|
Size: 11400
Comment:
|
Size: 26154
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| == Current topics == ==== Functionnal connectivity ==== * Integration of different metrics to study the brain connectivity: <<BR>>Correlation, coherence, Granger causality, phase locking value * Development of new ways to represent the connectivity between sensors or brain regions ==== EEG / epilepsy / intra-cranial recordings ==== * Editing the position of intracranial electrodes in the MRI viewer ==== Source modeling ==== * Computation of equivalent current dipoles * Beamformers ==== Large scale analysis ==== * Parallel processing: Reduce the computation times using the parallel processing toolbox * Distributed processing: Integrate tools for sending Brainstorm processes on computation clusters <<BR>><<BR>><<BR>> |
<<TableOfContents(2,2)>> |
| Line 23: | Line 7: |
| * RAW file viewer: | * RAW file viewer:<<BR>> * Bottom bar: display extend events as segments instead of dots (to see their extension) * Downsample before filtering? (attention to the filter design) * Add parameter to make the visual downsampling more or less aggressive |
| Line 25: | Line 12: |
| * Documentation: Add definition of bad segments * 2DLayout: Doesn't work when changing page => need refresh of GlobalData.Preferences.TopoLayoutOptions.TimeWindow * EEG reference/storage: * Intracranial electrodes: Define and display in the MRI viewer + 3D figures * Bad locations: indicate with [NaN NaN NaN] instead of [0 0 0] * Bad channels that can be specified at the program level (for sites that have permanently bad channels) => AS Dubarry * RAW processing: * Make it work for all the file formats (at least bandpass filter + sin removal) * Events: advanced process for recombining. Example: http://www.erpinfo.org/erplab/erplab-documentation/manual/Binlister.html |
* Keep the filter specifications in memory instead of recomputing for every page * Add field "comment" to markers: For clinicians to add notes (Marcel) * Events: Change the category of a selected event easily, instead of deleting/marking new * Events: Advanced process for recombining.<<BR>>Example: http://www.erpinfo.org/erplab/erplab-documentation/manual/Binlister.html * Bad trials: When changing the status of bad to good: remove the bad segments as well, otherwise it is not processed by processes like the PSD. * Review clinical recordings: Reduce the dimensionality of the data with a simple inverse problem, similar to what we do for the magnetic extrapolation ("Regional sources" in BESA, cf S Rampp) * MEG/EEG registration: Apply the same transformation to multiple runs * 2DLayout: * Does not work when DC offset is not removed * Add a proper amplitude scale that gets updated when shift+scroll, to compare figures * Create heat maps: Maybe with matlab function heatmap? == Interface == * Add a warning when computing a forward model with > 100000 sources (check selection) * Snapshot: Save as image / all figures (similar to Movie/all figure) * Generalize the use of the units (field .DisplayUnits): Rewrite processes to save the units correctly |
| Line 35: | Line 29: |
| * Create a colormap similar to MNE, where extrema are bright * Import data: * Save properties "baseline" and "resample" at the level of the protocol (to re-use for all the files) * NIRS: * Add new data type * Display of sensors by pairs oxy/deoxy (red/blue), overlaid * Images of amplitude: [sensor x time], [trial x time], scout: [trial x time] * Can be done with Matrix > View as image: extract cluster, concatenate for all trials * 2D Layout for multiple conditions |
* Allow brightness/contrast manipulations on the custom colormaps * Global colormap max: Should get the maximum across all the open files * Copy figures to clipboard (with the screencapture function) * Display warning before opening files that are too big * Smooth display from figure_image (ERPimage, raster plot...) * Contact sheets & movies: use average of time windows instead of single instants, for each picture. * Contact sheets: Allow explicit list of times in input (+ display as in MNE-Python with TS) * Display CTF coils: Show discs instead of squares * Use boundary() instead of conhull() in all the display functions (ie. 2DDisc) * Progress bar: * Add different levels (to handle sub-processes) * Uniformize calls in bst_process/Run * Add a "Cancel" button * Error message: Add a link to report directly the bug on the forum * Reorganize menus (Dannie's suggestion): {{attachment:dannie_menus.png||width="382",height="237"}} |
| Line 46: | Line 47: |
| * Tutorial coherence [1xN] * t-tests on connectivity measures |
* Thresholding and stat tests the connectivity matrices * Connectivity on unconstrained sources: "Default signal extraction for volume grids should be the time series of the first principal component of the triplet signals after each has been zero-meaned" (SB) * Display of connectivity graphs: * Display as straight lines * Recode 2D graphs * 3D display with anatomical constrains * Display using real position of EEG electrodes * Use new band-pass filters in bst_connectivity ('bst-hfilter' instead of 'bst-fft-fir') * Review by Jan-Mathijs: http://journal.frontiersin.org/article/10.3389/fnsys.2015.00175/full * Connectivity based on band limited power (Sylvain): * Compute Hilbert/Bandpass + correlation of the envelopes * Bandpass envelopes before computing correlations? * Compute Hilbert(sensors) and then project to source space? * Matrix view of NxN graphs: Add legend of the elements along X and Y axis |
| Line 49: | Line 62: |
| * Does not display negative values correctly (correlation or difference of coherence) * Re-write using pure Matlab code and smoothed graphics |
|
| Line 50: | Line 65: |
| * Fix zoom in one region | |
| Line 53: | Line 67: |
| * Fixed scales for intensity sliders * Add "=" shortcut for having graphs with similar configurations * Disable zoom in one region (serious bugs) * NxN on sensors: does not place the sensors correctly in space * Coherence: * Average cross-spectra instead of concatenating epochs (to avoid discontinuities)<<BR>>Explore inter-trial approaches (Esther refers to chronux toolbox) * Granger: * Crashes sometimes: improve stability * Check for minimum time window (Esther: min around 500-1000 data points) * Re-write and optimize code * Add progress bar * PLV: * Add p-values * Remove evoked * Optimize code * Add time integration * Unconstrained sources * Add warning when running of short windows (because of filters) * PAC: * Add input TF , to disconnect TF decomposition and PAC computation (Peter) * Refine frequency vector of low frequencies * How many central frequencies to use in bst_pac? * Change filters: no chirplet functions * bst_freqfilter: Use nfcomponents like in bst_pac * Esther recommended a larger frequency binning of the PAC estimation * PAC maps: Display all sensors at once (like TF and DynamicPAC) * Hui-Ling's PAC: * https://bsp.hackpad.com/Cross-Frequency-Coupling-cChe95lhDHz * https://github.com/NCTU-BSP/MEEG * Time-resolved correlation/coherence: Display as time bands |
