Granger causality
Authors: Hossein Shahabi and Raymundo Cassani
Granger causality (GC) is a method of ?functional connectivity, adapted by ?Clive Granger in the 1960s, but later refined by ?form that is used today. Granger causality was originally formulated in economics, but has caught the attention of the neuroscience community in recent years. Before this, neuroscience traditionally relied on lesions and applying stimuli on a part of the nervous system to study it’s effect on another part. However, Granger causality made it possible to estimate the statistical influence without requiring direct intervention (Bressler and Seth, 2011).
Contents
Mathematical background
By definition, Granger causality is a measure of linear dependence, which tests whether the variance of error for a ?linear autoregressive model estimation of a signal (A) can be reduced when adding a linear model estimation of a second signal (B). If this is true, signal B has a Granger causal effect on the first signal A, i.e., independent information of the past of B improves the prediction of A above and beyond the information contained in the past of A alone. The term independent is emphasized because it creates some interesting properties for GC, such as that it is invariant under rescaling of A and B, as well as the addition of a multiple of A to B. The measure of Granger Causality is nonnegative, and zero when there is no Granger causality(Geweke, 1982).
The main advantage of Granger Causality is that it is an asymmetrical measure, in that it can dissociate between A->B versus B->A. It is important to note however that though the directionality of Granger Causality is a step closer towards measuring effective connectivity compared to symmetrical measures, it should still not be confused with “true causality”. Effective connectivity estimates the effective mechanism generating the observed data (model based approach), whereas GC is a measure of causal effect based on prediction, i.e., how well the model is improved when taking variables into account that are interacting (data-driven approach) (Barrett and Barnett, 2013).
The difference with causality is best illustrated when there are more variables interacting in a system than those taken into account in the model. For example, if a variable C is causing both A and B, but with a smaller delay for B than for A, then a GC measure between A and B would show a non-zero GC for B->A, even though B is not truly causing A (Bressler and Seth, 2011).
IMAGE
GC in time domain
Even though GC has been extended for nonlinear, multivariate and time-varying conditions, in this tutorial we will stick to the basic case, which is a linear, bivariate and stationary model defined in both the time and spectral domain. In the time domain this can be represented in the following way. If x represents a signal that can be modeled using a linear autoregressive model estimation (AR model) in the following two ways:
![\begin{eqnarray*}
x(t) & = & \sum_{k=1}^{p} [A_{k}x(t-k)] +e_1 \\
x(t) & = & \sum_{k=1}^{p} [A_{k}x(t-k)+B_{k}y(t-k)] +e_2
\end{eqnarray*} \begin{eqnarray*}
x(t) & = & \sum_{k=1}^{p} [A_{k}x(t-k)] +e_1 \\
x(t) & = & \sum_{k=1}^{p} [A_{k}x(t-k)+B_{k}y(t-k)] +e_2
\end{eqnarray*}](/brainstorm/GrangerCausality?action=AttachFile&do=get&target=latex_ff96bd67f304dfa4455accf112cb1620facee597_p1.png)
Where 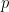 represent the amount of past information that will be included in the prediction of the future sample, and is called the model order. In these two equations, the first models $x$ using the past (and present) of only itself whereas the second includes the past (and present) of a second signal $y$. Note that when only past measures of signals are taken into account ($k\geq 1$), the model ignores simultaneous connectivity, which makes it less susceptible to volume conduction (Cohen, 2014). Then, according to the original formulation of GC, the measure of GC is defined as follows:
represent the amount of past information that will be included in the prediction of the future sample, and is called the model order. In these two equations, the first models $x$ using the past (and present) of only itself whereas the second includes the past (and present) of a second signal $y$. Note that when only past measures of signals are taken into account ($k\geq 1$), the model ignores simultaneous connectivity, which makes it less susceptible to volume conduction (Cohen, 2014). Then, according to the original formulation of GC, the measure of GC is defined as follows:
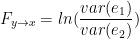
Which is 0 if $var(e_{1})=var(e_{2})$ and a non-negative value if $var(e_{1}) > var(e_{2})$. Note that $var(e+{1}) \geq var(e_{2})$ always holds, as the model can only improve when adding new information.
