Tutorial 23: Scouts
Authors: Francois Tadel, Elizabeth Bock, Raymundo Cassani, John C Mosher, Richard Leahy, Sylvain Baillet
In Brainstorm jargon, a scout represents a region of interest (ROI) in the available source space. It is a subset of dipoles defined on the cortex surface or the head volume. This tutorial explains how to create one or several scouts, use them to represent the activity in specific brain regions and compare the responses between different experimental conditions.
Here, all the steps are done from the scout panel, but scout time series can also be saved with the process "Extract > Scout time series" for further analysis, or computed directly within various processes, e.g. many under Frequency or Connectivity.
Contents
Hypothesis
For all the brain imaging experiments, it is highly recommended to have a clear hypothesis to test before starting the analysis of the recordings. With this auditory oddball experiment, we would like to explore the temporal dynamics of the auditory network, the deviant detection and the motor response. According to the literature, we expect to observe at least the following effects:
Bilateral response in the primary auditory cortex (P50, N100), in both experimental conditions (standard and deviant beeps).
Bilateral activity in the inferior frontal gyrus and the auditory cortex corresponding to the detection of an abnormality (latency: 150-250ms) for the deviant beeps only.
Decision making and motor preparation, for the deviant beeps only (after 300ms).
We will start by creating regions of interest corresponding to the auditory cortices to illustrate the tools, then define other regions to better explore the dynamics of the brain response.
Creating a scout
Almost all the features related to scout manipulation are accessible in the Scout tab in the main Brainstorm window. The scouts are automatically saved in the surface file from which they are created, and they are loaded and automatically displayed each time the surface is loaded.
An atlas designates, in this context, a list of scouts. For one cortex surface, we can have as many atlases as needed. An atlas can be an anatomical parcellation (like the ones loaded when using FreeSurfer), a random parcellation generated by Brainstorm, or a user-defined list of ROIs. All the surfaces contain, by default, an empty atlas called "User scouts", for the user to create new regions of interest.
First vertex (seed)
In Intra-subject, double-click on the normalized standard average.
Open the average recordings for the standard condition, to have a time reference.
Go to the first peak in the response, around 90ms.
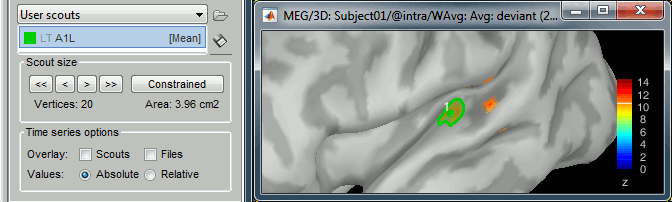
In the Surface tab, increase the amplitude threshold to see something relatively focal. The response is larger in the left hemisphere, so let's start with the left.
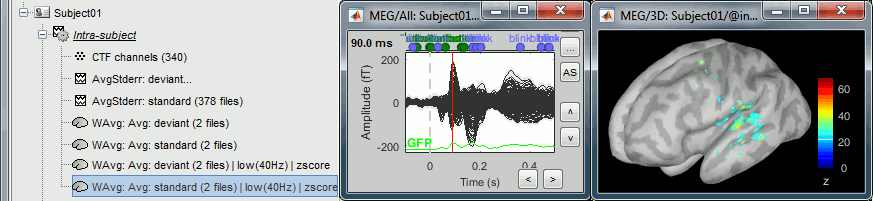
Switch to the Scout tab, click on the first button in the toolbar (the big cross).
In the 3D figure, click where we expect to find the primary auditory cortex (rotate and zoom before if necessary). A small green dot with the label "1" appears where you clicked. Your first scout is created in the User scouts atlas, and contains only one vertex for the moment.
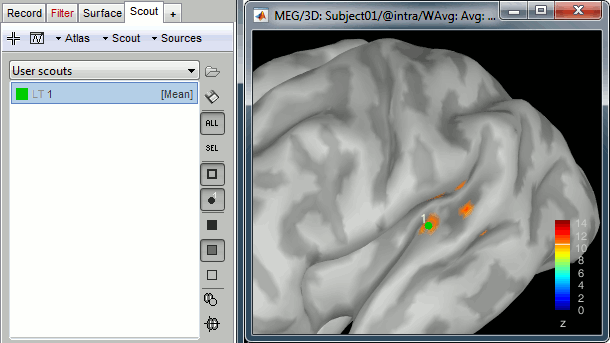
If you are not satisfied with the position of the vertex you selected, delete the new scout (select it in the list and press the Delete key, or menu Scout > Delete) and try again.
Rename it "A1L", for primary auditory left: double-click on it in the list, or menu Scout > Rename.
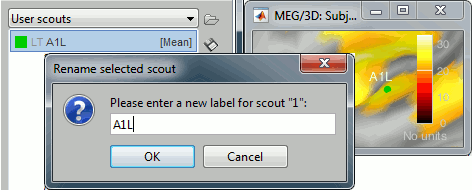
In light grey, you can see in the list the letters "LT", this means that based on the anatomical atlases imported from FreeSurfer, the point you clicked is in the left temporal lobe.
Growing a scout
For now, our scout contains only one vertex of the cortex surface. Most of the time, the aim of a scout is to extract the average activity over a larger region. The buttons in the Scout size section offer basic operations to define the scout extension.
[>]: Add the closest vertex (with respect to the seed) to the current scout.
[<]: Remove the furthest vertex (with respect to the seed) from the current scout.
[>>]: Grow scout in all directions.
[<<]: Shrink scout in all directions.
