have an intermediate storage type: continuous recordings saved in "epoched" files. The files are saved as small blocks of recordings of a constant time length (1 second in this case). All these time blocks are contiguous, there is no gap between them. * Brainstorm can consider this file either as a continuous or an epoched file. By default it imports the regular .ds folders as epoched, but we need to change this manually. * Right-click on the "Link to raw file" for '''AEF#01''' > '''Switch epoched/continuous'''<<BR>>You should get a message: "File converted to: continuous". * Double-click on the "Link to raw file" again. Now you can navigate in the file without interruptions. The box "Epoch" is disabled and all the events in the file are displayed at once.<<BR>><<BR>> {{attachment:review_continuous.gif}} * With the red square at the bottom of the figure, you can navigate in time (click in the middle and drag with the mouse) or change the size of the current page (click on the left or right edge of the red square and move your mouse). <<BR>><<BR>> {{attachment:resize_page.gif}} * Increase the duration of the displayed window to '''3 seconds''' (Page settings > Duration). <<BR>><<BR>> {{attachment:review_setpage.gif}} * Close the figure. * Repeat this operation with the other files to convert them all to a continuous mode. * '''AEF#02 > Switch epoched/continuous ''' * '''Noise''' '''> Switch epoched/continuous ''' == Display mode: Butterfly/Column == * Close all the figures. * Double-click on the AEF#01 Link to raw file to open the MEG recordings. * What we see are all the traces of the 274 sensors overlaid on top of each other. * Click on the "Display mode" button in the toolbar of the Record tab. <<BR>><<BR>> {{attachment:review_switch.gif}} * All the signals are now displayed, one below the other, but because we have 274 MEG channels the figure is still unreadable. We need to select only a subset of these sensors. <<BR>><<BR>> {{attachment:review_column.jpg||width="381",height="186"}} == Montage selection == * You can use the montage menu to select a group of sensors. This menu is accessible in two ways: * Record toolbar > Drop-down menu. * Figure popup menu > Right-click on the figure > Montage * Pre-defined groups of channels are available for some common MEG and EEG systems.<<BR>>Notice the keyboard shortcut on the right for All channels (Shift+A). You can define your own (Shift+B, C...) if you go to Edit montages. * You can also use this menu to create your own sensor selections or more complex montages.<<BR>>A separate tutorial is dedicated to the montage editor. * Select the group: '''CTF LT''' (Left Temporal). <<BR>><<BR>> {{attachment:review_montage.gif||width="527",height="190"}} * More information about the [[Tutorials/MontageEditor|Montage editor]]. == Channel selection == If you click on the white or grey areas of the figure, it changes the current time. <<BR>>If you click on the lines representing the recorded signals instead, it selects the corresponding channels. * When some channels are selected, an additional menu "Channels" is visible in the figure popup. * Select "View selected" or press [Enter] to open the selected channels in a separate window. * The management of the bad channels will be introduced in a [[Tutorials/BadChannels|separate tutorial]].<<BR>><<BR>> {{attachment:channel_select.gif||width="615",height="246"}} == Amplitude scale == A variety of display options allows you to adjust the amplitude scale for the recordings (vertical axis). Most of these options are available in the right part of the time series figure, some are repeated in the Record tab of the Brainstorm window. . {{attachment:review_scale.gif}} * '''Increase/decrease gain''': Buttons '''[+]''' and '''[-]''' on the right side of the figure. The shortcuts for these buttons are indicated in the tooltips (leave the mouse for a short while over a button): right-click and move your mouse, hold the '''Shift key''' and scroll, or use the keys '''"+"''' and '''"-"'''. * '''Auto-scale amplitude''': Button '''[AS]''' in the figure. <<BR>>Selected: the vertical scale is adapted to the new maximum amplitude when you scroll in the file. <<BR>>Not selected: The vertical scale is fixed, scrolling in the file does not affect the axis resolution. * '''Flip Y axis''': Exchange the direction