|
| Line 57: | Line 101: |
| * Work on progress bars | * Tutorial coherence [1xN] : Reproduce FieldTrip results? * Connect NxN: Display as time series > Display warning before trying to open too many signals |
| Line 60: | Line 105: |
| * Decoding/Classifiers: Implement Dimitrios and scikit-learn algorithms * Allow processes in Python and Java * Add MNE-Python functions: * scikit-learn classifiers * Implement data exchange with MNE-Python: write FIF files from Brainstorm and/or pass python objects in memory instead of FIF files * SSS/tSSS cleaning * Reproduce other tutorials / examples * Change the graphic renderer from Matlab * Add FieldTrip functions: * ft_sourceanalysis: * Check noise covariance * Check all the options of all the methods * Single trial reconstructions + noise covariance? * Filters?? http://www.fieldtriptoolbox.org/example/common_filters_in_beamforming * Beamformers: Save ftSource.avg.mom <<BR>>http://www.fieldtriptoolbox.org/workshop/meg-uk-2015/fieldtrip-beamformer-demo * http://www.natmeg.se/ft_beamformer/beamformer.html * http://www.fieldtriptoolbox.org/tutorial/beamformingextended * Baseline? Two inputs? * ft_prepare_sourcemodel: Compute MNI transformation (linear and non-linear) => Peter * Freqanalysis: ITC * ft_read_atlas('TTatlas+tlrc.BRICK'); * ft_volumereslice: http://www.fieldtriptoolbox.org/faq/how_change_mri_orientation_size_fov * ft_freqanalysis * ft_combineplanar * Use CUDA for speeding up some operations (filtering, wavelets, etc) * Pipeline editor: * Bug: After "convert to continuous", the time of the following processes should change * Add loops over subjects/conditions/trial groups * Events: Allow selection from a drop-down list (similar to option "channelname" in panel_process_selection) * ITC: Inter-trial coherence (see MNE reports for group tutorial)<<BR>>http://www.sciencedirect.com/science/article/pii/S1053811916304232 * ICA: * Why doesn't the ICA process converge when using 25 components in the EEG tutorial? * Add an option to resample the signals before computing the ICA decomposition * Add a stand-alone tutorial * Exploration: Add window with spectral decomposition (useful for muscle artifacts) * Export IC time series (and then compute their spectrum): solves the problem above * Comparison JADE/Infomax: <<BR>> http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0030135 * Use faster methods (MNE-Python?) * Add methods: SOBI, Fastica, AMICA/CUDICA (recommended by S Makeig) * Dimension reduction with PCA adds artifacts: Not done by default in EEGLAB<<BR>>Contact: Stephen Shall Jones ( shall-jones@infoscience.otago.ac.nz )<<BR>>Student Carl Leichter detailed this in his thesis * S Makeig: Use ICA to select the IC of interest instead of only removing artifacts * Display of spectrum for components (PSD/FFT) * Use FastICA (algo crashing) * Add components preselection: Correlation with EOG/ECG * Import ICA matrices available in EEGLAB .set files * EEGLAB recommends ICA + trial rejection + ICA again: Impossible right now with Brainstorm<<BR>>(http://sccn.ucsd.edu/wiki/Chapter_09:_Decomposing_Data_Using_ICA) * ICA+machine learning: https://www.ncbi.nlm.nih.gov/pubmed/28497769 * Automated artifact rejection: https://arxiv.org/abs/1612.08194 * Save IC time series in database * Use EYE-EEG: EEGLAB toolbox for eye-tracker guided ICA (Olaf Dimigen): http://www2.hu-berlin.de/eyetracking-eeg/ * Other EEGLAB functions: * Step function detection: https://github.com/lucklab/erplab/wiki/Artifact-Detection:-Tutorial * SSP: * Display warning if changing the ChannelFlag while there is a Projector applied * Pipeline editor: * When computing sources from the pipeline editor: doesn't reselect the options if you click twice on "edit" (works for minnorm, but not for lcmv) * When computing time-frequency/hilbert/psd: Find a way not to force the user to click on Edit * Bandpass: * Use new filters in all the functions using a bandpass ('bst-hfilter' instead of 'bst-fft-fir'): process_evt_detect_threshold * Weird bug: Filter(import) != Import(Filter) in the HCP tutorial... to investigate * Spectral flattening (John): * ARIMA(5,0,1): Apply on the signal before any frequency/connectivity/PAC analysis * PSD: * Rewrite to have the same input as coherence (frequency resolution instead of window length) * Use the progress bar * Allow display of Avg+StdErr * Remove line noise: http://www.nitrc.org/projects/cleanline * Use Matlab Coder to optimize some processes: Wavelets, bandpass filter, sinusoid removal |
|
| Line 61: | Line 178: |
| * Frequency bands: extended syntax (ex: [2 3 4], 10:5:90, ...) * How to combine 3 orientations for unconstrained sources * Display logs as negative |
* Optimization: bst_timefreq (around l.136), remove evoked in source space: Average should be computed in sensor space instead of source space (requested by Dimitrios) * Short-time Fourier transform: http://www.mikexcohen.com/lectures.html * Matching pursuit: http://m.jneurosci.org/content/36/12/3399.abstract?etoc * Bug: Display logs as negative * Bug: 3D figures: Colormaps with "log" option doesn't work * Bug: Difference of power displayed in log: problems (Soheila) |
| Line 66: | Line 186: |
| * Smooth display of TF/PAC maps (option) | |
| Line 68: | Line 187: |
| * Bandpass: Show warning when using inappropriate high-pass filter (precision too high) | * Impossible to keep complex values for unconstrained sources * Pad short epochs with zero values for getting lower frequencies * Hilbert with time bands very slow on very long files (eg. 3600s at 1000Hz) because the time vector is still full (10^7 values): save compressed time vector instead. * Extend clusters tab to display of TF to overlay TF signals (Svet) * When normalizing with baseline: Propagate with the edge effects marked in TFmask * Allow baseline normalization of files computed with time bands * Allow running TF on montages * Review continuous files in time-frequency space (for epilepsy) * Bug when computing TF on constrained and unconstrained scouts at the same time (in mixed head models for instance): uses only the constrained information and doesn't sum the 3 orientations for the unconstrained regions. |
| Line 70: | Line 197: |
| * Detection of bad segments in the RAW files | |
| Line 73: | Line 199: |
| * Distributed processing: Brainstorm that can run without Java * SSP: * Display warning if changing the !ChannelFlag while there is a Projector applied * Show where the attenuation is projected:<<BR>>(sum(IK,2)-sum(SSP(k,:)*IK,2)./sum(IK,2) * Average: * Remember how many trials were used per channel * Save standard deviation * Display standard deviation as a halo around the time series * Co-registration of MEG runs: * SSP: Group projectors coming from different files * Finish validation of the method * Apply to continuous recordings for correcting head movements * Current Source Density (CSD) => Ghislaine<<BR>>http://psychophysiology.cpmc.columbia.edu/software/CSDtoolbox/index.html * Other processes: * Detrending * Moving average * Max * Median * Significance test (Dimitrios, Leo) * Spatial smoothing: check / document parameters * Contact sheets & movies: use average of time windows instead of single instants, for each picture. * Optical flow |
* Events detection: Add option "std" vs "amplitude" * Simulation: * Fix units in simulation processes => no *1e-9 in "simulate recordings" * Use "add noise" process from Hui-Ling (in Work/Dev/Divers) * Use field process field "Group" to separate Input/Processing/Output options * Use new Matlab functions: movmean, movsum, movmedian, movmax, movmin, movvar, movstd * Process ft_prepare_heamodel: Add support from BEM surfaces from the Brainstorm database |
| Line 97: | Line 208: |
| * Rename protocol * Faster DB searches (for Emily) * Add buttons to sort files: by name, by comment, by date |
|
| Line 98: | Line 212: |
| * Group matrix files => allow to process matrix files by trial types | * Matrix files: Allow to be dependent from other files |
| Line 104: | Line 218: |
| * Rename multiple files * Default headmodel lost when reloaded: Keep selection on the hard drive (in brainstormstudy.mat) * Auto-save: * protocol.mat can be too big: do not store the results links in it (and recreate when loading)- http://neuroimage.usc.edu/forums/t/abnormally-slow-behavior/2065/10 * Improve auto-save: add tracking file next to protocol.mat, do not save all the time, only when closing app, and reload protocol at stratup if tracking file is still there == Distributed computing == * Options from FieldTrip: * Loose collection of computers: https://github.com/fieldtrip/fieldtrip/tree/master/peer * Alternative, with less limitations: http://research.cs.wisc.edu/htcondor/ * Single multicore machine: https://github.com/fieldtrip/fieldtrip/tree/master/engine * Batch system: https://github.com/fieldtrip/fieldtrip/tree/master/qsub * Documentation: http://fieldtrip.fcdonders.nl/faq#distributed_computing_with_fieldtrip_and_matlab * PSOM: http://psom.simexp-lab.org/ * Various initiatives: http://samirdas.github.io/Data_sharing.html#/ |
|
| Line 105: | Line 238: |
| * Stenroos 2014 paper: Include the following methods * Inner and outer skull surfaces generator from !FieldTrip (needs SPM, probably not so different from BST) * Nolte corrected-sphere model (good model re:Alex) * Fast BEM models * Dipole fitting * Visualize Beamformer results: * Read CTF SAM .svl * Display as layers in the MRI viewer |
* Dipoles: * Project individual dipoles files on a template * panel_dipoles: Doesn't work with multiple figures * Project sources: Very poor algorithm to project sub-cortical regions and cerebellum (algorithm to fit surfaces should be imrpoved) * Menu head model > Copy to other conditions/subjects (check if applicable first) * Menu Sources > Maximum value: Doesn't work with volume or mixed head models * Mixed head models: * Set loose parameter from the interface * Volume grid: * Optimize: 3D display (better than 9x9 cubes) * Optimize: vol_dilate (with 26 neighbors) * Menu Sources > Simulate recordings: * Do not close the 3D figures after generating a new file * Add a process equivalent to this menu * Panel Get coordinates: Add button "find maximum" * BEM single sphere: Get implementation from MNE |
| Line 114: | Line 255: |
| * Compute unconstrained and then project on the normal ? * Difference and stat should be: norm(A) - norm(B) |
|
| Line 118: | Line 257: |
| * Volume grid: * Scouts 3D * Test volume sources with all the subsequent processes (timefreq, stat...) * Optimize: 3D display (better that 9x9 cubes) * Optimize: vol_dilate (with 26 neighbors) * Magnetic extrapolation: Do the same thing with EEG |
|
| Line 125: | Line 258: |
| * Display with figure_image() | |
| Line 129: | Line 263: |
| * When deploying to other conditions: Apply destination SSP (!NoiseCov = SSP . !NoiseCov . SSP' ) | |
| Line 131: | Line 264: |
| * Simulation: synthesize pseudo data-files from a cortex patch (duration, amplitude, noise) * Calculate !ImagingKernel * Gain for a scout * EEG Source modeling: Manage references and bipolar montages properly (maybe not necessary) * MEG source modeling: Do reconstruction only for a subset of sensors for estimating dipoles? * Processes compute head model and sources: Additional option to set the file comment |
* Time-frequency beamformers: * Band-pass everything in different frequency bands + Source estimation + TF * Ask data to Sarang where he sees effects that cannot be extracted with MN followed by TF * Process "Extract scouts time series": Add PCA option (replace isnorm with choice PCA/Norm) * Add eyes models to attract eye activity |
| Line 138: | Line 271: |
| * Segmentation with SPM12 * BrainVISA: Add support for MarsAtlas * MNI transformation: Use SPM non-linear MNI transformation y_... * Registration: * Getting electrode positions from 3D scanners: https://sccn.ucsd.edu/wiki/Get_chanlocs * GARDEL: http://meg.univ-amu.fr/wiki/GARDEL:presentation * Use the same registration for multiple recording sessions that have already re-registered previously (eg. with MaxFilter) * When linking multiple EEG recordings including 3D positions, do the registration only once and copy it to all the runs * Compute non-linear MNI registration instead of linear * Select and remove bad digitized head points before automatic coregistration * Load the MNE -transf.fif: http://neuroimage.usc.edu/forums/showthread.php?2830 * MRI Viewer: * Pan in zoomed view (shift + click + move?) * Zoom in/out with mouse (shift + scroll?) * Ruler tool to measure distances * Display scouts as additional volumes * Render surface envelope in the MRI as a thin line instead of the full interpolation matrix * Edit fiducials: Replace 6 text boxes with 1 for easy copy-paste (see fiducials.m) * Optimize computation interpolation MRI-surface (tess_tri_interp) => spm_mesh_to_grid * BrainSuite: * Add new labels to all BrainSuite anatomy templates * Use BrainSuite inner skull for surface generation * Use same colors for left and right for anatomical atlases |
|
| Line 139: | Line 295: |
| * Mix constrained/unconstrained/volume sources, using the "Source model" atlas | |
| Line 141: | Line 296: |
| * Project scouts betweens subjects and between hemispheres | |
| Line 143: | Line 297: |
| * Sort scouts by region in process options | |
| Line 144: | Line 299: |
| * Sort scouts by region in process options * Generate mixed density surfaces * Import / registration: * Major bug when importing surfaces for an MRI that was re-oriented manually * Use mid-gray instead of pial surface? |