'Constrained': If this button is pressed, only the vertices that have a source value above the threshold will be added to the scout (its growth will be limited to the colored patch on the surface).
Add vertex manually: Select the Select point button, then select again the "A1L" scout. When you click on a vertex on the surface, it is added to the selected scout.
Remove vertex manually: Same as previously, but holding the SHIFT key at the same time.
Grow the scout A1L to 20 vertices, not in constrained mode. You can read the number of vertices and the estimated cortical area just below the [<<] and [>>] buttons.
Display time series
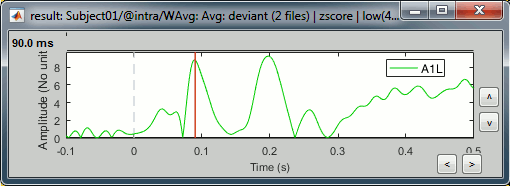
Select the scout in the list, click on the second button in the toolbar [Display scouts time series]. It displays the signal associated with this region for the entire time window (-100ms to 500ms).
We can now observe the early response at 50ms (P50) that was not very obvious before.
3D display options
In the toolbar on the right side of the scouts list, you can find a list of display options. Leave your mouse over each button for a few seconds to get a short description.
 Load Atlas
Load Atlas  Save selected scouts
Save selected scouts  Show all the scouts
Show all the scouts  Show only the selected scouts
Show only the selected scouts Hide all: Uncheck both buttons above to hide the scouts in the 3D figure.
 Show / hide the contour line
Show / hide the contour line  Show / hide the scouts labels
Show / hide the scouts labels  Scout patch: Opaque
Scout patch: Opaque  Scout patch: Transparent
Scout patch: Transparent  Scout patch: None
Scout patch: None  Display the color of the region instead of the scout color (only for anatomical atlases)
Display the color of the region instead of the scout color (only for anatomical atlases) 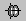 Center MRI on scout (open the MRI viewer and show the position of the scout's seed)
Center MRI on scout (open the MRI viewer and show the position of the scout's seed)
Scout function
We have extended the scout A1L to 20 vertices. Because this is a source model with constrained dipoles orientations, we have one source only per vertex. The region A1L corresponds to 20 signals.
The signals are grouped together into one unique signal that is then used to represent the activity of the region of interest. In the list of available scouts, you can see the indication [Mean] next to the name of the scout. It represents the name of the function that is used for combining all the source signals into one. This function can be changed individually for each scout, with the menu Scout > Set function.
Here is a description of the different options. In the case of unconstrained sources (3 signals for each vertex, one for each orientation), the function is applied separately for each orientation and produces 3 time series instead of one. For more details, see the code of bst_scout_value.m.
Mean: Average all the signals.
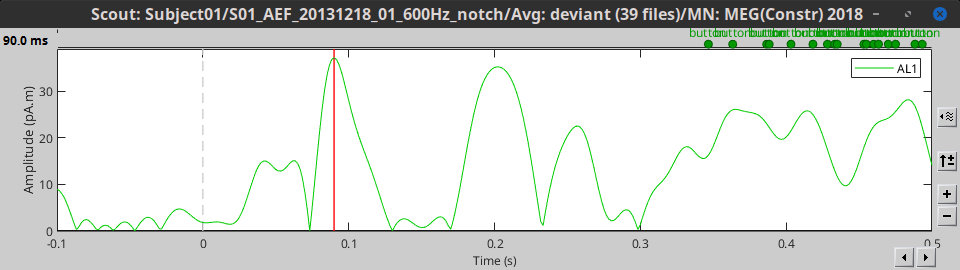
PCA: Take the first mode of the Principal Component Analysis decomposition of all the signals. Here, this is computed on a single file for display, but PCA works best when all epochs and conditions are processed together, through the process tabs. See PCA tutorial.
Fast PCA: Same as PCA function, but computes the first PCA decomposition based only on a subset of the strongest signals. It gives very similar results, but its computation is much faster for scouts that group a large number of dipoles (>50).
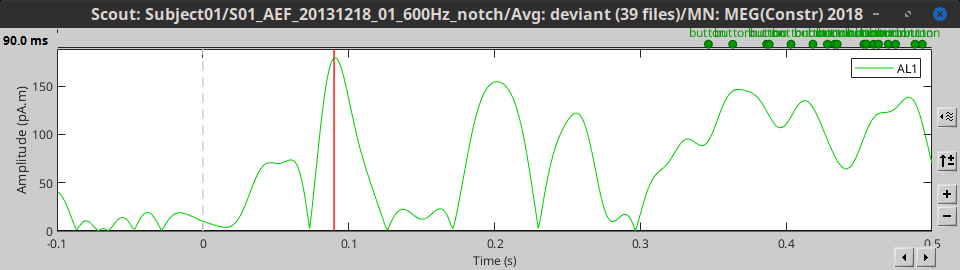
Mean(norm): Average the absolute values of all the signals: mean(abs(signal))
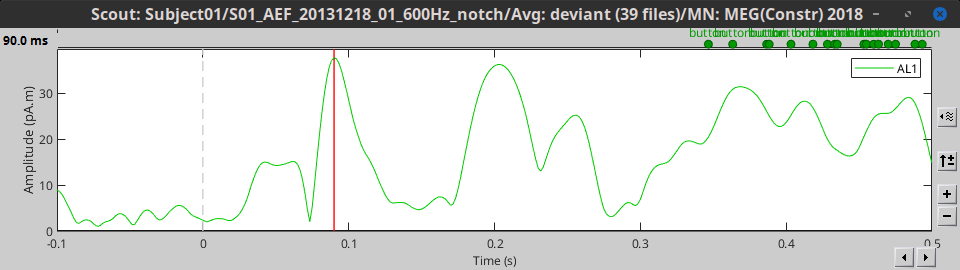
Max: For each time point, get the maximum across all the vertices.
Signed maximum: m = max(abs(signal)), scout = sign(signal(m)) * m
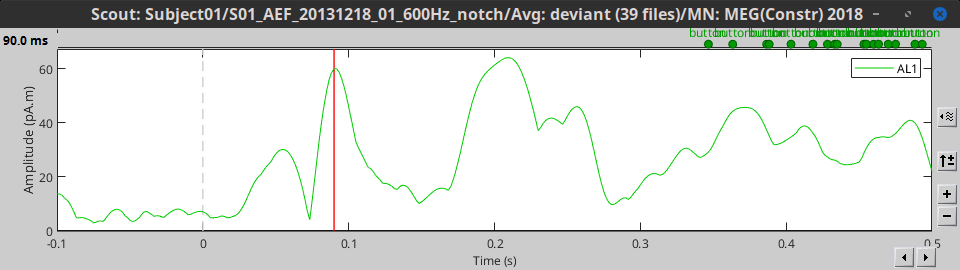
Power: Average the square of all the signals: mean(signal^2)
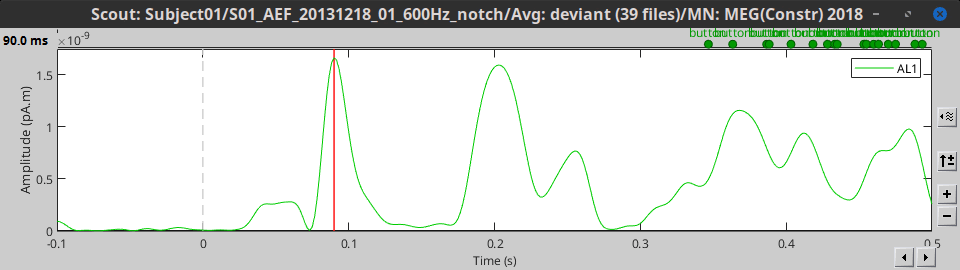
RMS: Square root of average the square of all the signals: sqrt(mean(signal^2))
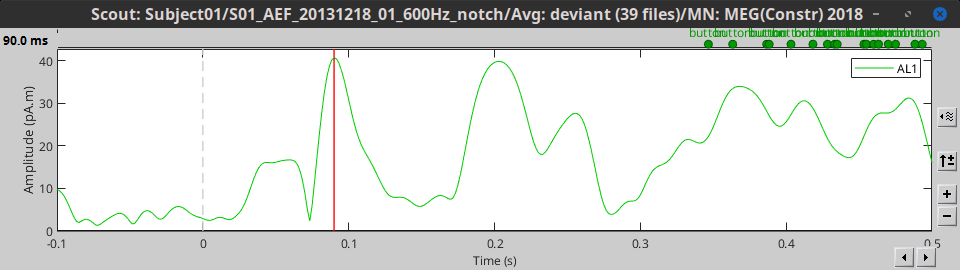
All: Do not apply any operation on the scouts signals, returns all the signals.
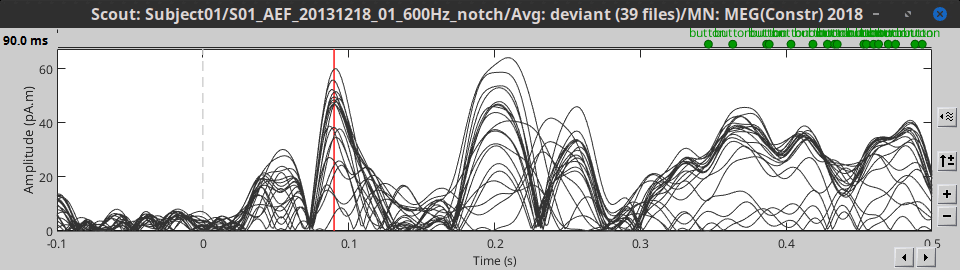
Note that the function selected here is ignored when running a process with a scout function option, like "Extract > Scout time series" or Connectivity.
Option: Absolute / relative
As said in the previous tutorial, the minimum norm current amplitudes can be positive or negative depending on the dipole orientation. This means that the values of the scouts, depending on the function that is used, may be positive or negative too. Most of the time we are interested in visualizing the absolute values of the scouts time series, to compare the activation level in different conditions or subjects. But it is sometimes easier to understand the temporal dynamic of a ROI with the relative values.
At the bottom of the Scout tab, you can choose to display either Absolute or Relative values.