of the Y axis, to have the peaks of negative values pointing up. Useful mostly for clinical EEG. * '''Set amplitude scale''': Opens a window to enter the amplitude scale manually. The value corresponds to the space between two horizontal lines in this figure. * '''Set axis resolution''': See section "Time and amplitude resolution" below. * '''Remove DC offset''': Button '''[DC]''' in the Record tab. When selected, the average value over the entire current time window is subtracted from each channel. This means that if you change the length of the time window, the value that is removed from each channel may change. Always keep this option selected for unprocessed MEG recordings, unless you use a high-pass filter. * '''Normalize signals''': Divide each signal by its maximal amplitude in the displayed time window. The signals displayed with this normalization are unitless. * '''Apply CTF compensation''': Button [CTF] in the Record tab. Enable/disable the CTF noise correction based on the reference sensors, when it is not already applied in the file. In the current file, the CTF 3rd order gradient compensation is already applied, therefore this option is not available. * '''Vertical zoom''': Use the zoom/scrolll buttons on the right of the figure or your mouse (CTRL+Mouse wheel to zoom, middle-click+move to scroll) in order to look at specific channels without having to change the montage. . {{attachment:review_vscroll.gif}} * '''Uniform amplitude scales''': Force all the time series figures to use the same amplitude scale. Option available in the Record tab with the button {{attachment:uniform_button.gif}} or from the figure options menu when at least two time series figures are visible. [[https://neuroimage.usc.edu/brainstorm/Tutorials/MultipleWindows#Uniform_amplitude_scales|More details]]. <<TAG(Advanced)>> == Time and amplitude resolution == In the Brainstorm interface, the axis resolution is usually set implicitly: you can set the size of the window, the duration or recordings reviewed at once and the maximum amplitude to show in the figure. These parameters are convenient to explore the recordings interactively but don't allow us to have reproducible displays with constant time and amplitude resolutions. However, some applications are very sensitive to the horizontal and vertical scaling, such as the visual detection of epileptic spikes. The shapes of traces the epileptologists try to identify are altered by the axes resolution. This is detailed in the tutorial [[http://neuroimage.usc.edu/brainstorm/Tutorials/Epilepsy|EEG and Epilepsy]]. For this reason, we also added an option to set the figure resolution explicitly. The distance unit on a screen is the pixel, we can set precisely how much time is represented by one pixel horizontally and how much amplitude is represented by one pixel vertically. <<BR>>Display menu in the right part of the figure > Amplitude > Set axis resolution (shortcut: CTRL+O) Note that this interface does not store the input values, it just modifies the other parameters (figure size, time window, max amplitude) to fit the resolution objectives. If you modify these parameters after setting the resolution (resize the figure, leave the button [AS] selected and scroll in time, etc) the resolution is lost, you have to set it again manually. {{attachment:review_resolution.gif}} == Filters for visualization == With the Filter tab, you can apply a band-pass filter to the recordings, or remove a set of specific frequencies (example: the 50Hz or 60Hz power lines contamination and their harmonics). The filters are applied only to the time window that is currently loaded. If the segment is too short for the required filters, the results might be inaccurate. These visualization filters provide a quick estimate for visualization only, the results are not saved anywhere. To filter properly the continuous files, please use the Process1 tab (see [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsFilter#What_filters_to_apply.3F|tutorial #10]]).