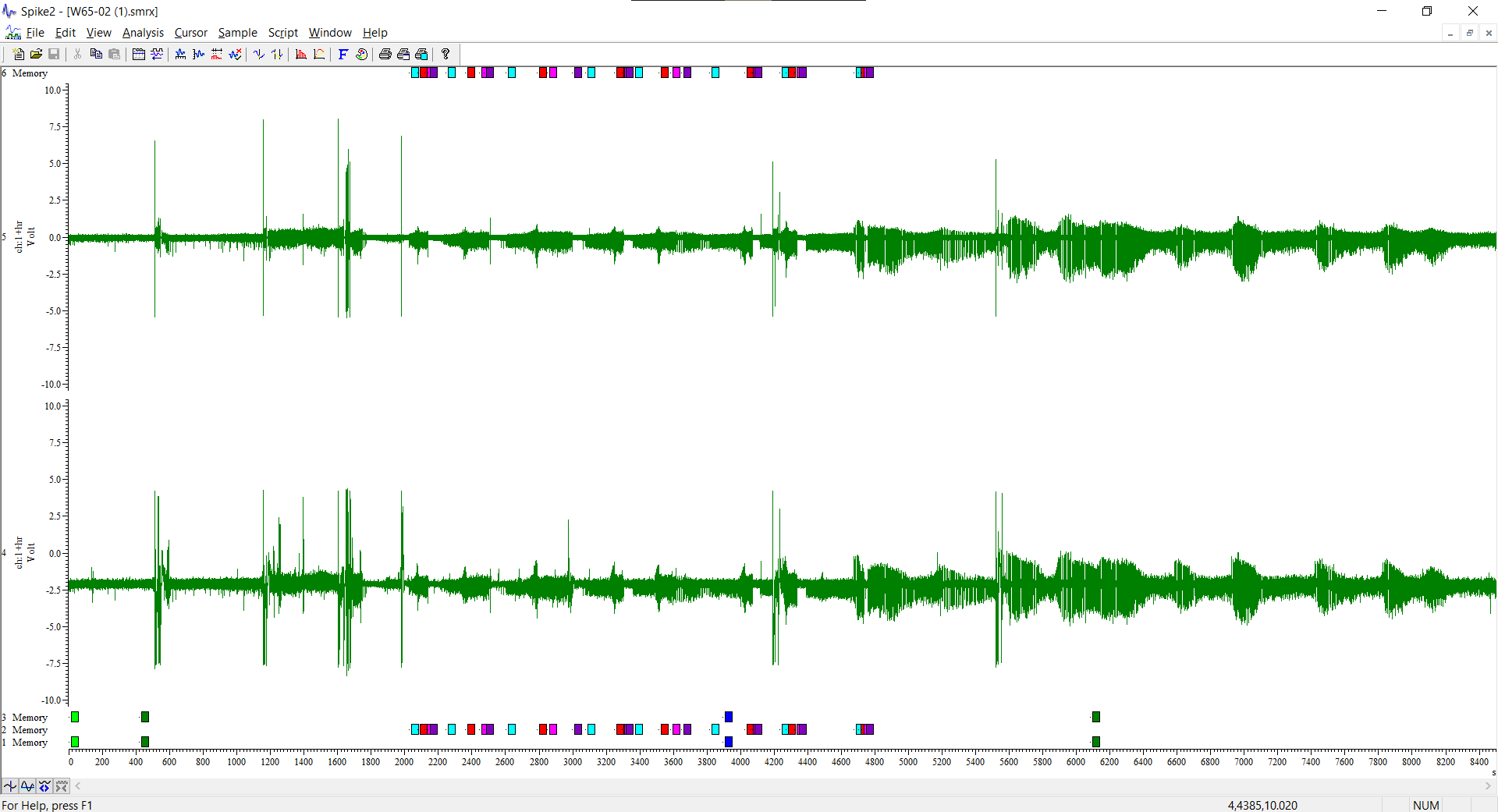
Hey! I am a new user. we want to calculate PSD in brainstorm, we search for individual spikes in Spike2 (using the same exact smrx file in both sowftares) and type in their position.
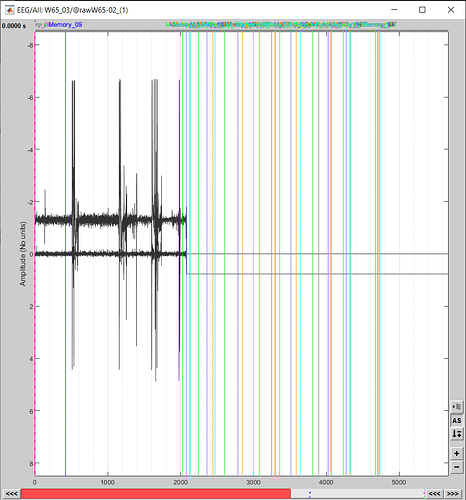
Is it possible to visually check these spikes (on an eeg) in brainstorm? (meaning, after entering the start&end, to see if brainstorm is "looking at" the same thing we want to "look at" based on Spike2)
Hi @Zsolt, welcome to Brainstorm.
For new users, we recommend you start by following the introduction tutorials first (section "Get started" on the tutorial page), using the example dataset provided: https://neuroimage.usc.edu/brainstorm/Tutorials
Once you gone through the Get started tutorials, you can check this advance tutorials on electrophysiology: https://neuroimage.usc.edu/brainstorm/e-phys/Introduction
Yes, it is possible. Just set those parameters in the Record tab
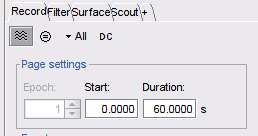
If you are working with already sorted spikes, it should possible to export the spikes information from Spik2 (type, channel, start, duration) and import it in Brainstorm. This is shown in the advanced electrophysiology tutorials
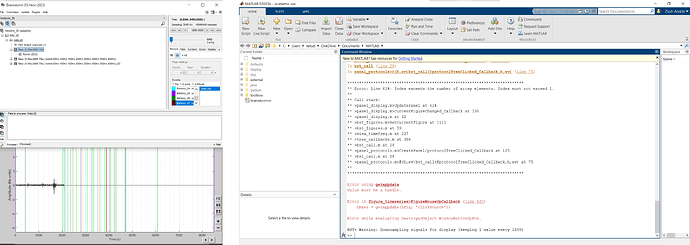
Is there any message in the Matlab Command Window?
What happens when the you double-click on the file Raw (0.00s, 8499.70s)?
Hey there!
I wanted to import an EEG (smrx format), but but only "managed" the first third of the EEG. What do I do now? Is there a solution to solve it? (There are other similar files that we have successfully imported, but there are a few cases in which it "cuts off" the second and third thirds of the eeg.)
Opening these files in the Spike2 displays the whole eeg recording.
@Zsolt, could you provide one of the files that is causing troubles?
Upload the file somewhere and post the download links here.
We managed to solve the problem!
@Zsolt, for sake of completeness.
Could you indicate what was the issue and how it was solved?
Thank you