The effect of this options changes whether you are processing source files with constrained (1 signal per vertex) or unconstrained (3 signals per vertex) dipoles orientations:
Constrained: Apply the scout function to all source signals, then:
Absolute: abs(ScoutFunc(sources))
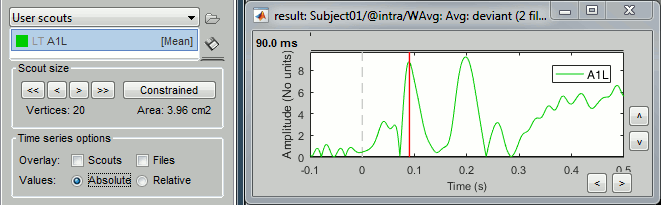
Relative: ScoutFunc(sources)
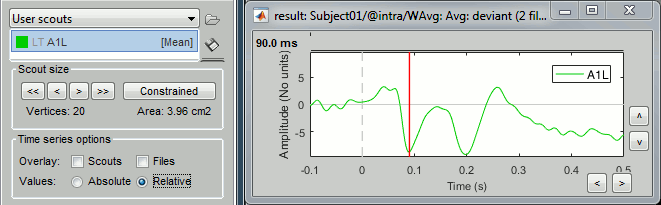
Unconstrained: Apply the scout function to the source signals for each orientation (Sx,Sy,Sz) separately, and then returns either the norm of the 3 orientations, or each orientation separately (except PCA, see below):
Absolute: Returns one time series:
sqrt( ScoutFunc(Sx)2 + ScoutFunc(Sy)2 + ScoutFunc(Sz)2)
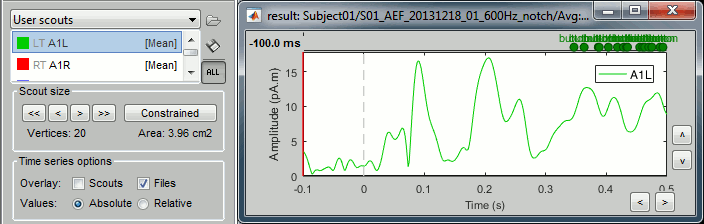
Relative: Returns three time series:
A1L1=ScoutFunc(Sx), A1L2=ScoutFunc(Sy), A1L3=ScoutFunc(Sz)
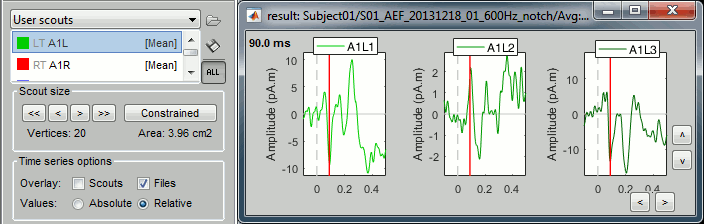
- Since PCA can also be used to combine the 3 orientations, it is applied here across all sources and orientations at once, producing a single time series, like in the constrained case above.
Display only: Note that this relative/absolute selection is a display option, it is not saved in the scouts themselves. It is used only when displaying the scouts time series with the [Display scouts time series] button of the Scout tab. In all other cases, such as the extraction of the scouts values from a script, this relative/absolute option is ignored. In those cases, there may also be an option to flatten unconstrained source orientations first, resulting in a single time series per scout instead of 3.
Multiple conditions
We can easily compare the activity between multiple conditions and multiple regions of interest.
In Intra-subject, open at the same time the normalized sources for deviant and standard condition.
Select the scout A1L and click on [Display scouts time series].
This computes the scouts time series for all the files that are currently open.
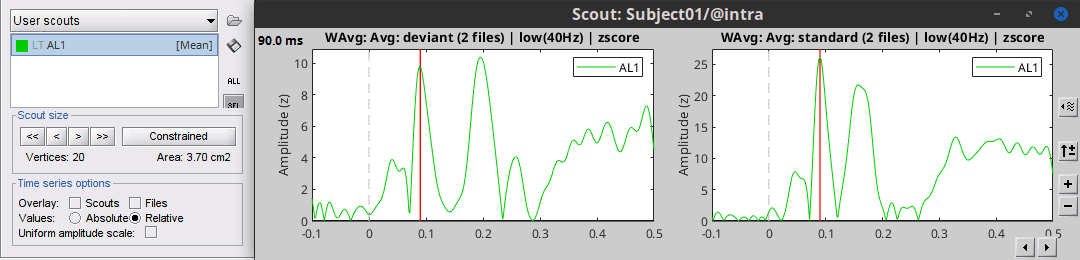
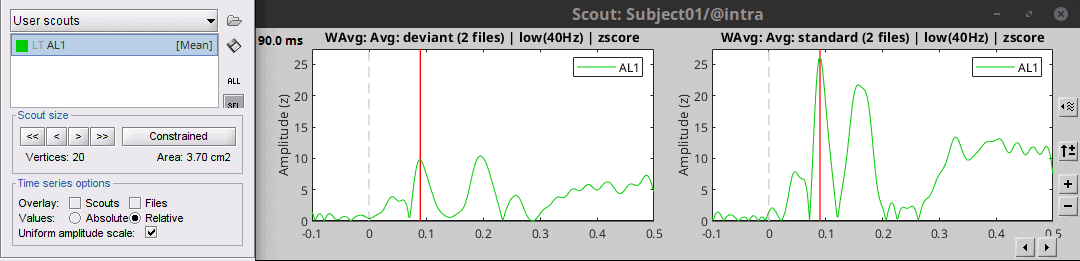
Use the checkbox Uniform amplitude scale to configure the amplitude of multiple axes.
Select it to force multiple graphs to use the same scale, unselect it to scale each graph separately.
At the bottom of the Scout tab, select the option "Overlay: Files", then click [Display] again.
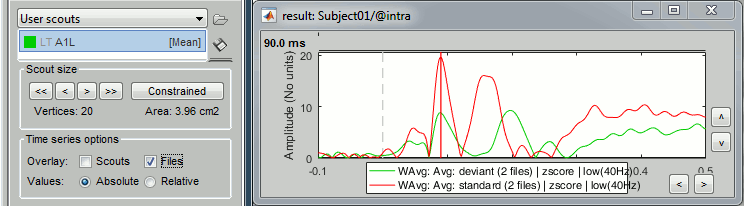
The Z-score value for the standard condition is a lot higher than the deviant condition because of the number of trials we used for computing the two averages (5x more trials for the standard). The SNR is higher with more trials, the baseline has less variance so the Z-score is higher.