* Project from one hemisphere to the other using registered spheres/squares (http://neuroimage.usc.edu/forums/t/how-to-create-mirror-roi-in-the-other-hemisphere/5910/8) * Parcellating volume grids: scikit-learn.cluster.Ward * Major bug when importing surfaces for an MRI that was re-oriented manually * Smooth surface: Fix little spikes due to irregularities in the mesh * Surface>Volume interpolation: Use spm_mesh_to_grid * Bug: Hide scouts in the preview of the grid for volume head models == ECOG/SEEG == * SEEG: Project contacts to SEEG / Compare with IntrAnat * Contact positions: Import / set / detect<<BR>> * New option: Align on none|inner|cortex to replace ECOG-mid * Add history: Save modifications and transformations applied to the channel files (Marcel) * Project contact positions across subjects or templates (Marcel) * ECOG: Default names of contacts? * Add menu to import implantation channel file in imported recordings * Automatic segmentation of CT: * GARDEL: http://meg.univ-amu.fr/wiki/GARDEL:presentation * Arnulfo: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6 * MAP07 / SPM: https://www.epi.ch/_files/Artikel_Epileptologie/Huppertz_2_13.pdf * ECOG: * Project and display contacts on cortex surface should consider the rigidity of the grids: Contacts cannot rotate, and distance between contacts should remain constant across runs * Method for contacts projection: https://pdfs.semanticscholar.org/f10d/6b899d851f3c4b115404298d7b997cf1d5ab.pdf * ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage) * Display: * Bad channels: Contacts greyed out instead of ignored (Marcel) * Display time in H:M:S * Display curved SEEG electrodes * Re-referencing: * Create new average reference montages with a specific list of channels, with the possibility to edit the order of the channels (for Jeremy) * Closest white reference (Arnulfo) * Detection CEEP stim artifacts: Use ImaGIN code ImaGIN_StimDetect * Alternatives to OpenMEEG: SimBio/FieldTrip? Matti Stenroos? NFT/NIST? |
| Line 151: | Line 333: |
| * ANOVA: Use LENA functions | * ANOVA: * Which functions to use? * Write panel similar to Process1 and Process2 to allow the |
| Line 154: | Line 338: |
| * Permutation tests: * t-test only (wilcoxon? sign-test?): paired, equal var, unequal var * http://www.adscience.fr/uploads/ckfiles/files/html_files/StatEL/statel_wilcoxon.htm * http://www.mathworks.fr/fr/help/stats/signrank.html * Less powerful than t-tests * nb permutations ~ 1000 * maximum statistic over "time" or "time and space" * Permutations / clustering: cf fieldtrip * http://fieldtrip.fcdonders.nl/tutorial/cluster_permutation_timelock * http://fieldtrip.fcdonders.nl/tutorial/cluster_permutation_freq * Threshold in time: keep only the regions that are significative for contiguous blocks of time, or over a certain number of time points<<BR>> => Process that creates a static representation of a temporal window * t-test on volume sources * Paired t-test on unconstrained sources: (convert to flat + Z-score) => !AnneSo * Question of Gaussianity of the samples: take a subset of samples + Kolmogorov-Smirnov / Shapiro-Wilk test * http://fr.wikipedia.org/wiki/Test_de_Shapiro-Wilk * http://stats.stackexchange.com/questions/362/what-is-the-difference-between-the-shapiro-wilk-test-of-normality-and-the-kolmog * http://www.mathworks.fr/fr/help/symbolic/mupad_ug/perform-shapiro-wilk-test.html * http://www.mathworks.fr/fr/help/symbolic/mupad_ref/stats-swgoft.html * http://stackoverflow.com/questions/14383115/shapiro-wilk-test-in-matlab * Create icons for Stat/PAC, Stat/Sprectrum, etc. * One sample t-test across subjects |
* New process to test for Gaussianity using swtest * Simulate recordings with specific properties, for stat validation * Quality control before statistics, on condition averages across subjects:<<BR>>mean(baseline)/std(baseline): shows bad subject quickly. * Use SurfStat: Impements interesting things, like an analytical cluster-based p-value correction (Random-field theory which is used in SPM) - Peter * Export to R or SPSS for advanced stat |
| Line 177: | Line 346: |
| * '''XDF import''': Use the EEGLAB plugin, contact Martin Bleichner (Oldenburg) * Output .nii have incorrect sform/qform when using the options to downsample the volume of cut the empty slices * DICOM converter: * Add dcm2nii (MRICron) * Add MRIConvert * FieldTrip: Import/Export time-frequency: * Export: http://neuroimage.usc.edu/forums/t/export-time-frequency-to-fieldtrip/1968 * Import: http://neuroimage.usc.edu/forums/t/import-time-frequency-data-from-fieldtrip/2644 * 4D file format: * Use reader from MNE-Python: mne.io.read_raw_kit (doesn't require Yokogawa slow library) * Reference gradiometers: Keep the orientation of the first or second coil? * Reference gradiometers: Add the sensor definition from coil_def.dat * Validate with phantom recordings that noise compensation is properly taken into account * References at too far from the head sensors in Marseille 4D system * The noise compensation is considered to be always applied on the recordings, not sure this assumption is always correct * 4D phantom tutorial (JM Badier?) |
|
| Line 178: | Line 363: |
| * EEG !CeeGraph | * EEG CeeGraph |
| Line 180: | Line 365: |
| * !FieldTrip structures: In / Out | * XLTEK: https://github.com/danielmhanover/OpenXLT * Persyst .lay: https://github.com/ieeg-portal/Persyst-Reader * Nervus .eeg: https://github.com/ieeg-portal/Nervus-Reader * Biopac .acq: https://github.com/ieeg-portal/Biopac-Reader * gTec EEG recordings: Read directly from the HDF5 files instead of the Matlab exports. |
| Line 182: | Line 371: |
| * Export TF maps to SPM / volumes * EEGLAB import: Selection of conditions in script mode |
* EEGLAB import: * Support for binary AND epoched files (now it's one or the other) * Allow epoched files with recordings saved in external files * BST-BIN: Add compressionto .bst * Review raw on all the file formats (ASCII EEG and Cartool missing) * SPM .mat/.dat: Fix the import of the EEG/SEEG coordinates * Get acquisition date from files: Missing for 4D |
| Line 186: | Line 380: |
| * Add Help buttons and menus (in popups, dialog windows...) => Links to the website. * Introduction tutorials: * Processes: Describe all the processes * Clusters * First steps: Brainstorm preferences * Headmodel: explain the fields + how to get the constrained leadfield * Sources: Modelized data * Sources: theshold min. size (not documented yet) * Import raw recordings: Add "detect bad trials/channels" in the pipeline * Temporary folder |
