You could try the following:
- Create a subject with NO default anatomy
- Right-click on the subject > Use template > MNI > ICBM152
- Run CAT12 on the ICBM152 MRI:
https://neuroimage.usc.edu/brainstorm/Tutorials/SegCAT12 - Use one of the output atlases that include the regions you are interested in (e.g. AAL3):
https://neuroimage.usc.edu/brainstorm/Tutorials/SegCAT12#Volume_parcellations - For using as scouts:
https://neuroimage.usc.edu/brainstorm/Tutorials/TutVolSource#Volume_atlases - For obtaining surfaces: Creating surfaces is not possible directly, but you can do the following:
- Right-click on the volume parcellation of interest > Export to file > .nii
- Right-click on the subject > Import surfaces > Select exported .nii with file format: Volume mask or atlas (subject space)
- Then manage this just like the ASEG surface in the FreeSurfer output (extract only the regions of interest and merge them with the cortex):
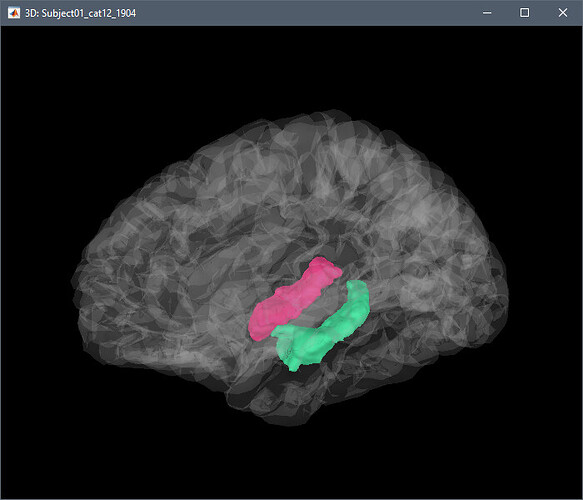
Example obtained for the hippocampus: