Hello everyone,
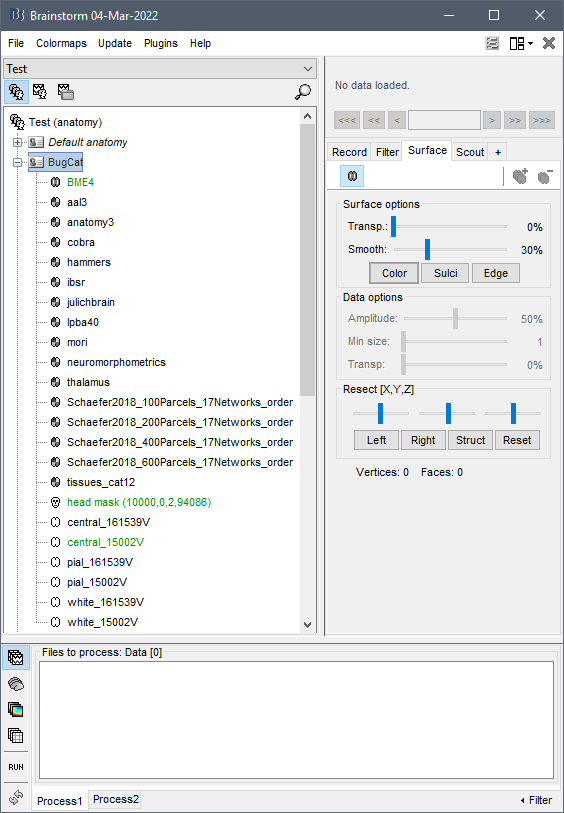
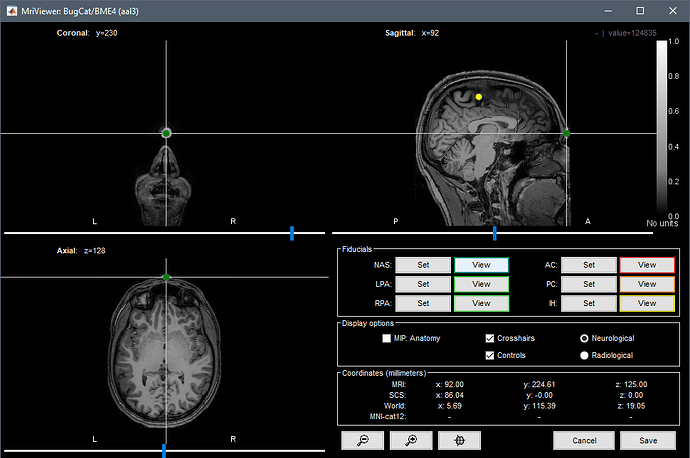
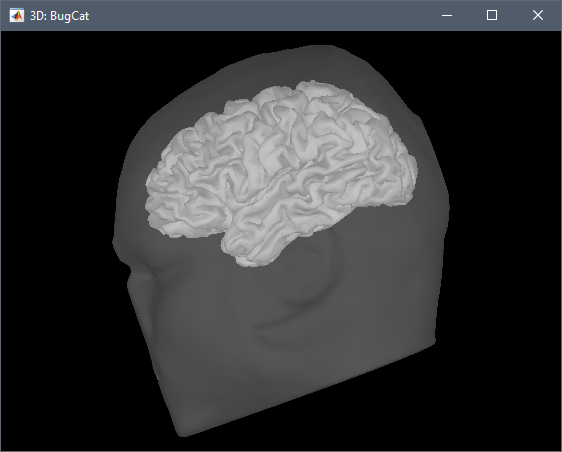
When I run CAT12 segmentation on this MRI, I get the following error:
CAT Preprocessing error for BME4:
Input to SVD must not contain NaN or Inf.
25 - encode_qform0
109 - fun
20 - subsasgn
109 - create_vol
17 - spm_create_vol
83 - spm_write_vol
518 - cat_vol_sanlm_filter
236 - cat_vol_sanlm_file
151 - cat_vol_sanlm
269 - cat_run_job1639
40 - cat_run_newcatch
1084 - run_job
639 - cat_run
29 - cfg_run_cm
1717 - local_runcj
972 - cfg_util
469 - fill_run_job
247 - spm_jobman
359 - Compute
442 - ComputeInteractive
28 - process_segment_cat12
28 - bst_call
3004 - @(h,ev)bst_call(@process_segment_cat12,'ComputeInteractive',iSubject,iAnatomy)
Error:cat_io_report:CATgui: Error in cat_io_report GUI parameter report creation > incomple image parameters.
Error:cat_io_report:Fig: Error in CAT report figure creation!
Warning: Matrix is singular, close to singular or badly scaled. Results may be inaccurate. RCOND = NaN.
My CAT12 and SPM are updated to the latest version. Is the issue with my MRI?
Thank you,
Jennifer