To overlay the results of two conditions averaged with very different numbers of trials, it makes more sense to display the scout time series for non-normalized maps (averaged current density).
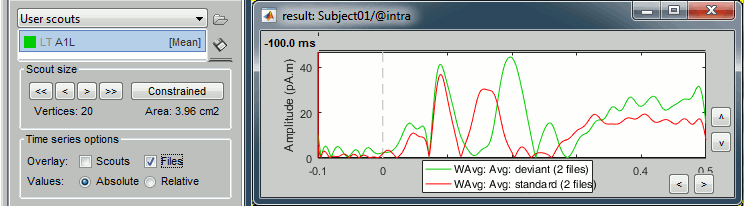
Other regions of interest
- Let's place all the regions of interest starting from the easiest to identify.
Open the normalized average source files (standard and deviant), together with the average recordings for the deviant and standard condition for Run#01 for an easier time navigation.
In the Surface tab, smooth the cortical surface at 70%.
- For each region: go to the indicated time point, adjust the amplitude threshold in the Surface tab, identify the area of interest, click on its center, grow the scout, rename it.
Grow all the regions to the same size: 20 vertices.
A1L: Left primary auditory cortex (Heschl gyrus) - Already marked.
- The most visible region in both conditions. Active during all the main steps of the auditory processing: P50, N100, MMN, P200, P300.
Standard condition, t=90ms, amplitude threshold=50%
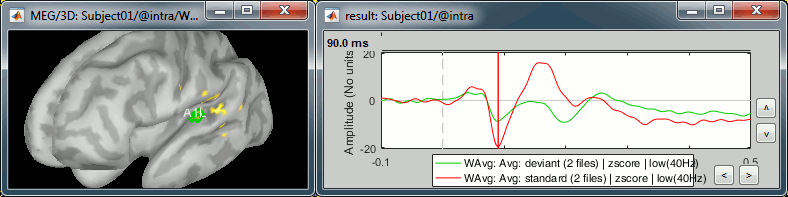
A1R: Right primary auditory cortex (Heschl gyrus)
- The response in this region is not as obvious than A1L. These binaural auditory stimulations should be generating similar responses in both left and right auditory cortices at early latencies. Possible explanations for this observation:
- The earplug was not adjusted on the right side and the sound was not well delivered.
- The subject's hearing from the right ear is impaired.
- The response is actually stronger in the left auditory cortex for this subject.
- The orientation of the source makes it more difficult for the MEG sensors to capture.
Deviant condition, t=90ms, amplitude threshold=50%
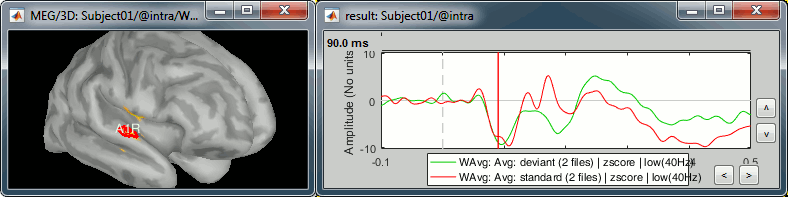
- The response in this region is not as obvious than A1L. These binaural auditory stimulations should be generating similar responses in both left and right auditory cortices at early latencies. Possible explanations for this observation:
IFGL: Left inferior frontal gyrus (Brodmann area 44)
- Involved in the auditory processing, particularly while processing irregularities.
- You can use the atlas "Brodmann-thresh" available in the Scout tab for identifying this region.
Deviant condition, t=130ms, amplitude threshold=30%
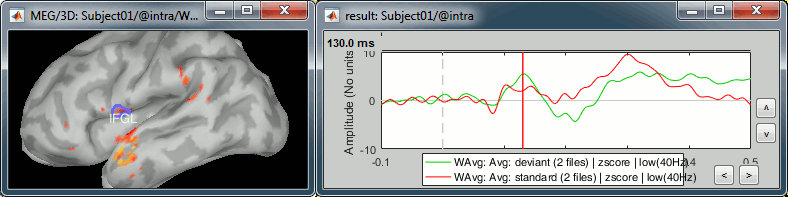
M1L: Left motor cortex
- The subject taps with the right index when a deviant is presented.
- The motor cortex responds at very early latencies together with the auditory cortex, in both conditions (50ms and 100ms). The subject is ready for a fast response to the task.
- At 170ms, the peak in the standard condition probably corresponds to an inhibition: the sound heard is not a deviant, there is no further motor processing required.
- At 230ms, the peak in the deviant condition is probably a motor preparation. At 350ms, the motor task begins, the subject moves the right hand (recorded reaction times 500ms +/- 108ms).
- We cannot expect to have clear responses during the motor response because of the averaging. The response times are variable, so in order to get a better representation of the regions involved we should import and average the trials based on the response triggers.
Deviant condition, t=440ms, amplitude threshold=25%
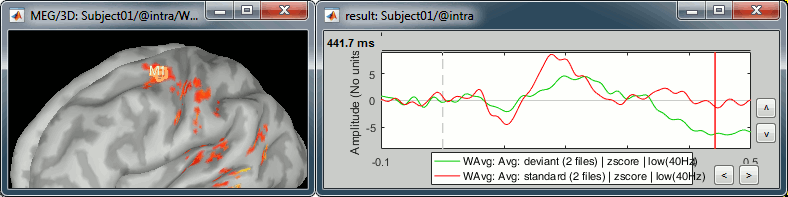
Multiple scouts
We can display the activity of multiple regions simultaneously.
- Close everything (toolbar button [X]).
Open the two normalized average source files (standard and deviant).
In the Scout tab, select the A1L and IFGL scouts simultaneously.
Select the option "Overlay:Scouts", do not select "Overlay:Files", Absolute values.
Click on [Display scouts time series].
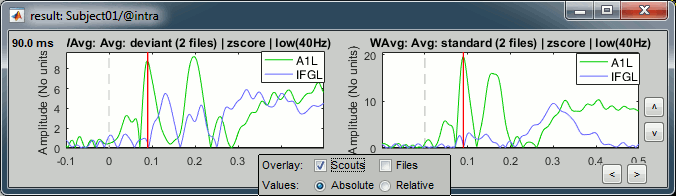
Now select the option "Overlay:Files", do not select "Overlay:Scouts", click on [Display].
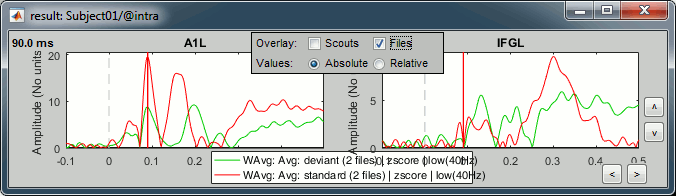
In both conditions, we observe similar early responses (P50 and N100), then it diverges.
In the deviant condition, we observe a pattern A1-IFG-A1 that is absent in the standard condition.
From the database explorer
You have to display the sources on the cortex to create the scouts. But once they are created, you can directly display the scouts time series from the database explorer. It means that you can quickly compare the values for a scout between many different conditions without having to open them all.
- Close everything (toolbar button [X]).
Select the option "Overlay:Files", do not select "Overlay:Scouts".
Select the two average recordings in Run#01 folder > right-click > Scouts time series.
Note that this menu is present at all the levels in the database explorer.
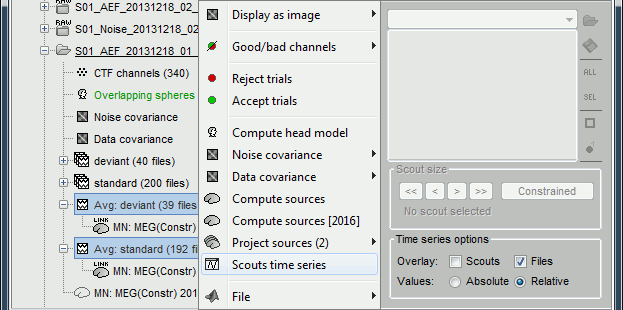
If no scout is currently loaded in the Scout tab, it shows all the scouts available in the surface.
If a surface is loaded, and at least one scout selected in the Scout tab, this popup menu would display only the selected scouts.
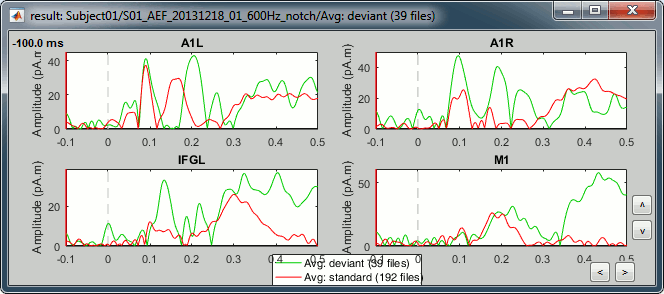
Once the list of scouts is loaded in the Scout tab, you can select one of them, and then display its values for all the trials of a single condition (overlay:files).
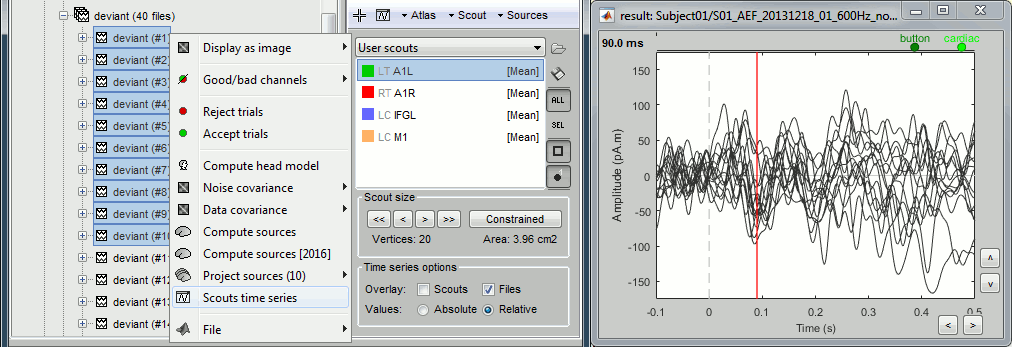
Sign flip
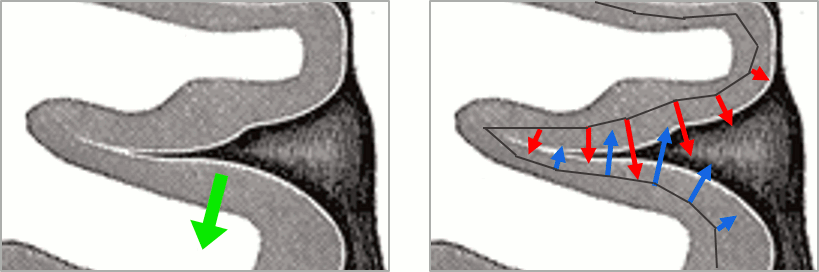
In the case of source models with constrained orientations (normal to the cortex), the sign of the current can be an issue. If the region is large enough to include vertices with normals in opposite directions, averaging the source values may cancel out the activity.
