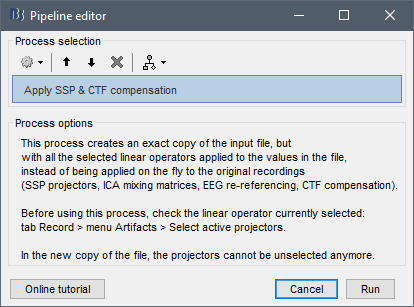
After applying the process_ssp_apply process, if you visualize the fif file and select Artifacts->Select active projectors, none of the projectors (autossp) have been applied. Specifically:
Also, process_ssp_apply generates a new link to raw file with extension '_clean', which seems unnecessary.
Step1: Review raw file.
Step 2: Run:
% Process: Apply SSP & CTF compensation
sFiles = bst_process('CallProcess', 'process_ssp_apply', sFiles, []);
Step 3: Artifacts->Select active projectors, this is still set to none.
I think I remember Matti explained at some point that the "autossp" projectors are meant to be used for visualization only, but should not be used for processing the MEG data.
This is the reason why they are disabled by default.
If you want to use them:
- Menu Artifacts > Select active projectors
- Select the projectors you want to use
- Then optionally call the process "Apply SSP" (but I'm not sure why you would do this).
In case I misunderstood your request, could you please post some screen captures illustrating the issue?
Hi Francois,
In general you are right, the autossp projectors are not used because we typically use Maxfilter instead (which also deletes the autossp projectors, so they cannot be accidentally applied). Of course this is with Megin systems, I do not know what happens with CTF.
I guess my point is that this is an unexpected behavior of the apply ssp process. If the projectors exist, one would expect them to be applied. It is also a discrepancy between the GUI and the scripting interface; applying the autossp projectors is possible with the GUI but not with scripts. [But this also generalizes to selecting individual projectors, etc]
I am also not sure why apply SSP projectors is together with apply CTF compensation, but I am not too familiar with the CTF machines to have an informed opinion about it.
I guess this is food for thought rather than change something at this point.
Cheers,
Dimitrios
There might some confusion around the purpose of the process "Apply SSP
- When linking the .fif files to the database you get the message below:
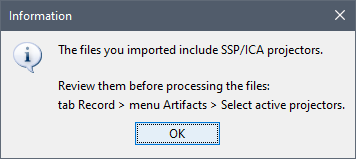
- We agreed a long time ago we would not use them by default. In case you want to use them on your data, you have to explicitly select them: menu Artifacts > Select active projectors, as the dialog box says.
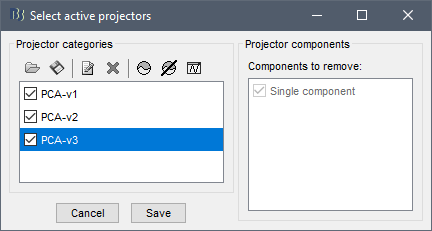
- The linear operations available for the data (SSP projectors, ICA mixing matrices, CTF 3rd-order gradient compensation matrices, EEG re-referencing matrices) are saved on the side, in the
Projector field of the channel file. These operators can be selected or unselected while reviewing the continuous files, and are applied dynamically to the MEG/EEG recordings when needed (ie. when importing, displaying or processing the files), but the original recordings are not altered.
- Applying these operators on the fly to the MEG/EEG recordings for one time sample require all the channels to be loaded simultaneously in memory. When running pre-processing functions that require reading all the time samples at once (eg frequency filtering), having linear operators (SSP, ICA, ...) on top of the file requires reading all the file at once (all channels x all time samples), and this might not fit in memory. In that case of memory limitations, you can call the process "Apply SSP & CTF compensation" to create a hard copy of the continuous recordings with all the linear operators apply to the recordings, so that later you can apply a frequency filter channel-by-channel, without having to load all the channels at once in memory. Another use case of this process is to run a feature detection on a single channel after cleaning with SSP/ICA. But I'm not sure I understood why you would need to use it in your pipeline.
- If you select the autossp projectors and then called the process "Apply SSP & CTF compensation", the only you get in output is a continuous with the same data, but now the projectors are applied hard to the data and can't be unselected:
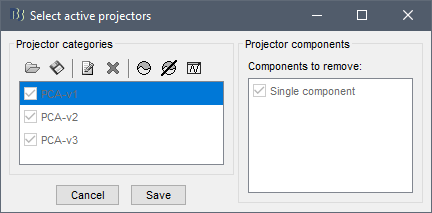
If you select the process "Apply SSP" and click on the Help button in the Pipeline editor window, it should open a browser pointing at this tutorial section, which introduces the logic of the process:
https://neuroimage.usc.edu/brainstorm/Tutorials/TutMindNeuromag?highlight=(Apply+SSP)#Remove:_60Hz_and_harmonics
For clarity, I added some in-line documentation to this process: