Hi Team,
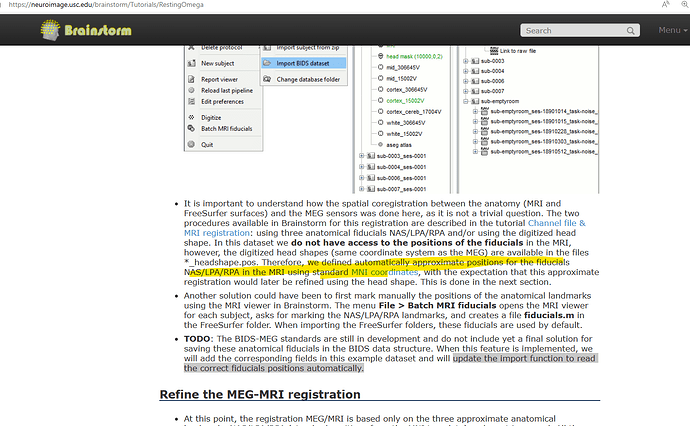
I am confused - if I should be editing the MRI Fiducials after importing the subjects. I am informed that the right procedure would be the same person who marked MRI fiducials should be editing it in Brainstorm as they know precisely where the fiducials were marked during recording.
In the tutorial; it's mentioned that the fiducials as seen are an approximation. So, should I edit the MRI Fiducials or just proceed with the default markings? The TODO enlists "the import function to read the correct fiducials positions automatically." - Is it implemented ?
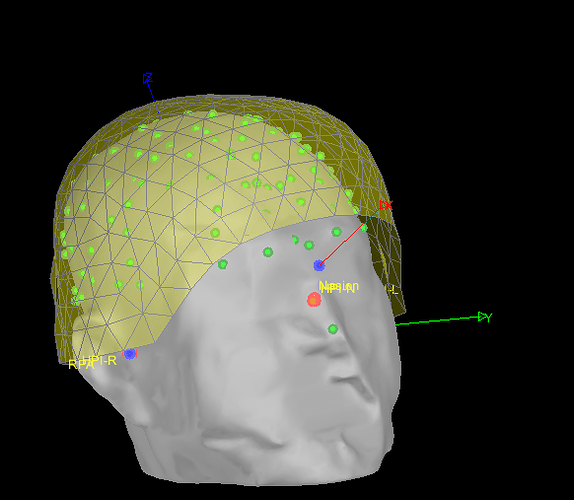
Also, which fiducials are the blue, red points representing? and the red/blue/green segments ?
Hello,
First, the comment about MEG-BIDS is slightly outdated. I'm not sure but I think there has been updates in Brainstorm to load fiducial coordinates if they are correctly specified in the BIDS files. This is something I will look into (confirm or implement) in the coming months after coregistration is done and saved in OMEGA. We should then update this tutorial's data, which does not yet have the coregistration saved, even if Brainstorm could read it.
As for what we recommend for coregistration in OMEGA in the meantime, it's as described here: use the approximate positions obtained from MNI registration first, then trust the automatic alignment with head points: "refine registration".
The blue dots are the fiducials marked (possibly automatically) on the MRI image, which are the ones used for defining the coordinate system. The green points are digitized "head points", and the "red" one the digitized fiducials (with labels). Note the inside of the red circles are yellow or orange for the two sets of digitized fiducials: head coils and anatomical.
Thanks Marc for the information.
Can you let me know when the feature for Brainstorm to able to load fiducial coordinates from the BIDS file if implemented/[planned to be?
Yes - For now, I will load the fiducials and won't edit them; refine registration and proceed for further analysis.
Thanks for the description of the different circles. I guess, the 3 protruding segments are X,Y,Z Axis as per MRI Co-ordinates. Is that right ?
Also, do the blue and red circles co-incidence mark a good co-registration?
As I said, the feature to read the coregistration may already be in Brainstorm, but I'll work on it (confirm or implement if not) in the coming months. But the OMEGA data doesn't have the coordinates saved, so nothing right now for Brainstorm to load.
Yes the segments are the axes of the Brainstorm "subject coordinate system" (SCS), defined from the MRI anatomical fiducials.
Your last question is more subtle. Before doing "refine with head points", the alignment between MRI and digitized points is based on both sets of anatomical fiducials (MRI and digitized), so they'll practically always line up well. But that doesn't mean the coregistration is any good if the MRI fiducials were only roughly placed (e.g. automatically). After refining with head points, the alignment is instead based on the head points and the scalp surface extracted from the MRI. At that stage, one can evaluate roughly the quality of the coregistration on where the digitized fiducials line up on the surface. Again, the blue MRI points are the same as before and we don't expect them to line up well with the now hopefully well aligned red digitized points. You can also look at where the digitized head coils are, but without pictures or detailed knowledge of where they were placed, it's not too informative.
Thank you so much for your answers Marc!
I understand the co-registration process much better now ! 
Hello!
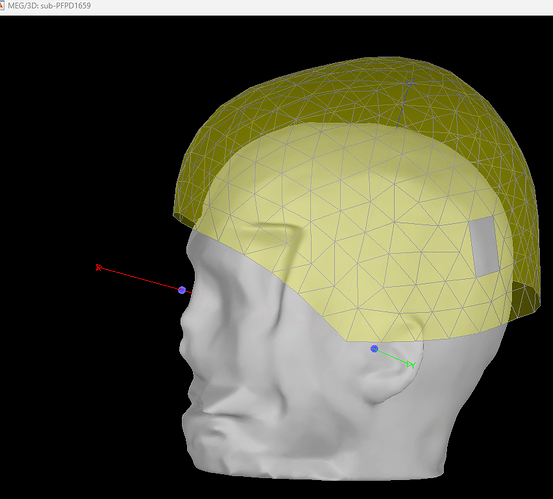
I observe that sub-PD1659 from OMEGA Dataset doesn't have the digitised head points. My goal is to analyse the reconstructed source time series using MRI Volume Head model. I am bit confused if this particular subject, the reconstructed source time series would be reliable or appropriate for further analysis. Can you help me with your advice ?
Thanks
@Sylvain @Raymundo.Cassani @Marc.Lalancette - can you help me with the above problem ?
I'll have to dig to find why there are no head points for this subject.
In the meantime, if there are no head points, you would need to mark the positions of the head coils on the MRI to align anatomy with MEG. Since the shared MRI is defaced and you don't have pictures of where those were placed, you can only guess and the alignment would be poor.
The blue points that show up are those that you, or the automated MNI registration, placed on the MRI. They do not come from the data. Note that for these subjects, the "nasion" MEG coil is usually placed on the forehead above the brow, and not where it appears in this figure. I would therefore suggest you may want to exclude this participant for now if precise localization is important.
Note that a student has been working on improving and validating coregistration in OMEGA, using original MRI images and pictures. This should be shared by the summer hopefully.
Thank you for the response Marc! I shall exclude this subject for time being.