* Tutorial OMEGA/BIDS: * Add review of literature for the resting state MEG * Download example datasets directly from the OMEGA repository * New tutorials: <<BR>> * Other public datasets: [[https://github.com/INCF/BIDS-examples/tree/bep008_meg|https://github.com/INCF/BIDS-examples/tree/bep008_meg/]] * Rat PAC + high gamma (Soheila) * EEG/research * FieldTrip ECOG tutorial: http://www.fieldtriptoolbox.org/tutorial/human_ecog * FieldTrip cortico-muscular coherence tutorial: http://www.fieldtriptoolbox.org/tutorial/coherence * Reproduce tutorials from MNE-Python: https://martinos.org/mne/stable/tutorials.html * Cam-CAN database: https://camcan-archive.mrc-cbu.cam.ac.uk/dataaccess/<<BR>>(download new datasets, including maxfiltered files and manual fiducial placements) * MEG steady-state / high-gamma visual / frequency tagging * BIDS-EEG example datasets * Stand-alone ICA tutorial * Move all the files to download to the cloud for faster download everywhere in the world * Provide secure way of sending password over HTTPS for: * Account creation * Forum exchanges * org.brainstorm.dialog.CloneControl * Workflows FieldTrip: http://www.fieldtriptoolbox.org/faq/what_types_of_datasets_and_their_respective_analyses_are_used_on_fieldtrip * Count GitHub clones in the the download stats * Google Analytics: Create template and update the section of the Community page * Deface the MRIs of all the tutorials * Clean up the wiki: * Remove all the wiki pages that are not used * Check all the links in all the pages * Check that all the TODO blocks have been properly handled * Remove useless images from all tutorials * Update page count on the main tutorials page |
| Line 198: | Line 412: |
| * Import anatomy folder menu crashes on MacOSX 10.8.5 / Matlab 2013b * Record tab: Text of epoch number is too big on MacOS * Menu "Use default EEG cap" doesn't work for a multiple selection (setting the same EEG cap for several subjects) |
* Image viewer: * Difficult to get to 100% * Buggy on some systems * 2DLayout: * (TF) Units are weird with % values * (TF) Difficult to navigate in frequencies: Scaling+changing frequency resets the scaling * Progress bar: * Doesn't close properly on some Linux systems * Focus requests change workspace when processing constantly (Linux systems) * MacOS bugs: * Buttons {Yes,No,Cancel} listed backwards * Record tab: Text of epoch number is too big * Colormap menus: Do not work well on compiled MacOSX 10.9.5 and 10.10 * Event markers are not visible anymore with the sequence: Open MEG, open EOG, close MEG |
| Line 202: | Line 427: |
| * tree_dependencies: sources files, reprojected on default anatomy; If based on data files that are bad trials, they should be ignored by tree_dependencies, and they are not * Image viewer has some bugs on some systems * Screen capture for reports never works: Find another solution * Screen capture when there is a fading effect in the window manager: captures the window * Close figure with coherence results should hide the "frequency" slider * Problems growing scouts on merged surfaces (Emily) |
|
| Line 209: | Line 428: |
| * Canolty maps computation: Fix progress bar | |
| Line 211: | Line 431: |
| * Use Matlab Coder to optimize some processes: Bandpass filter, sinusoid removal | * bst_bsxfun: After 2016b, we can use directly the scalar operators (./ .* ...) instead of bsxfun. Update bst_bsxfun to skip the use of bsxfun when possible. * Interface scaling: Rewrite class IconLoader to scale only once the icons at startup instead of at each request of an icon (might improve the speed of the rendering of the tree) |
| Line 213: | Line 434: |
| * mri2scs: convert arguments to meters | * Processes with "radio" and "radio_line" options: Replace with "radio_label" and "radio_linelabel" * Interpolations: Use scatteredInterpolant, griddedInterpolant, triangulation.nearestNeighbor (2014b) |
| Line 215: | Line 437: |
| * Shared kernels: do the "get bad channels" operation in a different way (reading all the files is too slow) | * Shared kernels: "get bad channels" operation in a different way (reading all the files is too slow) |
| Line 219: | Line 441: |
| * Progress bar: * Add different levels (to handle sub-processes) * Make work correctly with RAW on resting tutorial * Uniformize calls in bst_process/Run * Add a "Cancel" button * Line smoothing / anti-aliasing (time series figures) |
* Fix all the 'todo' blocks in the code |
What's next
A roadmap to the future developments of Brainstorm.
Contents
Recordings
RAW file viewer:
- Bottom bar: display extend events as segments instead of dots (to see their extension)
- Downsample before filtering? (attention to the filter design)
- Add parameter to make the visual downsampling more or less aggressive
- Pre-load next page of recordings
- Keep the filter specifications in memory instead of recomputing for every page
- Add field "comment" to markers: For clinicians to add notes (Marcel)
- Events: Change the category of a selected event easily, instead of deleting/marking new
Events: Advanced process for recombining.
Example: http://www.erpinfo.org/erplab/erplab-documentation/manual/Binlister.html
- Bad trials: When changing the status of bad to good: remove the bad segments as well, otherwise it is not processed by processes like the PSD.
- Review clinical recordings: Reduce the dimensionality of the data with a simple inverse problem, similar to what we do for the magnetic extrapolation ("Regional sources" in BESA, cf S Rampp)
- MEG/EEG registration: Apply the same transformation to multiple runs
- 2DLayout:
- Does not work when DC offset is not removed
- Add a proper amplitude scale that gets updated when shift+scroll, to compare figures
- Create heat maps: Maybe with matlab function heatmap?
Interface
Add a warning when computing a forward model with > 100000 sources (check selection)
- Snapshot: Save as image / all figures (similar to Movie/all figure)
Generalize the use of the units (field .DisplayUnits): Rewrite processes to save the units correctly
- Colormaps:
- Allow brightness/contrast manipulations on the custom colormaps
- Global colormap max: Should get the maximum across all the open files
- Copy figures to clipboard (with the screencapture function)
- Display warning before opening files that are too big
- Smooth display from figure_image (ERPimage, raster plot...)
Contact sheets & movies: use average of time windows instead of single instants, for each picture.
- Contact sheets: Allow explicit list of times in input (+ display as in MNE-Python with TS)
- Display CTF coils: Show discs instead of squares
- Use boundary() instead of conhull() in all the display functions (ie. 2DDisc)
- Progress bar:
- Add different levels (to handle sub-processes)
- Uniformize calls in bst_process/Run
- Add a "Cancel" button
- Error message: Add a link to report directly the bug on the forum
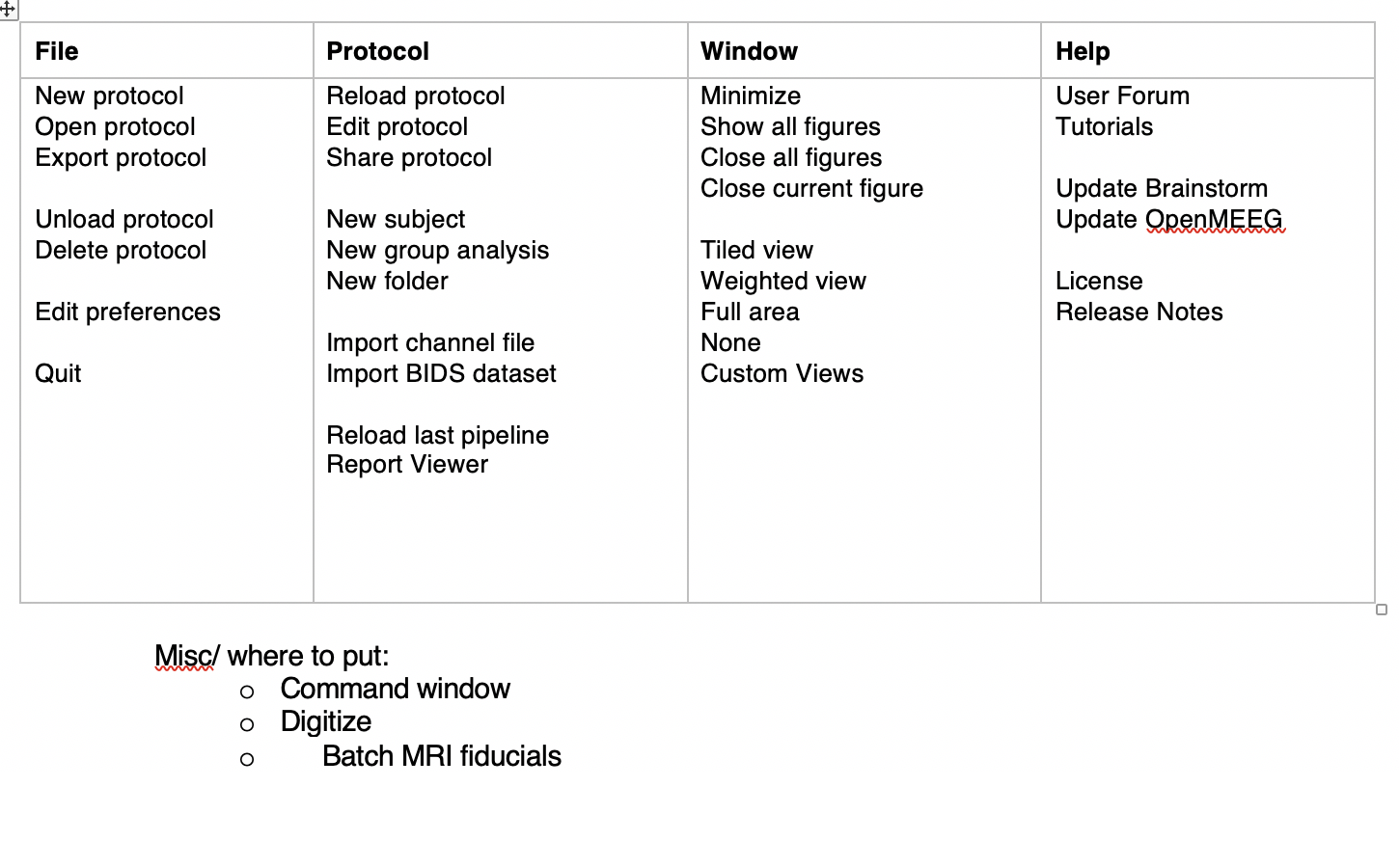
Reorganize menus (Dannie's suggestion):
Connectivity
- Thresholding and stat tests the connectivity matrices
- Connectivity on unconstrained sources: "Default signal extraction for volume grids should be the time series of the first principal component of the triplet signals after each has been zero-meaned" (SB)
- Display of connectivity graphs:
- Display as straight lines
- Recode 2D graphs
- 3D display with anatomical constrains
- Display using real position of EEG electrodes
- Use new band-pass filters in bst_connectivity ('bst-hfilter' instead of 'bst-fft-fir')
Review by Jan-Mathijs: http://journal.frontiersin.org/article/10.3389/fnsys.2015.00175/full
- Connectivity based on band limited power (Sylvain):
- Compute Hilbert/Bandpass + correlation of the envelopes
- Bandpass envelopes before computing correlations?
