Hi Brainstorm experts!
I am currently trying to compute the head model from the FEM mesh generated by SimNIBS with DTI tensor calculated. My data is 256 channel EEG, I compute the head model with the template grid(head), it takes literally around ~ 72 hours per one subject on my Mac, which is too long for me. Therefore, I am going to compute it on a cluster that has a couple of nodes with multiple cores each. I've got some questions as follows:
- Previously I did the batch process for DTI and mesh generation in a
For loopexecuting GUI generated script in every iteration. I did the same for the head model this time, the code is attached also. I noticed that the subject name is not passed to the process, but the channel file. It seems the further process will find corresponding FEM mesh and other relevent files. I just wanted to confirm with you that passing the channel file path to the loop is correct, right?
% Script generated by Brainstorm (27-Apr-2021)
% Input files
sFiles = [];
% Start a new report
bst_report('Start', sFiles);
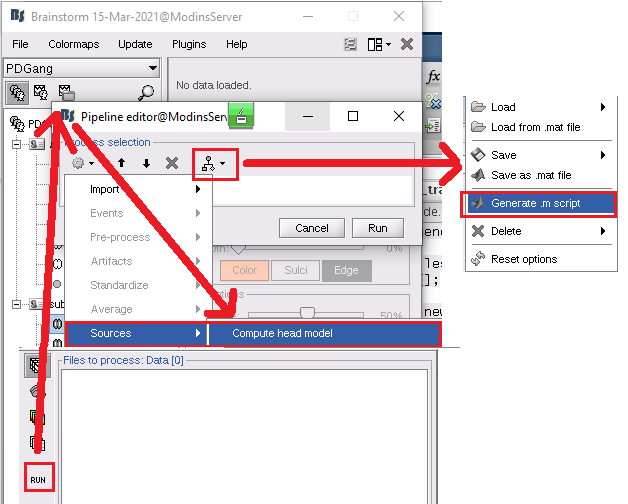
% Process: Compute head model
sFiles = bst_process('CallProcess', 'process_headmodel', sFiles, [], ...
'sourcespace', 2, ... % MRI volume
'volumegrid', struct(...
'Method', 'group', ...
'nLayers', 17, ...
'Reduction', 3, ...
'nVerticesInit', 4000, ...
'Resolution', 0.005, ...
'FileName', 'Group_analysis/@default_study/headmodel_grid_210514_1233.mat'), ...
'meg', 1, ... %
'eeg', 4, ... % DUNEuro FEM
'ecog', 1, ... %
'seeg', 1, ... %
'duneuro', struct(...
'FemCond', [], ...
'FemSelect', [], ...
'UseTensor', 0, ...
'Isotropic', 1, ...
'SrcShrink', 0, ...
'SrcForceInGM', 0, ...
'FemType', 'fitted', ...
'SolverType', 'cg', ...
'GeometryAdapted', 0, ...
'Tolerance', 1e-08, ...
'ElecType', 'normal', ...
'MegIntorderadd', 0, ...
'MegType', 'physical', ...
'SolvSolverType', 'cg', ...
'SolvPrecond', 'amg', ...
'SolvSmootherType', 'ssor', ...
'SolvIntorderadd', 0, ...
'DgSmootherType', 'ssor', ...
'DgScheme', 'sipg', ...
'DgPenalty', 20, ...
'DgEdgeNormType', 'houston', ...
'DgWeights', 1, ...
'DgReduction', 1, ...
'SolPostProcess', 1, ...
'SolSubstractMean', 0, ...
'SolSolverReduction', 1e-10, ...
'SrcModel', 'venant', ...
'SrcIntorderadd', 0, ...
'SrcIntorderadd_lb', 2, ...
'SrcNbMoments', 3, ...
'SrcRefLen', 20, ...
'SrcWeightExp', 1, ...
'SrcRelaxFactor', 6, ...
'SrcMixedMoments', 1, ...
'SrcRestrict', 1, ...
'SrcInit', 'closest_vertex', ...
'BstSaveTransfer', 0, ...
'BstEegTransferFile', 'eeg_transfer.dat', ...
'BstMegTransferFile', 'meg_transfer.dat', ...
'BstEegLfFile', 'eeg_lf.dat', ...
'BstMegLfFile', 'meg_lf.dat', ...
'UseIntegrationPoint', 1, ...
'EnableCacheMemory', 0, ...
'MegPerBlockOfSensor', 0), ...
'channelfile', '');
% Save and display report
ReportFile = bst_report('Save', sFiles);
bst_report('Open', ReportFile);
% bst_report('Export', ReportFile, ExportDir);
-
I got a warning when starting computing headmodel:
WARNING: You are using AMG without SuperLU! Please consider installing SuperLU, or set the usesuperlu flag to false to suppress this warning.How much faster would it be if I installed SuperLU? Is there a tutorial from Brainstorm that guides me to set SuperLu for DUNE in this case? -
In the anatomy files, each subject has
tess_cortex_pial_highandtess_cortex_pial_low, shall I activate the higher one before computing headmodel for better result?