Hello!
I am trying to calculate sources of ERP data but the results seem to be abnormal. Here are the procedures I followed:
- Right click “CNT 2D channels” – compute head model – OpenMEEG BEM (cortex surface)
- Copy the OpenMEEG BEM file to every subject
- Drag preprocessed files to process 1 – source – compute covariance (noise covariance, no noise modeling, merge).
- Drag average files (baseline corrected) to process 1 – source – compute source (minimum norm imaging, sLoreta, constrained)
- Perform t-test between two different conditions.
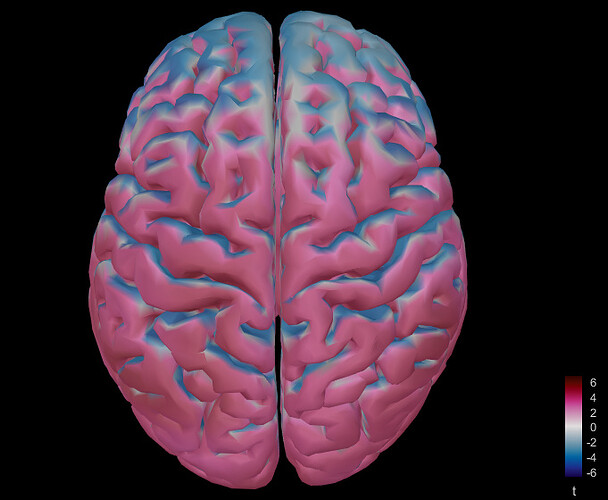
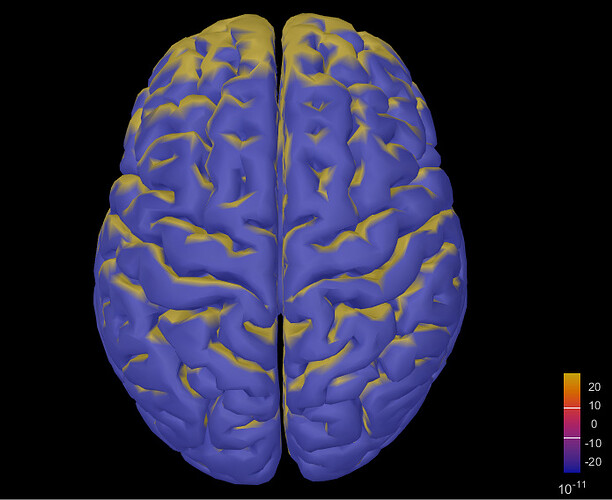
However, the t-test result was like this:
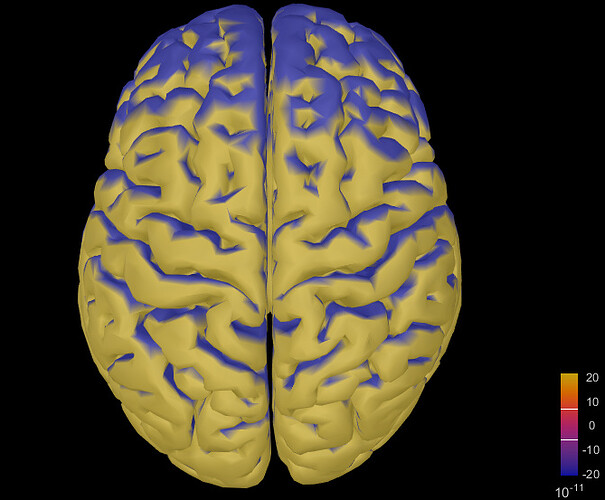
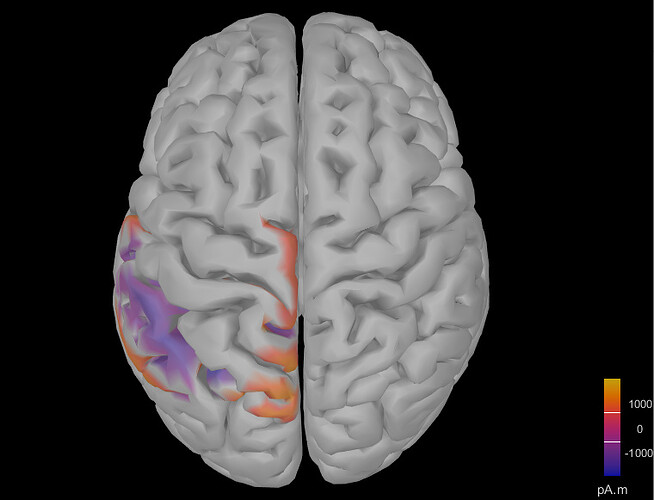
When I opened the source file for a single participant, it looked like this:
Although the activation changed as I moved through different time windows in the ERP time series figure, it was always like the activation was spreading all over the cortex. Any ideas what is wrong here? Thank you very much!