- Compute Hilbert(sensors) and then project to source space?
- Matrix view of NxN graphs: Add legend of the elements along X and Y axis
- Graph view:
- Does not display negative values correctly (correlation or difference of coherence)
- Re-write using pure Matlab code and smoothed graphics
- Fixed scales for intensity sliders
- Text bigger
- Too much data in appdata
- Fixed scales for intensity sliders
- Add "=" shortcut for having graphs with similar configurations
- Disable zoom in one region (serious bugs)
- NxN on sensors: does not place the sensors correctly in space
- Coherence:
Average cross-spectra instead of concatenating epochs (to avoid discontinuities)
Explore inter-trial approaches (Esther refers to chronux toolbox)
- Granger:
- Crashes sometimes: improve stability
- Check for minimum time window (Esther: min around 500-1000 data points)
- Re-write and optimize code
- Add progress bar
- PLV:
- Add p-values
- Remove evoked
- Optimize code
- Add time integration
- Unconstrained sources
- Add warning when running of short windows (because of filters)
- PAC:
- Add input TF , to disconnect TF decomposition and PAC computation (Peter)
- Refine frequency vector of low frequencies
- How many central frequencies to use in bst_pac?
- Change filters: no chirplet functions
- bst_freqfilter: Use nfcomponents like in bst_pac
- Esther recommended a larger frequency binning of the PAC estimation
- PAC maps: Display all sensors at once (like TF and DynamicPAC)
- Hui-Ling's PAC:
- Time-resolved correlation/coherence: Display as time bands
- Other metrics:
- Coherence by bands: bst_coherence_band_welch.m
- Granger by bands: bst_granger_band.m
- Inter-trial coherence
Tutorial coherence [1xN] : Reproduce FieldTrip results?
Connect NxN: Display as time series > Display warning before trying to open too many signals
Processes
- Decoding/Classifiers: Implement Dimitrios and scikit-learn algorithms
- Allow processes in Python and Java
- Add MNE-Python functions:
- scikit-learn classifiers
- Implement data exchange with MNE-Python: write FIF files from Brainstorm and/or pass python objects in memory instead of FIF files
- SSS/tSSS cleaning
- Reproduce other tutorials / examples
- Change the graphic renderer from Matlab
Add FieldTrip functions:
- ft_sourceanalysis:
- Check noise covariance
- Check all the options of all the methods
- Single trial reconstructions + noise covariance?
Filters?? http://www.fieldtriptoolbox.org/example/common_filters_in_beamforming
Beamformers: Save ftSource.avg.mom
http://www.fieldtriptoolbox.org/workshop/meg-uk-2015/fieldtrip-beamformer-demohttp://www.fieldtriptoolbox.org/tutorial/beamformingextended
- Baseline? Two inputs?
ft_prepare_sourcemodel: Compute MNI transformation (linear and non-linear) => Peter
- Freqanalysis: ITC
- ft_read_atlas('TTatlas+tlrc.BRICK');
ft_volumereslice: http://www.fieldtriptoolbox.org/faq/how_change_mri_orientation_size_fov
- ft_freqanalysis
- ft_combineplanar
- ft_sourceanalysis:
- Use CUDA for speeding up some operations (filtering, wavelets, etc)
- Pipeline editor:
- Bug: After "convert to continuous", the time of the following processes should change
- Add loops over subjects/conditions/trial groups
- Events: Allow selection from a drop-down list (similar to option "channelname" in panel_process_selection)
ITC: Inter-trial coherence (see MNE reports for group tutorial)
http://www.sciencedirect.com/science/article/pii/S1053811916304232- ICA:
- Why doesn't the ICA process converge when using 25 components in the EEG tutorial?
- Add an option to resample the signals before computing the ICA decomposition
- Add a stand-alone tutorial
- Exploration: Add window with spectral decomposition (useful for muscle artifacts)
- Export IC time series (and then compute their spectrum): solves the problem above
Comparison JADE/Infomax:
http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0030135- Use faster methods (MNE-Python?)
