Hello everyone,
I am trying to compute sources of scalp EEG (27 electrodes) using sLORETA and I encounter some unexpected results in BST which makes me question if I am doing something fundamentally wrong.
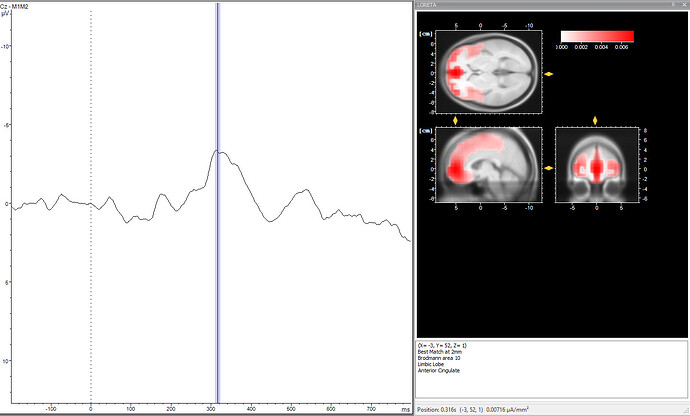
I have tried sLORETA on an EEG file that is a grand average (n=12) of an ERP-component (RewP, occurs 250-350ms after feedback). The file I have imported was already completely pre-processed (so imported as a 27x256 file, 27 channels and sampling rate of 256 Hz). I did the pre-processing in BrainVision Analyzer, which also allows LORETA. This is their result (depicted at channel Cz):
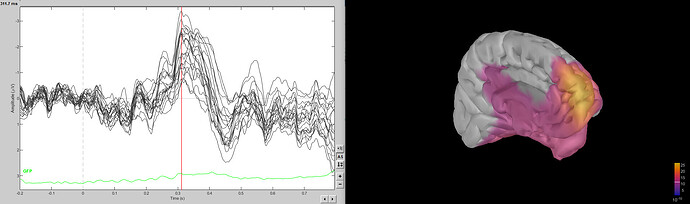
So this designates the ACC as the source of this ERP component (which is in line with our hypothesis). However, when I try to replicate this results in BST, I get the following (sLORETA, unconstrained because I used a template):
It doesn't point out the ACC, but more a frontopolar source.
This made me question if I was doing something wrong. Could you help me with the following questions:
- Does the reference of the imported GA file matter? Right now it is referenced to the linked mastoids, not the average reference.
- I have computed the noise covariance matrix from the recording, but after reviewing the tutorial https://neuroimage.usc.edu/brainstorm/Tutorials/NoiseCovariance?highlight=(covariance) this does not seem like a good idea. Should I select the option "no noise modelling" in this case?
- Are there any other steps that I am missing to correctly perform the sLORETA?
Additionally, if I want to compute the source by averaging the results of the source localization of the individual subjects together, do I need to do some kind of standardisation before I can average them together?
Many thanks for your help.
Hello Sylvain,
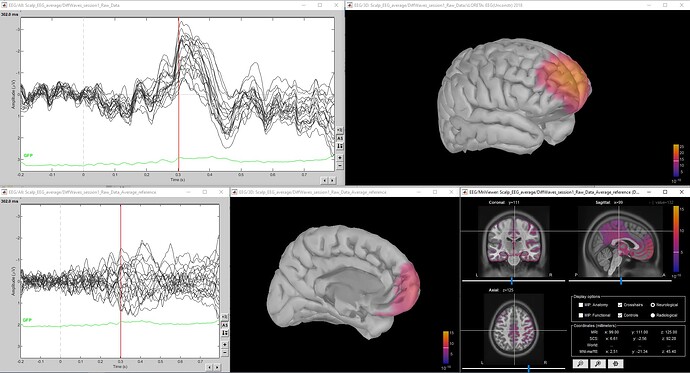
Thank you for your suggestions. I have re-referenced the data by running the process "standardize --> apply average reference" and used the option "no noise modelling" to compute the sLORETA (unconstrained), but the results don't look better (top is original data, bottom is re-referenced to average reference).