This "CNT 2D channels" is very suspicious: it looks like you don't have 3D positions for your sensors.
Please post a screen capture showing the electrodes aligned on the head surface, as in this tutorial:
https://neuroimage.usc.edu/brainstorm/Tutorials/Epilepsy#Prepare_the_channel_file
If you don't have electrodes positions, you can use templates.
Right-click on the channel file > Add EEG positions > ICBM152 > ...
If you do have defined correctly the positions of the electodres, please post screen captures of your ERP and the noise covariance matrix image.
Hello Francois,
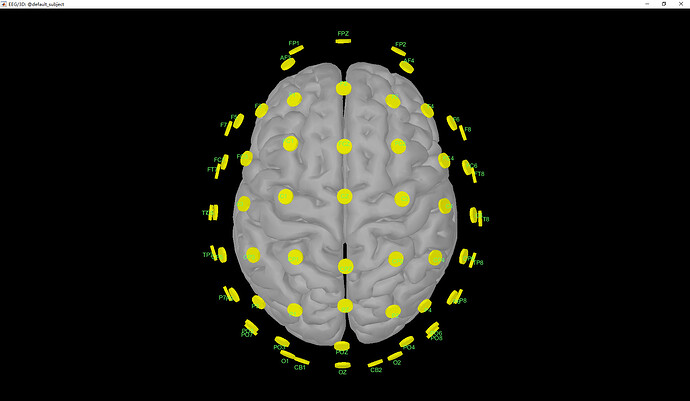
Here is the screen capture showing the electrodes aligned on the head surface:
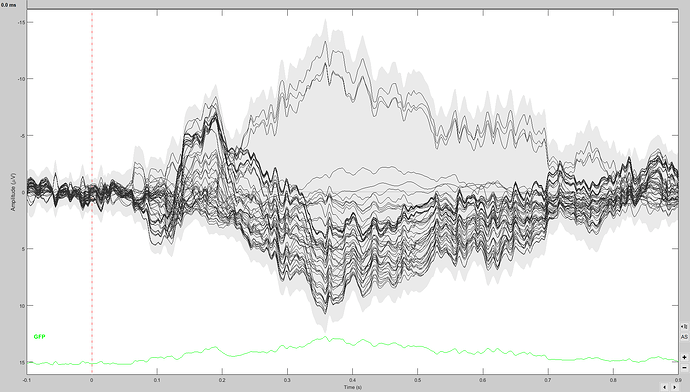
Here is the screen capture of the ERP waveforms of one of the participants:
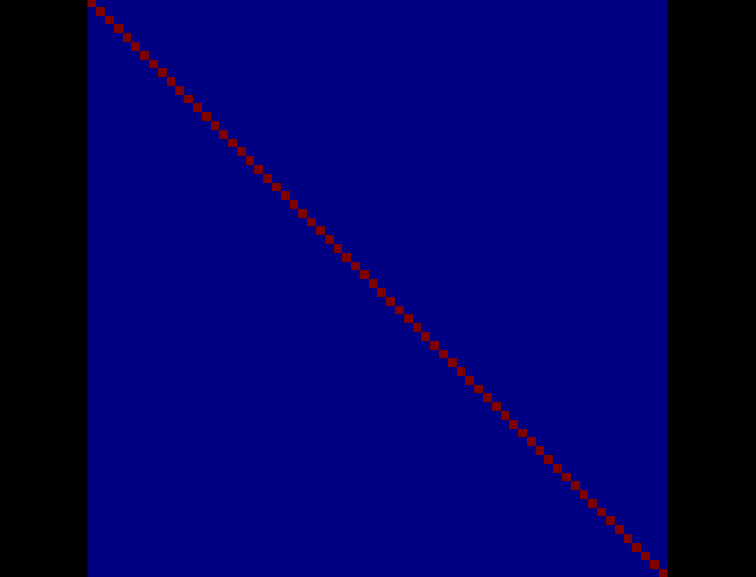
Here is the screen capture of the noise covariance matrix image:
This looks OK to me.
Have you tried with other forward models (a 3-shell spherical model instead of OpenMEEG)?
@John_Mosher @Sylvain What could be the problem here?
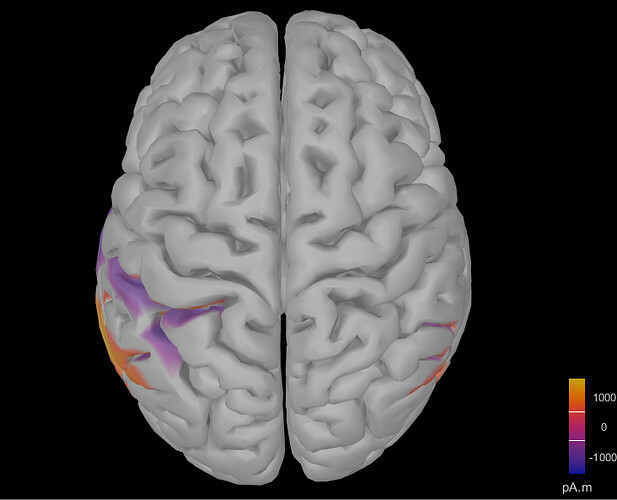
I tried to use the 3-shell spherical model but the results were similar:
What latency are we looking at ?