GC in Spectral Domain
An important improvement by Geweke was to show that under fairly general conditions, $F_{y \to x}$ can be decomposed by frequency. If the two autoregressive models in time domain are specified as:
![\begin{eqnarray*}
x(t) &=\sum_{k=1}^{p} [A_{k_{xx}}x(t-k) + A_{k_{xy}}y(t-k)] +\sigma_{xy} \\
y(t) &=\sum_{k=1}^{p} [A_{k_{yy}}y(t-k) + A_{k_{yx}}y(t-k)] +\sigma_{yx}
\end{eqnarray*} \begin{eqnarray*}
x(t) &=\sum_{k=1}^{p} [A_{k_{xx}}x(t-k) + A_{k_{xy}}y(t-k)] +\sigma_{xy} \\
y(t) &=\sum_{k=1}^{p} [A_{k_{yy}}y(t-k) + A_{k_{yx}}y(t-k)] +\sigma_{yx}
\end{eqnarray*}](/brainstorm/GrangerCausality?action=AttachFile&do=get&target=latex_7053c1d2e3c64970fd2e3c0b13042ca58f9fcdc7_p1.png)
In each equation the reduced model can be defined when each signal is an autoregressive model of only it’s own past, with error terms $\sigma_{xx}$ and $\sigma_{yy}$. Then we can defined the variance-covariance matrix of the whole system as such:
![\begin{eqnarray*}
\Sigma =
\left[\begin{array}{cc}
\Sigma_{xx} & \Sigma_{xy} \\
\Sigma_{yx} & \Sigma_{yy}
\end{array}\right] \\
where \Sigma_{xx}=var(\sigma_{xx}), etc
\end{eqnarray*} \begin{eqnarray*}
\Sigma =
\left[\begin{array}{cc}
\Sigma_{xx} & \Sigma_{xy} \\
\Sigma_{yx} & \Sigma_{yy}
\end{array}\right] \\
where \Sigma_{xx}=var(\sigma_{xx}), etc
\end{eqnarray*}](/brainstorm/GrangerCausality?action=AttachFile&do=get&target=latex_2572ecc5db09aa7ba97841aeef5dc4c963c2a107_p1.png)
Now the Fourier transform of these equations for a give frequency $\omega$ can be expressed as:
\begin{equation}
- \begin{pmatrix}
A_{xx}(w) & A_{xy}(w) \\ A_{yx}(w) & A_{yy}(w) \\ \end{pmatrix} \begin{pmatrix} x(w)\\ y(w)\\ \end{pmatrix}
=
- \begin{pmatrix} \varepsilon_{1}(w) \\ \varepsilon_{2}(w) \\ \end{pmatrix}
\end{equation}
Rewriting this as
\begin{equation}
- \begin{pmatrix} x(w) \\ y(w) \\ \end{pmatrix}
=
- \begin{pmatrix}
H_{xx}(w) & H_{xy}(w)\\ H_{yx}(w) & H_{yy}(w)\\ \end{pmatrix} \begin{pmatrix} \varepsilon_{1}(w) \\ \varepsilon_{2}(w) \\ \end{pmatrix}
\end{equation}
Where H is the transfer matrix. The spectral matrix is then defined as:
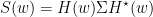
Finally, assuming independence of $x$ and $y$, and $\Sigma_{xy}=\Sigma_{yx}=0$ , we can define GC for a give frequency as:
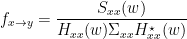
Note that the ability to reformulate the equations in spectral domain also makes it very easy to generalize to a multivariate GC. That is why all multivariate measures of GC are defined in spectral domain. Alternative measures derived from these multivariate autoregressive estimates are Partial Directed Coherence (PDC) and the Directed Transfer Function (DTF).
A few noteworthy practical issues about GC
1. Temporal resolution: the high time resolution offered by MEG,EEG, and intracranial EEG allows for a very powerful application of GC and also offers the important advantage of spectral analysis.
1. Stationarity: the GC methods described so far are all based on AR models, and therefore assume stationarity of the signal (constant auto-correlation over time). However neuroscience data, especially task based data such as event-related potentials are mostly nonstationary. There are two possible approaches to solve this problem. The first is to apply methods such as differencing, filtering, and smoothing to make the data stationary (see recommendation for time domain GC). Dynamical changes in the connectivity profile cannot be detected with the first approach. The second approach is to turn to versions of GC that have been adapted for nonstationary data, either by using a non-parametric estimation of GC, or through measures of time-varying GC, which estimate dynamic parameters with adaptive or short-time window methods (Bressler and Seth, 2011).
1. Number of variables: Granger causality is very time-consuming in the multivariate case for many variables ($O(m^2)$) where m represents the number of variables). Since each connection pair results in two values, there will also be a large number of statistical comparisons that need to be controlled for. When performing GC in the spectral domain, this number increases even more as statistical tests have to be performed per frequency. Therefore it is usually recommended to select a limited number of ROIs or electrodes based on some hypothesis found in previous literature, or on some initial processing with a more simple and less computationally heavy measure of connectivity.
1. Pre-processing: The influence of pre-processing steps such as filtering and smoothing on GC estimates is a crucial issue. Studies have generally suggested to limit filtering only for artefact removal or to improve the stationarity of the data, but cautioned against band-pass filtering to isolate causal influence within a specific frequency band (Barnett and Seth, 2011).