Also, according to this post BST automatically re-references your data to the AVG when computing the head model, so I am a bit confused.
Also, when I tried to do source localization for similar data for another group of participants, corpus callosum is activated instead of ACC. Since this is not a cortical structure, this is definitely surprising to me.
Could you thinks of any other reasons why the source localization does not seem to work on my data?
Best,
Joyce
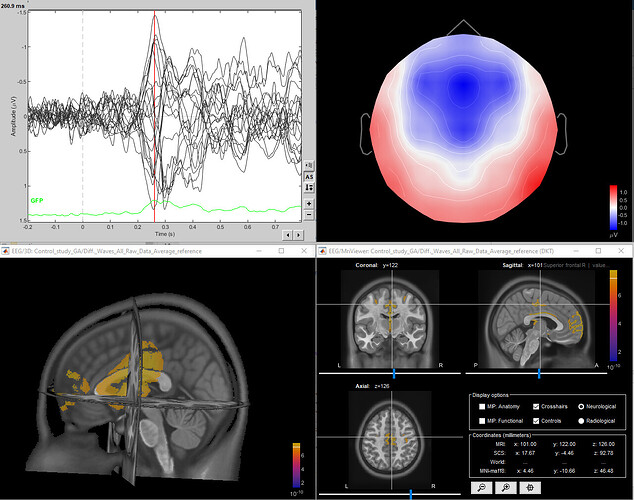
I just wanted to add that this is the result of one of my source localizations:
To me, the shape looks weird and it is strange that the corpus callosum is also highlighted.
Thank you, Joyce and sorry for putting you on a wrong irrelevant track re: re-referencing.
I am not sure what the issue is with the second cohort. The first batch of results are close, I think, of the original sLORETA data you saw in publications. You can try another source mapping technique, for confirmation. I recommend: Mini-norm, followed by Abs values and three z-scoring based on pre-stim baseline.
1 Like
Hi @joerlema
Adding to @Sylvain comments, the difference can also be due to how you computed the forward head.
Do you know what method is used in BrainVision and what are the values of the conductivity?
Were you able to check the location of the imported electrodes on the head surface and the source space?
The difference in these parameters can lead to differences in the final results.
So to be clear: I do not need to re-reference to the average explicitly before doing source localization?
And what kind of mini-norm source localization do you recommend? CDM or sLORETA? And if I choose sLORETA, is z-scoring then still necessary because I thought that standardisation was already done in this method?
I don't know what is used in BrainVision, just that it is a LORETA approach. I can try to find out more information.
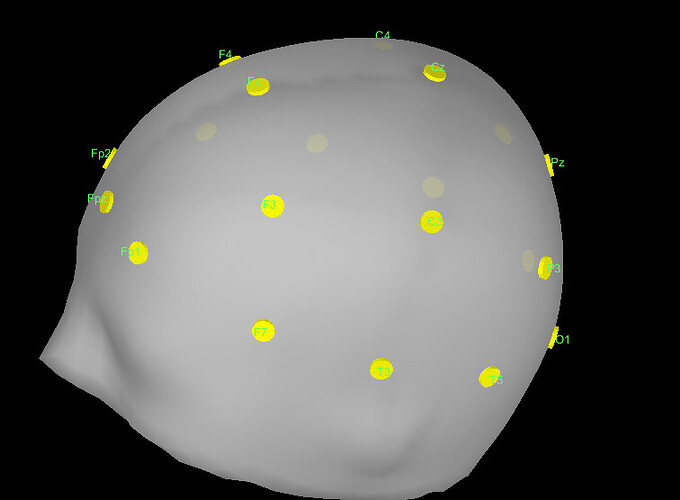
I checked the locations of the electrodes and they seem to be correct:
Any ideas as to why the corpus callosum pops up in my source localization results?
Thanks!
1 Like