Place the time cursor around 0.1s or .2s where there are peak in your data.
Also, please make sure the EEG reference information is correct.
Hello Sylvain,
I tried to move the time cursor but the activation is always spreading all over the cortex. I used M1 and M2 as the EEG reference and I have marked them as EEG REF in channel selections.
I can recommend that you re-reference your recordings to "Average Reference": there is a process in brainstorm to that purpose. You can then compute the source map again. This may help. The other thing would be to display a few EEG scalp topographies before moving to source space: what do they look like?
I can recommend that you re-reference your recordings to "Average Reference": there is a process in brainstorm to that purpose. You can then compute the source map again.
The sources are always estimated in average reference, so it shouldn't change anything.
Here are a couple of EEG scalp topographies:
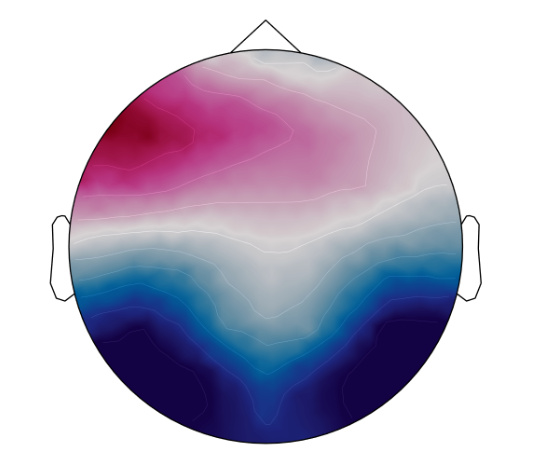
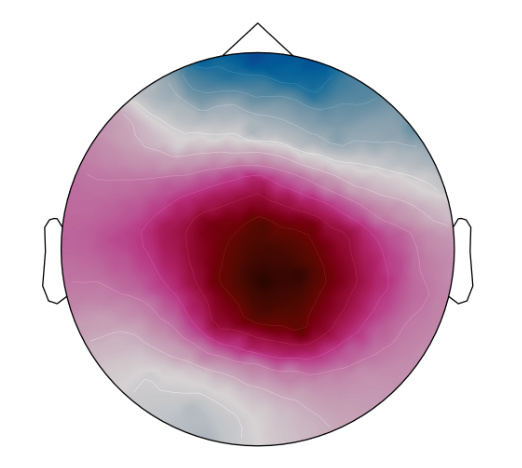
I think the EEG waveforms and topographies are fine. Just the source seems to be confusing...
Thanks. That's right. When you visualize the EEG scalp topographies, do you activate the EEG AVG REF button in the main display panel? If yes, please deactivate it and capture a couple more screenshots.
Hello Sylvain,
Do you mean the "Average reference" button in the Montage menu when I right click on the topography? It has always been deactivated and "All channels" has been selected.
OK great. You may want to re-reference your EEG data via the process Standardize > Re-Reference EEG > then type in "AVERAGE" in the reference text box as explained in the panel.
I also notice that you have 3 channels than seem off from the rest in the time series plot above. The variance around these time series is very large, which may also be indicative that these channels are noisy and could be declared as "bad". But please try to re-reference first. thanks.
Hello Sylvain,
I tried with the average reference. Now the figure looks normal:
However, when I use M1 and M2 as reference and follow the same procedure, the results are still problematic as before. In the channel file I have marked M1 and M2 as EEG REF, I am not sure if it has something to do with the abnormality in source estimation.
Great!
Modeling linked mastoids as EEG reference is not an entirely trivial issue, which we have not looked at in brainstorm. We recommend you change your data to Average Reference following the process steps you just ran.
I think I've figured out the reason. It's due to M1 and M2 being marked as EEG REF. I changed them back to EEG channels and now the sources are normal. Thank you very much for your help and patience!
1 Like
This is great - glad it worked out!
Happy brainstorming!
Indeed, all the the EEG channels, including the references, should have their type set to "EEG".
Otherwise, the forward model is not estimated for the references, and the average reference is not properly applied.
In general, the type "EEG REF" should not been used, except for some file formats that have the reference information duplicated (in that case, it should be already defined as "EEG REF" automatically when loading the file in the Brainstorm database).
@gubei2008 Can you please confirm that after setting the type of M1 and M2 to "EEG", you don't need to apply the average reference explicitly before estimating the sources?
(so that future readers of this thread do not try to duplicate their data if they don't need to)
Thanks
Hello Francois,
Yes, applying the average reference is not necessary. Just changing M1 and M2 back to "EEG" will work.