Dear community,
I have conducted fNIRS signal reconstruction on an individual surface (mid_15000v surface) and obtained an activation map using Nirstorm. Subsequently, I manually created a scout based on the fMRI map of the same individual. I am now seeking to determine the spatial relationship between the activation area (or activated vertices) and the scout. Specifically, I aim to identify
- How many vertices were activated?
- How many activated vertices are located within the scout and,
- if any activated vertices lie outside the scout, how to pinpoint the vertex with the highest activation level (peak value) and calculate the geodesic distance between this vertex and the scout.
Would it be advisable to utilize a script to accomplish these tasks? Hopefully, I clearly described my questions. Any guidance or assistance in this regard would be greatly appreciated.
Thank you for your support.
Best regards,
Adam
Hi Adam,
You can perform all what you need by combining some function on the Brainstrom interface together with some basic scripts.
Once you have the activation at the source level (i.e., at each vertex over time), you can find the index of the activated vertex and visualize it on the MRI viewer.
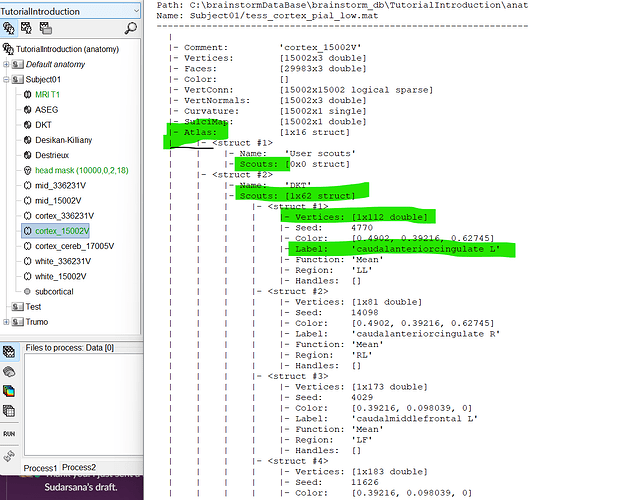
To get all the vertices of the scouts, go to the surface file of the cortex in the Anatomy tab. You can export it to MATLAB to extract the vertices belonging to each scout (see attached figure). All scouts are listed under the Atlas field.
- How many vertices were activated?
- How many activated vertices are located within the scout and,
you can either do it manually by drawing a scout around the activated area
Or you can export the result file, set a threshold, and then extract all the vertices that show a value higher than the threshold.
- if any activated vertices lie outside the scout, how to pinpoint the vertex with the highest activation level (peak value) and calculate the geodesic distance between this vertex and the scout.
To find the highest activation: display the cortical activation on the MRI viewer, press "M" to get the coordinates of that point, then compute the geodesic distance to any other point of interest.