1. Volume Conduction: Granger causality can be performed both in the scalp domain or in the source domain. Though spectral domain GC generally does not incorporate present values of the signals in the model, it is still not immune from spurious connectivity measures due to volume conduction (for a discussion see (Steen et al., 2016)). Therefore it is recommended to reduce the problem of signal mixing using additional processing steps such as performing source localization, and doing connectivity in source domain.
1. Data length: because of the extent of parameters that need to be estimated, the number of data points should be sufficient for good fit of the model. This is especially true for windowing approaches, where data is cut into smaller epochs. A rule of thumb is that the number of estimated parameters should be at least (~10) several times smaller than the number of data points.
Simulate signals
Since the linear parametric GC is based on autoregressive modeling of signals, a simulation example based on AR modeling provides a nice ground truth to validate the results. In this example, we generate a dataset for a 3-variate autoregressive model (example 2 in the study of (Baccala and Sameshima, 2001)). These observed time series are assumed to be generated by the equations:

Where $w(n)$ are zero-mean uncorrelated white processes with identical variances. Since the model order in this example is one, we can rewrite the equations in matrix form as such:
\begin{equation} X=
- \begin{bmatrix}
0.5 & 0.3 & 0.4 \\ -0.5 & 0.3 & 1 \\ 0 & -0.3 7 -0.2 \\ \end{bmatrix}
X_1+
- \begin{bmatrix}
1 & 0 & 0 \\ 0 & 1 & 0 \\ 0 & 0 & 1 \\ \end{bmatrix}
W \end{equation}
To do simulate this in Brainstorm, we use the following steps: \item Select the menu File > Create new protocol > "TutorialGC" and select the options:
- \begin{itemize}
- \item "Yes, use protocol's default anatomy", \item "No, use one channel file per acquisition run (MEG/EEG)".
- \item In the Process1 tab, leave the file list empty and click on the button [Run]
\item Select the process "Simulate > Simulate AR signals". \item Set all the parameters as follows:
(figure 1)
Which is a 10 second simulation with 100Hz sampling rate. The process will give you a data file presented in Brainstorm as:
(figure 2)
When displaying the data as time-series:
(figure 3)
Process options
Subject name:} Specify in which subject the simulated file is saved. Condition name:} Specify the folder in which the file is saved (default: "Simulation"). Number of time samples:} number of data samples per simulation Signal sampling frequency:} Sampling frequency of the simulated signal in Hz. Nested and nesting frequency:} Definition of the phase-amplitude coupling. Coefficient matrix:} coefficient matrix as defined in the matrices in equations above. There are p (model order) number of A matrices, each the size of variable by variables. These are then combined in the following fashion: $A=[A_1;A_2;…;A_p ]$; Intercept:} constant offset for each variable, as defined by a matrix with dimensions 1 by variables. Noise covariance matrix C:} covariance matrix for the error (noise) term. Since we are modeling an uncorrelated white process with identical variance, this is equivalent to an identity matrix with dimensions variables by variables.
Bivariate Granger Causality NxN in time domain}
The data generated above is then used to estimate GC. To estimate GC, drag the simulated file to the Process1 tab:
(figure 4)
Select the process \textbf{“Connectivity $>$ Bivariate Granger Causality NxN”} Set the parameters as follows:
(figure 5)
The connectivity profile calculated is saved in the file Granger: full. Displaying this as an image will show the following:
(figure 6)
In this image, an asymmetrical matrix of GC estimates is shown, with nonnegative values. Note that this map is not a multivariate analysis of our multivariate system, but instead a 2by2 bivariate analysis. It is known that bivariate connectivity measures to estimate the correct pattern of propagation in a multivariate system can be problematic, nevertheless the results still show a pretty good estimation of connectivity, as can be seen in the strong propagation of activity going from variable 3 $\to$ 2, as is the case in the originally described model. The next strong connections are from 1 $\to$ 2, and from 3 $\to$ 1. Connections 2 $\to$ 1 and 2 $\to$ 3 show the same value, as they have the same value though with negative signs. Note that the relative strengths of connectivity in the results are reserved, but the value itself is a ratio of the residual errors of the full and reduced model as described in the introduction, and should not be confused with the actual coefficient values. You also have the option of showing the results as a graph by right-clicking on the Granger file > Display as graph.
(figure 7)
1. You can choose the threshold for the coupling intensity shown in the right hand Display panel (Intensity Thresh).
1. To set link transparency and size, right-click on the figure: Display options
1. To detect incoming and outgoing connections, select the variable you want to see by clicking on it, and in the right hand Display panel, section direction, select Out or In.
