Hello Brainstorm team,
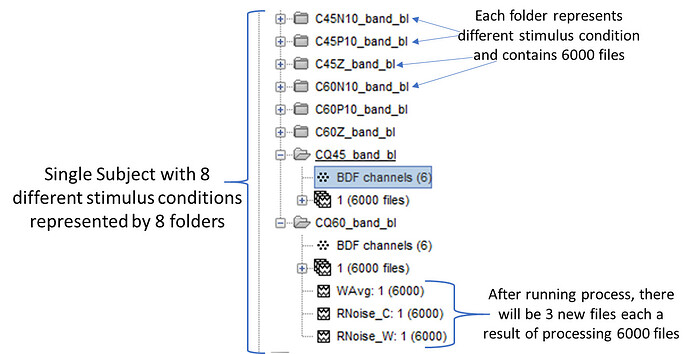
I have a process which generates three files from X number of epochs (in this case 6000 files).
Each subject (and I have 20 subjects) has 8 different stimulus conditions represented by 8 different folders and each folder contains 6000 files of epochs.
The process works well but it does take some time to run (approximately 15 mins).
Therefore rather than waiting for each folder to finish going through the process, I would like to select and drag 6000 files from each folder into the Process tab and run the process and leave the computer on overnight to do this.
The end result would be that each folder would contain 6000 files and underneath 3 different new files for every single folder for every subject (I do not want processing across folders or across subjects).
I hope the explanation is clear.
Looking at the "Averaging" tutorial (https://neuroimage.usc.edu/brainstorm/Tutorials/Averaging#Process_options:_Average), it seems like the condition that I need is By trial group (folder average) would you agree?
And also what would be the code required to do this?
I looked into "process_average" within the Brainstorm process folder to get some help and it seems to be under %% ===== SORT FILES =====.
How would I implement this into my custom code? I attach my code to this query for review.
process_weighted_average_and_residual_noise.m (8.5 KB)
You have two possible approaches:
1 - Within your process: Add a classification of the input files in multiple groups (by folder) to be processed separately. You can call directly the sorting function from process_average.
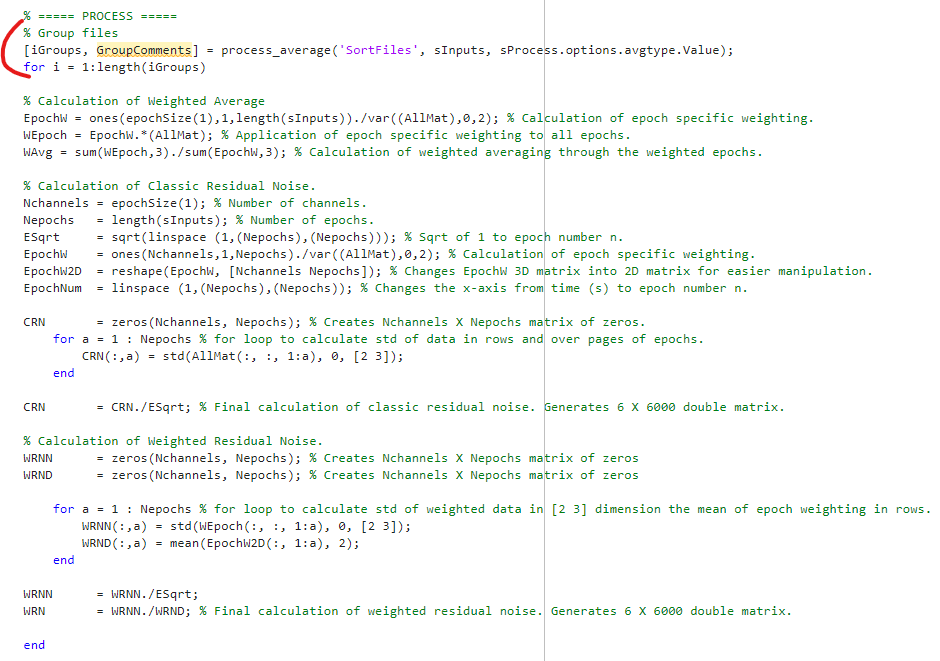
% Group files
avgtype = 3;
[iGroups, GroupComments] = process_average('SortFiles', sInputs, avgtype);
% Average each group
for i = 1:length(iGroups)
% Your code, adapted from lines 175-190 of process_average.m
end
This solution would be easy to add to your current process function. If you select this option, then I recommend you keep all the possible options available for avgtype in process_average. If you implement only what you need (ie. avgtype=3), it would be more difficult to share your process with other users, as they might not expect it to run ONLY in one configuration (per folder and per subject).
2 - The other option is to keep your process as it is (processing all the input files together at once), and write a script that calls in a loop. Your script would loop over subjects and folders in your database, search all the data files in each subject/folder, and call the process on it.
This would require a bit more reading about Brainstorm scripting and database requests:
Thank you Francois - I think the first option sounds more reasonable for my situation.
Based on your recommendation this is what I've done so far.
-
Included the following under %%GET DESCRIPTION to keep all the possible options available for avgtype.
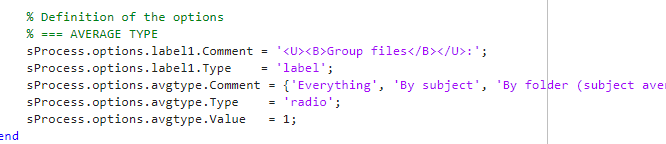
-
Added the following code based on your suggestion. This is just before where all the calculations take place (i.e. my custom code). I understand that selecting a specific avgtype will give a "value" output which "sProcess.options.avgtype.Value" will use to decide what kind of avgtype to run.
Is this what you had in mind? Can you see any issues?
I was not sure where and how to include the "code" you adapted.
You wrote in your comment "% Your code" so I assumed my code should be inserted between the "for" and "end".
Hello Francois,
So I've modified the code that you've provided to keep all the possible options available for avgtype. [iGroups, GroupComments] = process_average('SortFiles', sInputs, sProcess.options.avgtype.Value);
I've added the "avgtype" description under "%% Get Description" so it should provide a value to specify the "avgtype".
I understand that (please correct me if I'm wrong):
- "process_average('SortFiles'" calls for the "SortFiles" function. I know that this means I don't have to copy and paste the entire "SortFiles" function from process_average.
- "sInputs" is the input structure.
- "sProcess.options.avgtype.Value" specifies the avgtype (i.e. everthing, by subject, by condition etc) based on the selection made.
Based on the above information I need to implement the following code somewhere within my code:
[iGroups, GroupComments] = process_average('SortFiles', sInputs, sProcess.options.avgtype.Value);
for i = 1:length(iGroups)
end
What I'm having trouble implementing is the [iGroups, GroupComments] part and the "for" loop with "i = 1:length(iGroups)" where and how do I implement this?
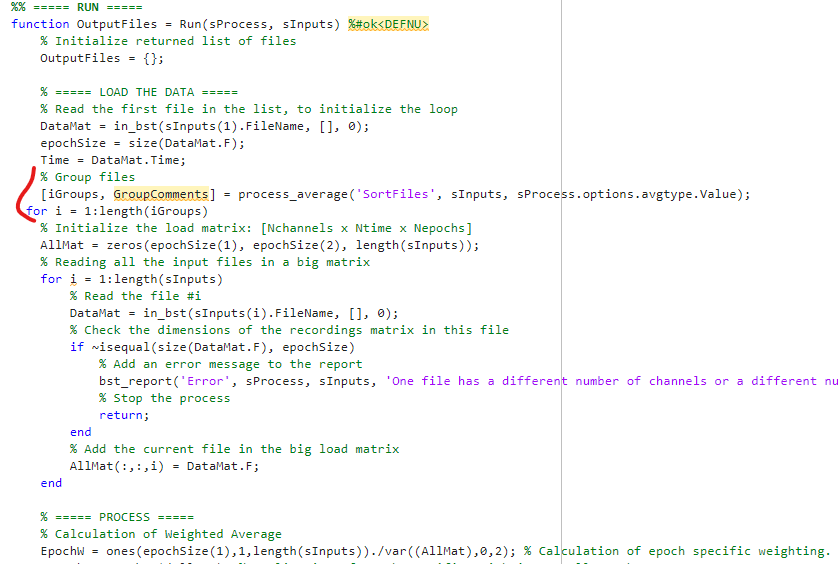
Looking at "process_average" the "%Group files" is under "%%Run" but I'm not sure where under the "%%Run" I should insert this code.
At the moment, I've included the code just before where the actual data manipulation and calculation starts. But I feel like this code needs to be somewhere higher up within "% Load the Data"
Any suggestions? Bear in mind I wrote the following code from "[process_example_customavg.m]".
Everything you've done so far looks good.