Let's use the example from the previous tutorial and consider that one scout contains all the dipoles corresponding to both the red arrows (positive values) and the blue arrows (negative values). If we average all the values from this scout, we get a value close to zero.
To avoid this, a mechanism was added in the scout calculation, to flip the sign of sources with opposite directions before the averaging. We start by finding the dominant orientation of the scout, then flip the sign of the values that are not in the same direction (scalar product of the orientations < 0).
If the sign of some sources is flipped, you get a message in the Matlab command window, for example:
BST> Flipped the sign of 7 sources.
Scout toolbar and menus
Menu: Atlas
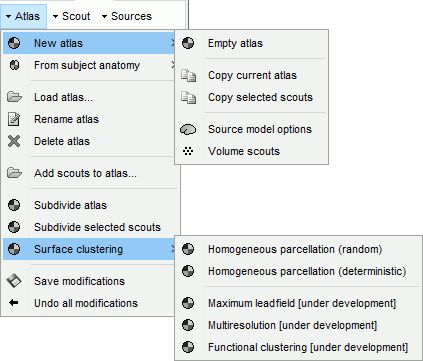
New atlas:
Empty atlas: Create a new empty atlas.
Copy atlas: Duplicate an atlas and all the scouts it contains.
Copy selected scouts: Create a new atlas and copy only the selected scouts to it.
Source model options: Create a special atlas "Source model", with which you can attribute different modeling constraints to each region, after merging the cortex with some subcortical regions. This is required to use the option "Custom head model" (mixed models).
Volume scouts: Create a volume atlas in order to define 3D scouts (volume head models).
From subject anatomy: Import as surface or volume scouts the ROIs defined in one of the volume parcellation available in the subject's anatomy (MNI-based atlases or subject parcellations from CAT12 or FreeSurfer).
Load atlas: Load ROIs or a cortical parcellation coming from FreeSurfer or BrainSuite as a new atlas.
Rename atlas: Change the name that appears in the atlas list to refer to the selected atlas.
Delete atlas: Delete the current atlas and all the scouts in contains.
Add scouts to atlas: Load ROIs or a cortical parcellation and add them to the current atlas.
Subdivide atlas: Splits all the scouts of the current atlas in smaller ROIs. Available options:
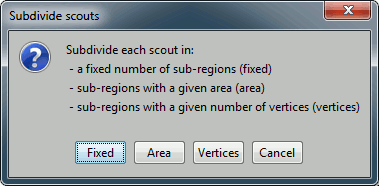
Subdivide selected scouts: Same thing, but processes only the selected scouts.
Surface clustering: Create a parcellation of the cortex surface and saves it as a new atlas.
Only the "Homogeneous parcellation" clustering is currently available.Save modifications: Force the current modifications to be saved to the cortex surface.
Undo all modifications: Restore the atlas the way it was the last time the surface was loaded.
Menu: Scout
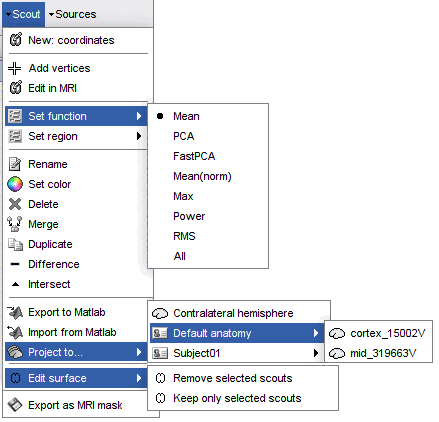
New: coordinates: Create a new scout starting from the vertex that is the closest to the specified coordinates. You can enter coordinates in MRI, SCS or MNI coordinates.
Add vertices: The user can select some points on the cortex surface and add them to the scout.
Equivalent: Click on Select point (first toolbar button) then select the scout in list.Edit in MRI: Open an interface to edit the selected scout as a 3D ROI, slice by slice. Only the vertices contained in the 3D mask are kept, all the volume information is lost. It means that the mask you intended to draw might be very different from what you get as a scout at the end. This is not a very reliable tool, as there is no direct correspondence between the volume and the surface.
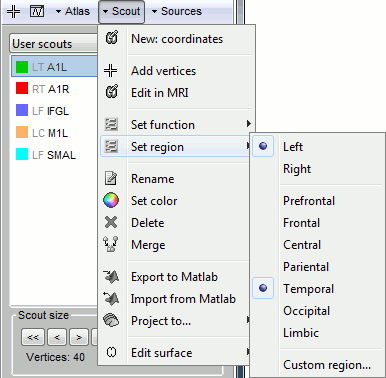
Set function: Set the scout function for the selected scouts.
Set region: Set the cortical region for the selected scouts.
Rename: Rename the selected scout. Shortcut: double-click.
Set color: Change the display color of the selected scout.
Delete: Delete selected scouts. Shortcut: Delete key
Merge: Join two or more selected scouts.
Duplicate: Duplicate selected scouts.
Difference: With two scouts A and B selected, this menu removes the vertices included scouts A from scout B, or vice versa. The direction of the difference is asked in a dialog window.
Intersect: With two or more scouts selected, this menu creates a new scout with the vertices that are common for all the selected scouts.
Export to Matlab: Export the structures of the selected scouts to the Matlab environment and makes them accessible from the command window. This menu can be useful to get quickly the list of vertex indices or modify a scout manually.