- Add methods: SOBI, Fastica, AMICA/CUDICA (recommended by S Makeig)
Dimension reduction with PCA adds artifacts: Not done by default in EEGLAB
Contact: Stephen Shall Jones ( shall-jones@infoscience.otago.ac.nz )
Student Carl Leichter detailed this in his thesis- S Makeig: Use ICA to select the IC of interest instead of only removing artifacts
- Display of spectrum for components (PSD/FFT)
- Use FastICA (algo crashing)
- Add components preselection: Correlation with EOG/ECG
- Import ICA matrices available in EEGLAB .set files
EEGLAB recommends ICA + trial rejection + ICA again: Impossible right now with Brainstorm
(http://sccn.ucsd.edu/wiki/Chapter_09:_Decomposing_Data_Using_ICA)ICA+machine learning: https://www.ncbi.nlm.nih.gov/pubmed/28497769
Automated artifact rejection: https://arxiv.org/abs/1612.08194
- Save IC time series in database
Use EYE-EEG: EEGLAB toolbox for eye-tracker guided ICA (Olaf Dimigen): http://www2.hu-berlin.de/eyetracking-eeg/
- Other EEGLAB functions:
Step function detection: https://github.com/lucklab/erplab/wiki/Artifact-Detection:-Tutorial
- SSP:
Display warning if changing the ChannelFlag while there is a Projector applied
- Pipeline editor:
- When computing sources from the pipeline editor: doesn't reselect the options if you click twice on "edit" (works for minnorm, but not for lcmv)
- When computing time-frequency/hilbert/psd: Find a way not to force the user to click on Edit
- Bandpass:
- Use new filters in all the functions using a bandpass ('bst-hfilter' instead of 'bst-fft-fir'): process_evt_detect_threshold
- Weird bug: Filter(import) != Import(Filter) in the HCP tutorial... to investigate
- Spectral flattening (John):
- ARIMA(5,0,1): Apply on the signal before any frequency/connectivity/PAC analysis
- PSD:
- Rewrite to have the same input as coherence (frequency resolution instead of window length)
- Use the progress bar
Allow display of Avg+StdErr
Remove line noise: http://www.nitrc.org/projects/cleanline
- Use Matlab Coder to optimize some processes: Wavelets, bandpass filter, sinusoid removal
- Time-frequency:
- Optimization: bst_timefreq (around l.136), remove evoked in source space: Average should be computed in sensor space instead of source space (requested by Dimitrios)
Short-time Fourier transform: http://www.mikexcohen.com/lectures.html
Matching pursuit: http://m.jneurosci.org/content/36/12/3399.abstract?etoc
- Bug: Display logs as negative
- Bug: 3D figures: Colormaps with "log" option doesn't work
- Bug: Difference of power displayed in log: problems (Soheila)
- 2D Layout in spectrum
- Make much faster and more memory efficient (C functions coded by Matti ?)
- TF scouts: should display average of TF maps
- Impossible to keep complex values for unconstrained sources
- Pad short epochs with zero values for getting lower frequencies
- Hilbert with time bands very slow on very long files (eg. 3600s at 1000Hz) because the time vector is still full (10^7 values): save compressed time vector instead.
- Extend clusters tab to display of TF to overlay TF signals (Svet)
- When normalizing with baseline: Propagate with the edge effects marked in TFmask
- Allow baseline normalization of files computed with time bands
- Allow running TF on montages
- Review continuous files in time-frequency space (for epilepsy)
- Bug when computing TF on constrained and unconstrained scouts at the same time (in mixed head models for instance): uses only the constrained information and doesn't sum the 3 orientations for the unconstrained regions.
- Artifact detection:
- Artifact rejection like SPM: if bad in 20%, bad everywhere
- Test difference between adjacent samples
- Events detection: Add option "std" vs "amplitude"
- Simulation:
Fix units in simulation processes => no *1e-9 in "simulate recordings"
- Use "add noise" process from Hui-Ling (in Work/Dev/Divers)
- Use field process field "Group" to separate Input/Processing/Output options
- Use new Matlab functions: movmean, movsum, movmedian, movmax, movmin, movvar, movstd
- Process ft_prepare_heamodel: Add support from BEM surfaces from the Brainstorm database
Database
- Rename protocol
- Faster DB searches (for Emily)
- Add buttons to sort files: by name, by comment, by date
- MEG protocols: More flexible organization of the database; sub-conditions to allow different runs X different conditions.
- Matrix files: Allow to be dependent from other files
- Add notes in the folders (text files, visible as nodes in the tree)
- Screen captures: save straight to the database
- Rename multiple files
- Default headmodel lost when reloaded: Keep selection on the hard drive (in brainstormstudy.mat)
- Auto-save:
protocol.mat can be too big: do not store the results links in it (and recreate when loading)- http://neuroimage.usc.edu/forums/t/abnormally-slow-behavior/2065/10
- Improve auto-save: add tracking file next to protocol.mat, do not save all the time, only when closing app, and reload protocol at stratup if tracking file is still there
Distributed computing
Options from FieldTrip:
Loose collection of computers: https://github.com/fieldtrip/fieldtrip/tree/master/peer
Alternative, with less limitations: http://research.cs.wisc.edu/htcondor/
Single multicore machine: https://github.com/fieldtrip/fieldtrip/tree/master/engine
Batch system: https://github.com/fieldtrip/fieldtrip/tree/master/qsub
Documentation: http://fieldtrip.fcdonders.nl/faq#distributed_computing_with_fieldtrip_and_matlab
Various initiatives: http://samirdas.github.io/Data_sharing.html#/
Source modeling
- Dipoles:
- Project individual dipoles files on a template
- panel_dipoles: Doesn't work with multiple figures
- Project sources: Very poor algorithm to project sub-cortical regions and cerebellum (algorithm to fit surfaces should be imrpoved)
Menu head model > Copy to other conditions/subjects (check if applicable first)
Menu Sources > Maximum value: Doesn't work with volume or mixed head models
- Mixed head models:
- Set loose parameter from the interface
- Volume grid:
- Optimize: 3D display (better than 9x9 cubes)
- Optimize: vol_dilate (with 26 neighbors)
Menu Sources > Simulate recordings:
- Do not close the 3D figures after generating a new file
- Add a process equivalent to this menu
- Panel Get coordinates: Add button "find maximum"
- BEM single sphere: Get implementation from MNE
- Unconstrained sources:
- Stat and connectivity: what to do? (re-send email John+Sylvain)
- Overlapping spheres: improve the estimation of the spheres for the frontal lobes
- Noise covariance matrix:
- Display with figure_image()
- Storage of multiple noise covariance matrices (just like the head models)
- Always save as full, then at inversion time, we can decide between full, heteroskedastic (diagonal) or homoskedastic (i.i.d, scalar)
Problem of having inividual trials + averages in the condition => Display warning or not?
- Save nAvg in noisecov file, to make it easier to scale to other recordings
Sources on surface: Display peak regions over time (time = color) => A.Gramfort
- Time-frequency beamformers:
- Band-pass everything in different frequency bands + Source estimation + TF
- Ask data to Sarang where he sees effects that cannot be extracted with MN followed by TF
- Process "Extract scouts time series": Add PCA option (replace isnorm with choice PCA/Norm)
- Add eyes models to attract eye activity
Anatomy
- Segmentation with SPM12
BrainVISA: Add support for MarsAtlas
- MNI transformation: Use SPM non-linear MNI transformation y_...
- Registration:
Getting electrode positions from 3D scanners: https://sccn.ucsd.edu/wiki/Get_chanlocs
Use the same registration for multiple recording sessions that have already re-registered previously (eg. with MaxFilter)
- When linking multiple EEG recordings including 3D positions, do the registration only once and copy it to all the runs
- Compute non-linear MNI registration instead of linear
- Select and remove bad digitized head points before automatic coregistration
Load the MNE -transf.fif: http://neuroimage.usc.edu/forums/showthread.php?2830
- MRI Viewer:
- Pan in zoomed view (shift + click + move?)
- Zoom in/out with mouse (shift + scroll?)