<<BR>>The option "'''Filter all results'''" is not useful for now, it will be described later. After testing the high-pass, low-pass and notch filters, uncheck them. Otherwise you may forget about them, they would stay on until you restart Brainstorm. Note that as long as there are visualization filters applied, the title of the Filter tab remains red. . {{attachment:review_filter.gif}} == Mouse and keyboard shortcuts == ==== Keyboard shortcuts ==== * '''Left / right arrows''': * Change current time, sample by sample * '''+Control '''key: Jump to previous/next epoch or page (same as [<<<] and [>>>]) * '''+Shift '''key: Jump to previous/next event (you need to have one event selected) * MacOS: These shortcuts are different, please read the tooltips for [>], [>>] and [>>>] * '''Page-up / page-down''': * Change current time, 10 samples at a time * '''+Control '''key: Jump to the next/previous epoch or page, 10x faster * '''F3/Shift+F3''': Jump to the next/previous epoch or page (10% overlap between 2 pages) * '''F4/Shift+F4''': Jump to the next/previous half-page (50% overlap) * '''F6/Shift+F6''': Jump to the next/previous page with no overlap (0% overlap) * '''Plus / minus''': Adjust the vertical scale of the time series * '''Shift + Letter''': Changes the montage * '''Control + B''': Mark selected time segment as bad * '''Control + D''': Dock figure * '''E''': Add / delete event marker * '''Control + E''': Add / delete event marker for the selected channels * '''Control + F''': Open a copy of the figure, not managed by the Brainstorm window manager * '''Control + H''': Hide/show selected event group * '''Control + I''': Save figure as image * '''Control + J''': Open a copy of the figure as an image * '''Control + O''': Set axes resolution * '''Control + L''': Change display mode of events (dots, lines or hidden) * '''Control + J''': Open a screen capture of the figure * '''Control + T''': Open a 2D topography window at the current time * '''Enter''': Display the selected channels in a separate figure * '''Escape''': Unselect all the selected channels * '''Delete''': Mark the selected channels as bad * '''1 2 3 4 5 6 7 8 9''': User-defined shortcuts for new events ([[https://neuroimage.usc.edu/brainstorm/Tutorials/EventMarkers#Custom_shortcuts|tutorial #7]]) ==== Mouse shortcuts ==== * '''Click on a channel''': Select the channel * '''Click''': Change current time * '''Shift + click''': Force the selection of the current time (even when clicking on a channel) * '''Click + move''': Select time range * '''Right-click''': Display popup menu * '''Right-click + move''': Adjust the vertical scale of the time series * '''Scroll''': Zoom around current time * '''Shift + scroll''': Adjust the vertical scale of the time series * '''Control + scroll''': Zoom vertically * '''Central click + move''': Move in a zoomed figure * '''Double click''': Restore initial zoom settings (or edit the notes associated to the clicked event) <<HTML(<!-- END-PAGE -->)>> <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/ChannelFile&next=Tutorials/MultipleWindows")>> |
Tutorial 5: Review continuous recordings
Authors: Francois Tadel, Elizabeth Bock, John C Mosher, Sylvain Baillet
Contents
Open the recordings
Let's look at the first file in the list: AEF#01.
Right-click on the Link to raw file. Below the first to menus, you have the list of channel types:
MEG: 274 axial gradiometers
ECG: 1 electrocadiogram, bipolar electrode across the chest
EOG: 2 electrooculograms (vertical and horizontal)
Misc: EEG electrodes Cz and Pz
ADC A: Unused
ADC V: Auditory signal sent to the subject
DAC: Unused
FitErr: Fitting error when trying to localize the three head localization coils (NAS, LPA, RPA)
HLU: Head Localizing Unit, displacements in the three directions (x,y,z) for the three coils
MEG REF: 26 reference sensors used for removing the environmental noise
Other: Unused
Stim: Stimulation channel, records the stim triggers generated by the Psychophysics toolbox and other input channels, such as button presses generated by the subject
SysClock: System clock, unused
Select > MEG > Display time series (or double-click on the file).
It will open a new figure and enable many controls in the Brainstorm window.
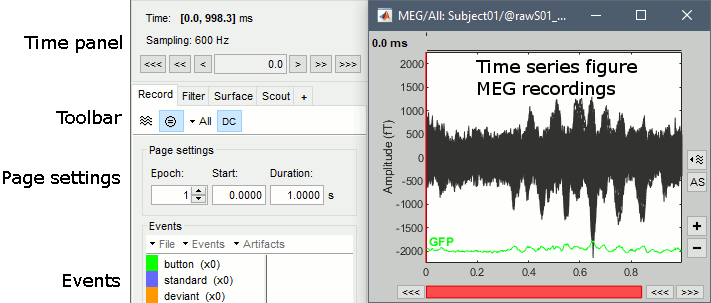
Navigate in time
The files we have imported here are shown the way they have been saved by the CTF MEG system: as contiguous epochs of 1 second each. These epochs are not related with the stimulus triggers or the subject's responses, they are just a way of saving the files. We will first explore the recordings in this epoched mode before switching to the continuous mode.
From the time series figure
Click: Click on the white or grey parts of figure to move the time cursor (red vertical line).
If you click on the signals, it selects the corresponding channels. Click again to unselect.Shortcuts: See the tooltips in the time panel for important keyboard shortcuts:
Left arrow, right arrow, page up, page down, F3, Shift+F3, etc...Bottom bar: The red square in the bottom bar represents the portion of the file that is currently displayed from the current file or epoch. Right now we show all the epoch #1. This will be more useful in the continuous mode.
Zoom: Scroll to zoom horizontally around the time cursor (mouse wheel or two-finger up/down).
[<<<] and [>>>]: Previous/next epoch or page
From the time panel
Time: [0, 998]ms is the time segment over which the first epoch is defined.
Sampling: We downsampled these files to 600Hz for easier processing in the tutorials.
Text box: Current time, can be edited manually.
[<] and [>]: Previous/next time sample - Read the tooltip for details and shortcuts
[<<] and [>>]: Previous/next time sample (x10) - Read the tooltip for details and shortcuts
[<<<] and [>>>]: Previous/next epoch or page - Read the tooltip for details and shortcuts
From the page settings
Epoch: Selects the current time block that is displayed in the time series figure.
Start: Starting point of the time segment displayed in the figure. Useful in continuous mode only.
Duration: Length of this time segment. Useful in continuous mode only.
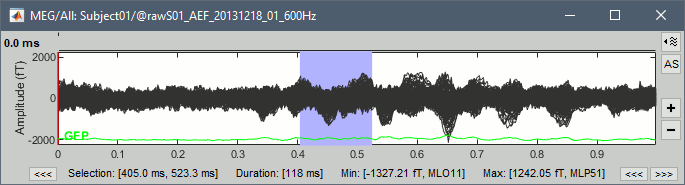
Time selection
- In the time series figure, click and drag your mouse for selecting a time segment.
- At the bottom of the figure, you will see the duration of the selected block, and min/max values.
- Useful for quickly estimating the latencies between two events, or the period of an oscillation.
Click anywhere on the figure to cancel this time selection.
Epoched vs. continuous
- The CTF MEG system can save two types of files: epoched (.ds) or continuous (_AUX.ds).
- Here we have an intermediate storage type: continuous recordings saved in "epoched" files. The files are saved as small blocks of recordings of a constant time length (1 second in this case). All these time blocks are contiguous, there is no gap between them.
- Brainstorm can consider this file either as a continuous or an epoched file. By default it imports the regular .ds folders as epoched, but we need to change this manually.
Right-click on the "Link to raw file" for AEF#01 > Switch epoched/continuous
You should get a message: "File converted to: continuous".Double-click on the "Link to raw file" again. Now you can navigate in the file without interruptions. The box "Epoch" is disabled and all the events in the file are displayed at once.
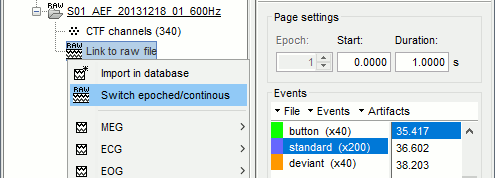
With the red square at the bottom of the figure, you can navigate in time (click in the middle and drag with the mouse) or change the size of the current page (click on the left or right edge of the red square and move your mouse).
