I am unable to launch BrSt. Here is the sequence of events:
- launched BrSt, did some exploration of reducing the # of vertices in the cortical mesh.
- I eventually tried the 3rd option, inhomogeneous, and was informed I needed to download the iso2mesh package
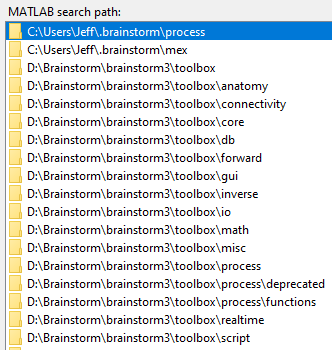
- I did so, and added its path to Matlab
- I closed and re-launched Matlab OK
- I tried to launch brainstorm, and got the following console response:
Tuesday, February 20, 2024
1:36 PM
>> brainstorm
BST> Starting Brainstorm:
BST> =================================
BST> Version: 06-Feb-2024
BST> Compiling main interface files...
BST> Deleting old process reports...
BST> Loading configuration file...
BST> Checking internet connectivity... ok
BST> Update available online: 20-Feb-2024
BST> Initializing user interface...
BST> Starting OpenGL engine... hardware
BST> Plugin mff: D:\downloads\EEGlab\eeglab_current\eeglab2023.1\plugins\MFFMatlabIO4.1
BST> Reading process folder...
BST> Loading current protocol...
Error using bst_get
Subject name not specified.
Error in node_create_db_subjects (line 58)
CurrentSubject = bst_get('Subject');
Error in panel_protocols>UpdateTree (line 432)
case 'Subjects', [selNode, dbNode, numNodes] = node_create_db_subjects(nodeRoot, iSearch);
Error in panel_protocols>CreatePanel/DatabaseTabChanged_Callback (line 316)
UpdateTree(0);
Error in panel_protocols>CloseAllDatabaseTabs (line 1569)
bakCallback();
Error in panel_protocols (line 46)
eval(macro_method);
Error in gui_brainstorm>SetCurrentProtocol (line 955)
panel_protocols('CloseAllDatabaseTabs');
Error in gui_brainstorm (line 34)
eval(macro_method);
Error in bst_startup (line 643)
gui_brainstorm('SetCurrentProtocol', GlobalData.DataBase.iProtocol);
Error in brainstorm (line 123)
bst_startup(BrainstormHomeDir, 1, BrainstormDbDir);
Here is what the screen showed. It never finished so I closed it down.
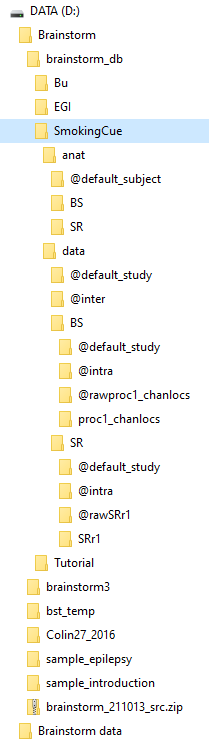
I thought the iso2mesh path might be the problem, so I removed that path and tried again and got the same result. I am now unable to continue my BrSt work. Am I going to have to re-install BrSt? Or perhaps rebuild my database? Cannot do that if I cannot start BrSt though.