Import from Matlab: Import scouts structures that you modified manually from your Matlab command window directly as new scouts.
Project to: Project the selected scout on another surface available in the database.
Project to: Contralateral hemisphere: Transfer scouts defined on one hemisphere to the other hemisphere. This feature is only available with FreeSurfer, and the recon-all segmentation pipeline must be executed with additional options. In the anatomy of the tutorial dataset, this option was not used, and therefore this menu generates an error. However, this option was used in the in the ICBM152 2023b template anatomy. To get this contralateral projection to work, FreeSurfer recon-all must be executed again with the extra option, and the anatomy of the subject must be imported again. More information.
Edit surface: Create a new surface containing only the desired parts (remove or keep only the selected scouts). This is useful for instance for selecting one sub-cortical region from the Aseg FreeSurfer atlas (see the FreeSurfer tutorial).
Export as MRI mask: Create a MRI mask/parcellation using selected scouts (all scouts are used if none is selected). For volume scouts, they are projected from the source grid to the MRI volume. For surface scouts, their cortex patch is projected to the MRI volume, this is based on the registration of the cortical surface and MRI volume.
Menu: Sources
Correlation with sensor: Create a new scout with all the sources that are strongly correlated with a given sensor.
Expand with correlation: Computes the correlation between the values for the scout's seed (first point) and all the other sources. The sources that have correlation coefficient superior to a given threshold are added to the scout.
Maximal value (new scout): Find the vertex with the maximal intensity at the current time, and create a scout centered on it.
Maximal value (selected scout): Move the scout's seed to the source that has the maximum amplitude in the scout, at the current time.
Simulate recordings: Multiply the selected scouts with the forward model. Simulate the scalp data that would be recorded if only the selected cortex region was activated; all the other sources are set to zero. Create a new data file in the database.
If no scout selected: simulate recordings produced by the activity of the entire cortex.
Scout region
A scout is defined by its name, and it has several properties: a list of vertices and an aggregating function. These are usually enough to explore the activity at the cortex level the way we did it in these tutorials. An extra property can be defined on the scout: the explicit classification in a brain region. This property is used only in more advanced functional connectivity analysis, for the representation of the NxN connection graphs. It is introduced here for reference purpose.
A brain region in Brainstorm is following a hierarchy with three levels: hemisphere / lobe / sub-region. The definition at each level is optional: a region can be classified only at the hemisphere level, or at the hemisphere+lobe level, or none of them. It depends on level or hierarchy you are interested in to explore the connectivity graphs.
The region for a scout can be set with the Scout > Set region menus, and is encoded in a string that contains at least 2 characters: "HLxxx". H represents the hemisphere (L,R,U), L stands for the lobe (F,PF,C,P,T,O,L,U), and xxx for the sub-region name (optional). For both the hemisphere and the lobe, the value "U" stands for "Undefined", meaning that the classification is simply not set. The menu Set region>Custom region... lets you directly edit this string.
When set, the region string is shown before the scout name in the list, representing only the defined levels. It doesn't show the letters U for "undefined".
On the hard drive
The scouts are saved in the surface file on which they have been defined.
In the anatomy view, right-click on the selected cortex surface (cortex_15002V) > View file contents.
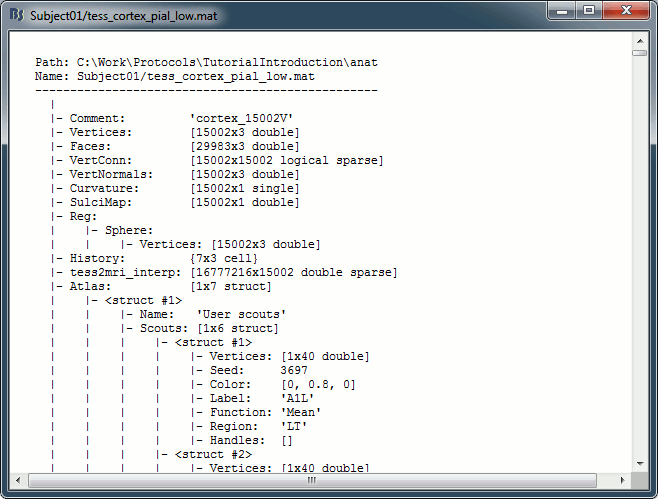
iAtlas: Index of the atlas that is currently selected for this surface.
Atlas: Array of structures, each entry is one menu in the drop-down list in the Scout tab.
Name: Label of the atlas (reserved names: "User scouts", "Structures", "Source model").
Scouts: Array of structures, one per scout in this atlas.
Vertices: Array of indices of the vertices that are part of the scout.
Seed: Index of the central point of the scout (or the most relevant one).
Color: [r,b,g] color array, with values between 0 and 1.
Label: Display name of the scout (must be unique in this atlas).
Function: Scout function {'Mean', 'PCA', 'FastPCA', 'Mean_norm', 'Max', 'Power', 'All'}
Region: Code name for indicating the anatomical region in which the scout is located.
Handles: Graphic handles if the scout is currently displayed (always empty in a file).
Useful functions
bst_scout_value: Combine multiple signals into one.
process_extract_scout: Process "Extract > Scouts time series"
view_scouts: Compute scouts time series and displays them in a new figure.
Additional documentation
Forum: Scout coordinates: http://neuroimage.usc.edu/forums/showthread.php?2375
Forum: Coordinates of the max value from a scout: https://neuroimage.usc.edu/forums/t/get-the-coordinates-of-a-max-value-from-scouts/4959/2
Forum: Importing other FreeSurfer atlases: https://neuroimage.usc.edu/forums/t/930