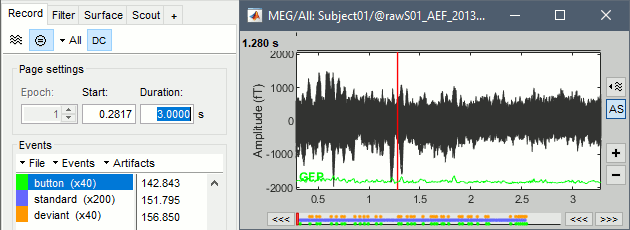
Increase the duration of the displayed window to 3 seconds (Page settings > Duration).
- Close the figure.
- Repeat this operation with the other files to convert them all to a continuous mode.
AEF#02 > Switch epoched/continuous
Noise > Switch epoched/continuous
Display mode: Butterfly/Column
- Close all the figures.
- Double-click on the AEF#01 Link to raw file to open the MEG recordings.
- What we see are all the traces of the 274 sensors overlaid on top of each other.
Click on the "Display mode" button in the toolbar of the Record tab.
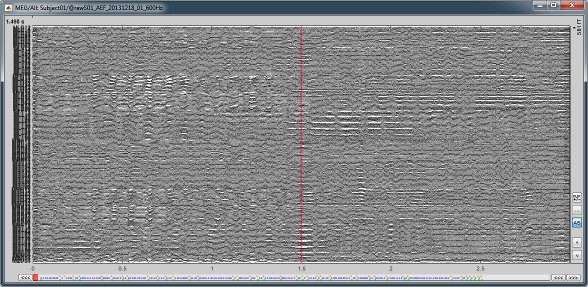
All the signals are now displayed, one below the other, but because we have 274 MEG channels the figure is still unreadable. We need to select only a subset of these sensors.
Montage selection
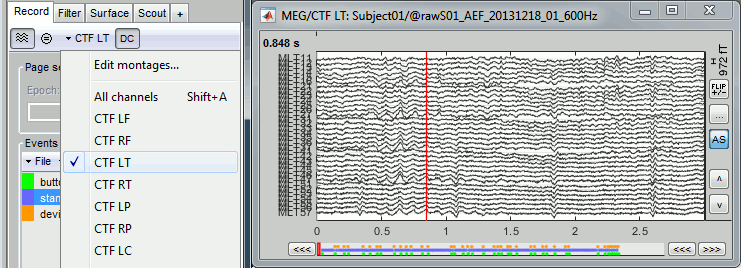
- You can use the montage menu to select a group of sensors. This menu is accessible in two ways:
Record toolbar > Drop-down menu.
Figure popup menu > Right-click on the figure > Montage
Pre-defined groups of channels are available for some common MEG and EEG systems.
Notice the keyboard shortcut on the right for All channels (Shift+A). You can define your own (Shift+B, C...) if you go to Edit montages.You can also use this menu to create your own sensor selections or more complex montages.
A separate tutorial is dedicated to the montage editor.Select the group: CTF LT (Left Temporal).
More information about the Montage editor.
Channel selection
If you click on the white or grey areas of the figure, it changes the current time.
If you click on the lines representing the recorded signals instead, it selects the corresponding channels.
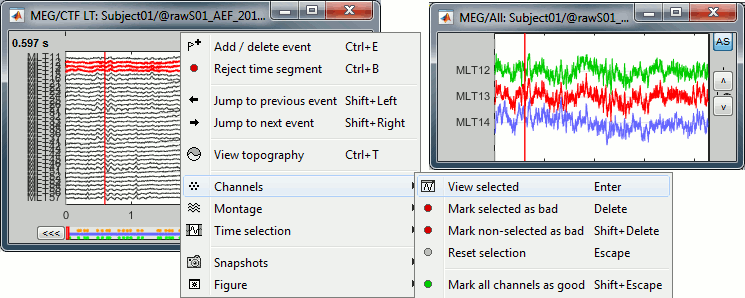
- When some channels are selected, an additional menu "Channels" is visible in the figure popup.
- Select "View selected" or press [Enter] to open the selected channels in a separate window.
The management of the bad channels will be introduced in a separate tutorial.