Process options
Input options
\item \textbf{Time window:} specifies the time window you want to use for your model. \item \textbf{Remove evoked response from each trial:} this option refers to subtracting the average of phase-locked activity (ERP) from each individual trial. Presently some studies measure interdependency of ongoing brain activity by removing the average event-related potential from each trial. It is also recommended by some as it meets the zero-mean stationarity requirement (improves stationarity of the system). However, the problem with this approach is that it does not account for trial-to-trial variability (For a discussion see (Wang et al., 2008)).
Estimator options:
Selection of model order is a critical issue and is typically evaluated from criteria derived from information theory. Several criteria have been proposed, of which the most used are Akaike’s information criterion, the Bayesian-Schwartz’s criterion, and the Hannan-Quinn criterion (Koichi and Antonio, 2014). Model fitting quality crucially depends on the proper model order selection. Too low orders may lack the necessary details, while too big orders tend to create spurious values of connectivity. Note that in our simulated example even though the simulation was created with an underlying model of 1, a Granger model order of 3 was selected with decent resulting connectivity.
Output options:
\item \textbf{Save individual results (one file per input file):} option to save GC estimates on several files separately. \item \textbf{Concatenate input files before processing (one file):} option to save GC estimates on several files as one concatenated matrix.
File contents
The output of the Bivariate Granger Causality NxN is an NxN matrix saved in the TF domain, and represent the coupling strength between variables. You can see these contents by right-clicking in the Granger file $>$ File $>$ View file contents.
(figure 8)
Recommendations
As mentioned before, a model order should be selected with care. In electrophysiological data, a model order should also make some biological sense. For example in EEG data with 2Khz sampling rate an order of 5 would correspond to a 2.5ms of past information incorporated to train your model, which is probably too little past information to make a decent judgement. Note that to take an arbitrary high model order is also not recommended, as it will increase the type I error. We will refrain from recommending a rule of thumb model order, but instead advise to use the information criteria such as AIC or BIC to compare model fits:

Where $\Sigma$ is the noise covariance matrix, $p$ model order, $k$ number of variables, and $N$ is the number of data samples
On a similar note, the sampling rate of the data is also not an easy choice to make. Even though too low sampling rate will eliminate possible dynamics of the data, if the sampling rate is too high the model will be computationally very expensive, as a very high model order will be required. For most M/EEG data, a sampling rate between 200 and 1000 Hz will probably suffice (Cohen, 2014). As mentioned in the introduction, the current model is based on the assumption of stationarity. Therefore it is important to first check if the time series we are using are stationary with statistical tests such as the KPSS test or the Dickey-Fuller (unit root test). If the data is shown to be nonstationary, certain techniques can still render it stationary (depending on the type of nonstationarity). A common approach is to difference the data to eliminate any trends in the time series (i.e., $x_t^{'}=x_{t}-x_{t-1}$). This can be done several times if necessary. A problem with this is that it could potentially change the interpretation of any resulting GC, specifically in the spectral domain where differencing acts as a high filter. Other commonly used methods are smoothing or filtering the data, but in the end if there are causal relation that vary over time these technique will make it impossible to find those. An alternative approach could be to divide the data into time-series that are (sufficiently) locally stationary, and apply the GC per time window. This method makes is possible to study changes in connectivity over time, but is of course not without it’s own problems, the most important of which is the choice of the time window. If the time window is too short, there will not be enough data points to make a decent estimation.
Bivariate Granger Causality NxN in spectral domain
To estimate GC values in spectral domain, drag your data to the Process1 tab as before. Select run $>$ Connectivity $>$ Bivariate Granger causality (spectral) NxN Set the parameters as the following
(figure 9)
To view the spectral profile for each bivariate connection pair, right-click on the SpGranger file $>$ Power Spectrum
(figure 10)
Where the first variable in the legend refers to the sender and the second to the receiver of information. As we can see from the figure the large coupling strength from 2->3 is highest in higher frequency ranges. You can select your desired connectivity pair, and then press enter to view it in a separate window.
Process options
Input options
\item \textbf{Time window:} same as time domain GC. \item \textbf{remove evoked response from each trial:} same as time domain GC.
Estimator options
\item \textbf{Maximum Granger model order:} same as time domain GC. \item \textbf{Maximum frequency resolution:} frequency steps of the spectral GC. A large number of steps (high resolution) will increase the number of statistical comparisons. Note that the frequency resolution has to be less than half the highest frequency of interest. \item \textbf{Highest frequency of interest:} maximum frequency evaluated. Based on the Nyquist theorem, this value needs to be less than half the sampling frequency
Output options
\item \textbf{Save individual results (one file per input file):} same as time domain GC. \item \textbf{Concatenate input files before processing (one file):} same as time domain GC.
File Contents
Files saved by this process are saved in the format links by trials by frequency
\textbf{Recommendations}
(figure 11)
