Thank you Francois, do you have subject for 10 years old ?
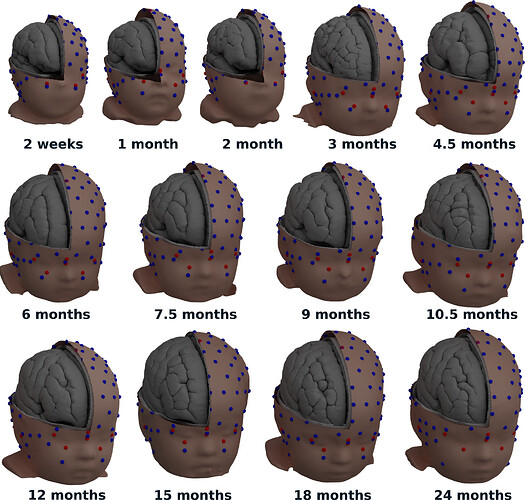
Not yet. All the templates are available for download directly with your up-to-date Brainstorm installation, and documented here:
https://neuroimage.usc.edu/brainstorm/Tutorials/DefaultAnatomy#FreeSurfer_templates
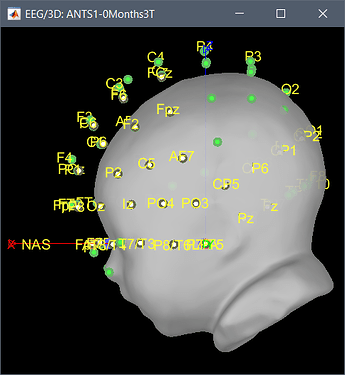
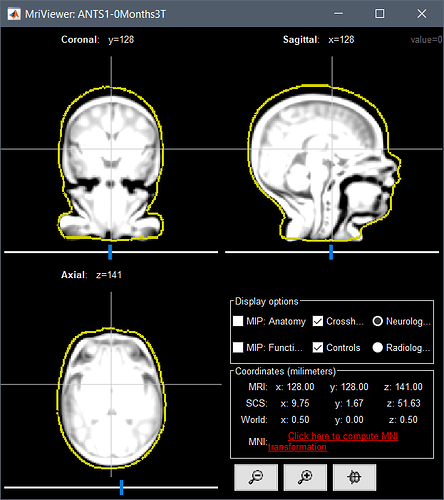
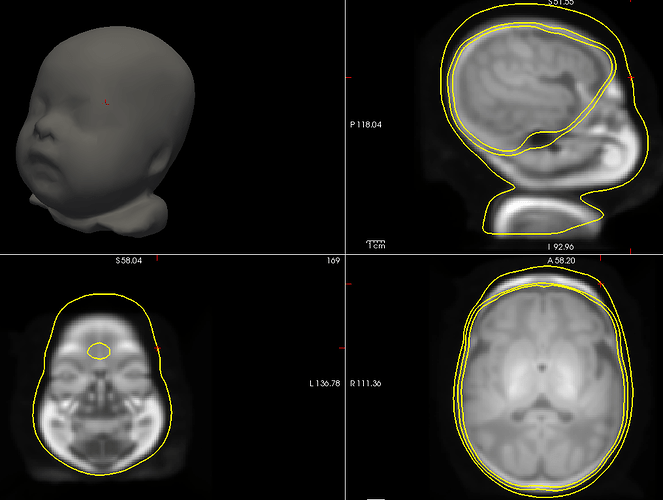
A solution that could be interesting for you, since you have a list of digitized head points that covers all the head uniformly, is to warp a template (ICBM152 or Oreilly_24m) to these head points, after manual/approximate registration.
https://neuroimage.usc.edu/brainstorm/Tutorials/TutWarping
it is correct to exported the meg squid (extracted from vecterview example in bst ) exported to matlab and imported again with this new subject
I'm sorry, I don't understand what you are trying to do.
If you need help, please try to describe what you are doing, starting from an empty database.
but yes thanks to the edit we can correct it
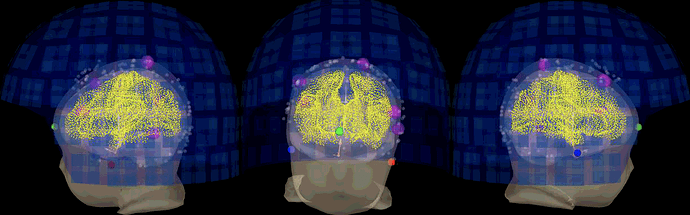
You are not really supposed to move the MEG sensors around the head... If the subject was not sitting correctly in the MEG, artificially moving the sensors to a "better looking" position in Brainstorm on't change the position of the head that the sensors were actually recording.
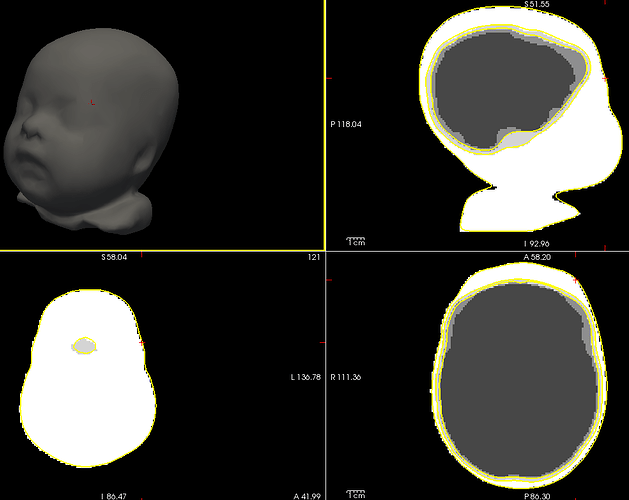
In your screen captures, the head looks very far from the sensors, are you sure it was really like this? If so, you might not be recording a lot of brain signal.
but fro the back ot seems the distance between sensor and scalp is big,
The actual head (green digitized head points) is much bigger than the template your are using. Try warping your template to these points.
despite that we have different type of sensor the display sensors have not the othe options
what should be the problem?
You don't have 3D positions available for the STI/EOG/ECG data channels, therefore you can't display their positions in space. This is normal.