Amplitude scale
A variety of display options allows you to adjust the amplitude scale for the recordings (vertical axis). Most of these options are available in the right part of the time series figure, some are repeated in the Record tab of the Brainstorm window.
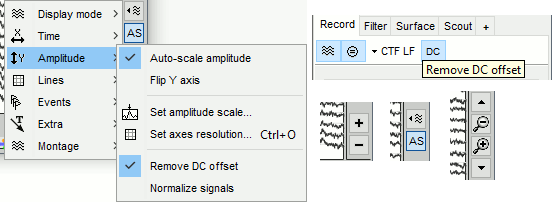
Increase/decrease gain: Buttons [+] and [-] on the right side of the figure. The shortcuts for these buttons are indicated in the tooltips (leave the mouse for a short while over a button): right-click and move your mouse, hold the Shift key and scroll, or use the keys "+" and "-".
Auto-scale amplitude: Button [AS] in the figure.
Selected: the vertical scale is adapted to the new maximum amplitude when you scroll in the file.
Not selected: The vertical scale is fixed, scrolling in the file does not affect the axis resolution.Flip Y axis: Exchange the direction of the Y axis, to have the peaks of negative values pointing up. Useful mostly for clinical EEG.
Set amplitude scale: Opens a window to enter the amplitude scale manually. The value corresponds to the space between two horizontal lines in this figure.
Set axis resolution: See section "Time and amplitude resolution" below.
Remove DC offset: Button [DC] in the Record tab. When selected, the average value over the entire current time window is subtracted from each channel. This means that if you change the length of the time window, the value that is removed from each channel may change. Always keep this option selected for unprocessed MEG recordings, unless you use a high-pass filter.
Normalize signals: Divide each signal by its maximal amplitude in the displayed time window. The signals displayed with this normalization are unitless.
Apply CTF compensation: Button [CTF] in the Record tab. Enable/disable the CTF noise correction based on the reference sensors, when it is not already applied in the file. In the current file, the CTF 3rd order gradient compensation is already applied, therefore this option is not available.
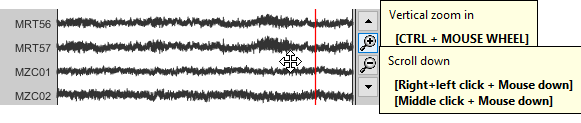
Vertical zoom: Use the zoom/scrolll buttons on the right of the figure or your mouse (CTRL+Mouse wheel to zoom, middle-click+move to scroll) in order to look at specific channels without having to change the montage.
Uniform amplitude scales: Force all the time series figures to use the same amplitude scale. Option available in the Record tab with the button
 or from the figure options menu when at least two time series figures are visible. More details.
or from the figure options menu when at least two time series figures are visible. More details.
Time and amplitude resolution
In the Brainstorm interface, the axis resolution is usually set implicitly: you can set the size of the window, the duration or recordings reviewed at once and the maximum amplitude to show in the figure. These parameters are convenient to explore the recordings interactively but don't allow us to have reproducible displays with constant time and amplitude resolutions.
However, some applications are very sensitive to the horizontal and vertical scaling, such as the visual detection of epileptic spikes. The shapes of traces the epileptologists try to identify are altered by the axes resolution. This is detailed in the tutorial EEG and Epilepsy.
For this reason, we also added an option to set the figure resolution explicitly. The distance unit on a screen is the pixel, we can set precisely how much time is represented by one pixel horizontally and how much amplitude is represented by one pixel vertically.