What I'm having trouble implementing is the [iGroups, GroupComments] part and the "for" loop with "i = 1:length(iGroups)" where and how do I implement this?
Your computation and the "SAVE THE RESULTS" block both go inside the for loop.
Instead of using sInputs, you must use sInputs(iGroups(i)), in order to select only the files within the input group at each iteration. Same goes for AllMat => AllMat(:, :, iGroups(i))
Alternatively, you could move the block that loads all the input files into AllMat inside the group for loop as well. This will be lighter in memory (not all the trials of all the subjects are loaded at once, only the ones that are averaged together), and probably easier to implement as you don't have to add any indices when accessing AllMat.
You need to be careful about the STUDY in which you save each average. Use GetOutputStudy, as in process_average.m, instead of using the path of the first input file, as in process_example_customavg (fileparts(sInputs(1).FileName))
[sStudy, iStudy, Comment, uniqueDataFile] = bst_process('GetOutputStudy', sProcess, sInputs(iGroups(i)));
In case you wonder how all these options work more precisely, this is all documented in this tutorial:
https://neuroimage.usc.edu/brainstorm/Tutorials/TutUserProcess
Thanks for your input Francois I appreciate it!
-
As per your suggestion I've included the block that loads all the input files into AllMat to the "SAVE THE RESULTS" section within the "for" loop for grouping. As I included the AllMat within the "for" loop for grouping I did NOT change AllMat(: , : , i) into AllMat(:, :, iGroups(i)).
I hope what I'm showing here is correct.
-
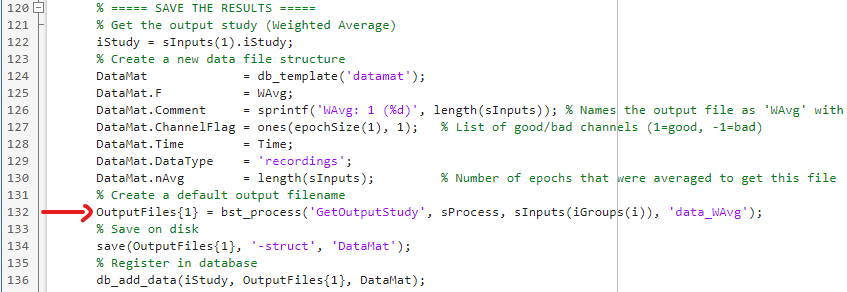
As per your suggestion I've used GetOutputStudy and changed line 132 from:
OutputFiles{1} = bst_process('GetNewFilename', fileparts(sInputs(1).FileName), 'data_WAvg');
to
OutputFiles{1} = bst_process('GetOutputStudy', sProcess, sInputs(iGroups(i)), 'data_WAvg');
However, I then get an error message saying that "Index exceeds the number of array elements. Index must not exceed 4."
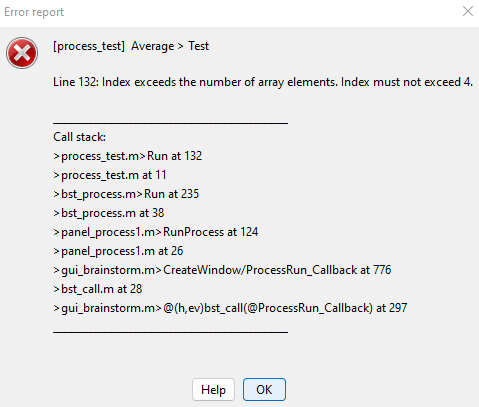
So I then went through "bst_process.m" and discovered that 'GetOutputStudy' only requires 3 inputs:
[sStudy, iStudy, Comment, uniqueDataFile] = GetOutputStudy(sProcess, [sInputA, sInputB], sOutput.Condition);
Is this why I am getting the error message? How do I solve this issue?
As per your suggestion I've included the block that loads all the input files into AllMat
But I can see you didn't replace all the sInputs with sInputs(iGroups(i)).
This is not going to work.
As per your suggestion I've used GetOutputStudy and changed line 132 from:
Please see how this is done in process_average.m.
These questions are related with Matlab programming, and not Brainstorm. There are beyond the scope of this forum, which is to provide help in using Brainstorm. Maybe you need to train deeper with Matlab coding before getting back to this task.
Read entirely the tutorial "How to write your own process". Then put a breakpoint at the beginning of the Run() function, start the process from the Brainstorm interface, and then execute step by step the code and inspect the variable to try and understand what this is doing.
Good luck!