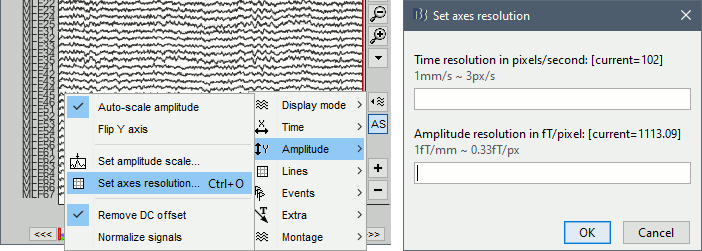
Display menu in the right part of the figure > Amplitude > Set axis resolution (shortcut: CTRL+O)
Note that this interface does not store the input values, it just modifies the other parameters (figure size, time window, max amplitude) to fit the resolution objectives. If you modify these parameters after setting the resolution (resize the figure, leave the button [AS] selected and scroll in time, etc) the resolution is lost, you have to set it again manually.
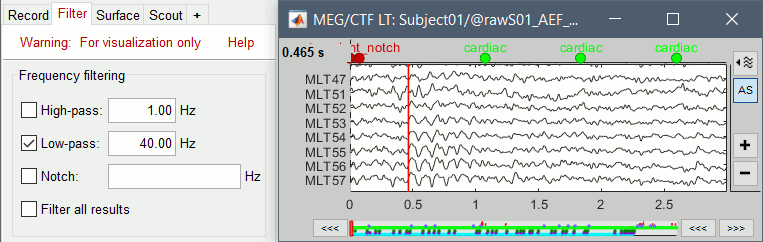
Filters for visualization
With the Filter tab, you can apply a band-pass filter to the recordings, or remove a set of specific frequencies (example: the 50Hz or 60Hz power lines contamination and their harmonics). The filters are applied only to the time window that is currently loaded. If the segment is too short for the required filters, the results might be inaccurate.
These visualization filters provide a quick estimate for visualization only, the results are not saved anywhere. To filter properly the continuous files, please use the Process1 tab (see tutorial #10).
The option "Filter all results" is not useful for now, it will be described later.
After testing the high-pass, low-pass and notch filters, uncheck them. Otherwise you may forget about them, they would stay on until you restart Brainstorm. Note that as long as there are visualization filters applied, the title of the Filter tab remains red.
Mouse and keyboard shortcuts
Keyboard shortcuts
Left / right arrows:
- Change current time, sample by sample
+Control key: Jump to previous/next epoch or page (same as [<<<] and [>>>])
+Shift key: Jump to previous/next event (you need to have one event selected)
MacOS: These shortcuts are different, please read the tooltips for [>], [>>] and [>>>]
Page-up / page-down:
- Change current time, 10 samples at a time
+Control key: Jump to the next/previous epoch or page, 10x faster
F3/Shift+F3: Jump to the next/previous epoch or page (10% overlap between 2 pages)
F4/Shift+F4: Jump to the next/previous half-page (50% overlap)
F6/Shift+F6: Jump to the next/previous page with no overlap (0% overlap)
Plus / minus: Adjust the vertical scale of the time series
Shift + Letter: Changes the montage
Control + B: Mark selected time segment as bad
Control + D: Dock figure
E: Add / delete event marker
Control + E: Add / delete event marker for the selected channels
Control + F: Open a copy of the figure, not managed by the Brainstorm window manager
Control + H: Hide/show selected event group
Control + I: Save figure as image
Control + J: Open a copy of the figure as an image
Control + O: Set axes resolution
Control + L: Change display mode of events (dots, lines or hidden)
Control + J: Open a screen capture of the figure
Control + T: Open a 2D topography window at the current time
Enter: Display the selected channels in a separate figure
Escape: Unselect all the selected channels
Delete: Mark the selected channels as bad
1 2 3 4 5 6 7 8 9: User-defined shortcuts for new events (tutorial #7)
Mouse shortcuts
Click on a channel: Select the channel
Click: Change current time
Shift + click: Force the selection of the current time (even when clicking on a channel)
Click + move: Select time range
Right-click: Display popup menu
Right-click + move: Adjust the vertical scale of the time series
Scroll: Zoom around current time
Shift + scroll: Adjust the vertical scale of the time series
Control + scroll: Zoom vertically
Central click + move: Move in a zoomed figure
Double click: Restore initial zoom settings (or edit the notes associated to the clicked event)