- Ruler tool to measure distances
- Display scouts as additional volumes
- Render surface envelope in the MRI as a thin line instead of the full interpolation matrix
- Edit fiducials: Replace 6 text boxes with 1 for easy copy-paste (see fiducials.m)
Optimize computation interpolation MRI-surface (tess_tri_interp) => spm_mesh_to_grid
BrainSuite:
Add new labels to all BrainSuite anatomy templates
Use BrainSuite inner skull for surface generation
- Use same colors for left and right for anatomical atlases
- Scouts:
- Display edges in the middle of the faces instead of the vertices
- Display scouts in a tree: hemisphere, region, subregion
- Sort scouts by region in process options
- Downsample to atlas: allow on timefreq/connect files
Project from one hemisphere to the other using registered spheres/squares (http://neuroimage.usc.edu/forums/t/how-to-create-mirror-roi-in-the-other-hemisphere/5910/8)
- Parcellating volume grids: scikit-learn.cluster.Ward
- Major bug when importing surfaces for an MRI that was re-oriented manually
- Smooth surface: Fix little spikes due to irregularities in the mesh
Surface>Volume interpolation: Use spm_mesh_to_grid
- Bug: Hide scouts in the preview of the grid for volume head models
ECOG/SEEG
SEEG: Project contacts to SEEG / Compare with IntrAnat
Contact positions: Import / set / detect
- New option: Align on none|inner|cortex to replace ECOG-mid
- Add history: Save modifications and transformations applied to the channel files (Marcel)
- Project contact positions across subjects or templates (Marcel)
- ECOG: Default names of contacts?
- Add menu to import implantation channel file in imported recordings
- Automatic segmentation of CT:
- ECOG:
- Project and display contacts on cortex surface should consider the rigidity of the grids: Contacts cannot rotate, and distance between contacts should remain constant across runs
Method for contacts projection: https://pdfs.semanticscholar.org/f10d/6b899d851f3c4b115404298d7b997cf1d5ab.pdf
ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage)
- Display:
- Bad channels: Contacts greyed out instead of ignored (Marcel)
- Display time in H:M:S
- Display curved SEEG electrodes
- Re-referencing:
- Create new average reference montages with a specific list of channels, with the possibility to edit the order of the channels (for Jeremy)
- Closest white reference (Arnulfo)
Detection CEEP stim artifacts: Use ImaGIN code ImaGIN_StimDetect
Alternatives to OpenMEEG: SimBio/FieldTrip? Matti Stenroos? NFT/NIST?
Statistics
- ANOVA:
- Which functions to use?
- Write panel similar to Process1 and Process2 to allow the
- Output = 1 file per effect, all grouped in a node "ANOVA"
- Display several ANOVA maps (from several files) on one single figure, using a "graphic accumulator", towards which one can send any type of graphic object
- New process to test for Gaussianity using swtest
- Simulate recordings with specific properties, for stat validation
Quality control before statistics, on condition averages across subjects:
mean(baseline)/std(baseline): shows bad subject quickly.Use SurfStat: Impements interesting things, like an analytical cluster-based p-value correction (Random-field theory which is used in SPM) - Peter
- Export to R or SPSS for advanced stat
Input / output
XDF import: Use the EEGLAB plugin, contact Martin Bleichner (Oldenburg)
- Output .nii have incorrect sform/qform when using the options to downsample the volume of cut the empty slices
- DICOM converter:
- Add dcm2nii (MRICron)
- Add MRIConvert
FieldTrip: Import/Export time-frequency:
- 4D file format:
- Use reader from MNE-Python: mne.io.read_raw_kit (doesn't require Yokogawa slow library)
- Reference gradiometers: Keep the orientation of the first or second coil?
- Reference gradiometers: Add the sensor definition from coil_def.dat
- Validate with phantom recordings that noise compensation is properly taken into account
- References at too far from the head sensors in Marseille 4D system
- The noise compensation is considered to be always applied on the recordings, not sure this assumption is always correct
- 4D phantom tutorial (JM Badier?)
- EEG File formats:
EEG CeeGraph
- EGI: Finish support for epoched files (formats 3,5,7)
Persyst .lay: https://github.com/ieeg-portal/Persyst-Reader
Nervus .eeg: https://github.com/ieeg-portal/Nervus-Reader
Biopac .acq: https://github.com/ieeg-portal/Biopac-Reader
- gTec EEG recordings: Read directly from the HDF5 files instead of the Matlab exports.
- BCI2000 Input (via EEGLAB plugin)
- EEGLAB import:
- Support for binary AND epoched files (now it's one or the other)
- Allow epoched files with recordings saved in external files
- BST-BIN: Add compressionto .bst
- Review raw on all the file formats (ASCII EEG and Cartool missing)
- SPM .mat/.dat: Fix the import of the EEG/SEEG coordinates
- Get acquisition date from files: Missing for 4D
Distribution & documentation
- Tutorial OMEGA/BIDS:
- Add review of literature for the resting state MEG
- Download example datasets directly from the OMEGA repository
New tutorials:
Other public datasets: https://github.com/INCF/BIDS-examples/tree/bep008_meg/
- Rat PAC + high gamma (Soheila)
- EEG/research
FieldTrip ECOG tutorial: http://www.fieldtriptoolbox.org/tutorial/human_ecog
FieldTrip cortico-muscular coherence tutorial: http://www.fieldtriptoolbox.org/tutorial/coherence
Reproduce tutorials from MNE-Python: https://martinos.org/mne/stable/tutorials.html
Cam-CAN database: https://camcan-archive.mrc-cbu.cam.ac.uk/dataaccess/<<BR>>(download new datasets, including maxfiltered files and manual fiducial placements)
- MEG steady-state / high-gamma visual / frequency tagging
- BIDS-EEG example datasets
- Stand-alone ICA tutorial
- Move all the files to download to the cloud for faster download everywhere in the world
- Provide secure way of sending password over HTTPS for:
- Account creation
- Forum exchanges
org.brainstorm.dialog.CloneControl
Workflows FieldTrip: http://www.fieldtriptoolbox.org/faq/what_types_of_datasets_and_their_respective_analyses_are_used_on_fieldtrip
Count GitHub clones in the the download stats
- Google Analytics: Create template and update the section of the Community page
- Deface the MRIs of all the tutorials
- Clean up the wiki:
- Remove all the wiki pages that are not used
- Check all the links in all the pages
- Check that all the TODO blocks have been properly handled
- Remove useless images from all tutorials
- Update page count on the main tutorials page
Current bugs
- Image viewer:
- Difficult to get to 100%
- Buggy on some systems
- 2DLayout:
- (TF) Units are weird with % values
- (TF) Difficult to navigate in frequencies: Scaling+changing frequency resets the scaling
- Progress bar:
- Doesn't close properly on some Linux systems
- Focus requests change workspace when processing constantly (Linux systems)
- MacOS bugs:
- Buttons {Yes,No,Cancel} listed backwards
- Record tab: Text of epoch number is too big
- Colormap menus: Do not work well on compiled MacOSX 10.9.5 and 10.10
- Event markers are not visible anymore with the sequence: Open MEG, open EOG, close MEG
- in_bst_data_multi: If trials have different sizes, output is random (the one of the first file)
- Edit scout in MRI: small modifications cause huge increase of the scout size
- Canolty maps computation: Fix progress bar
Geeky programming details
- bst_bsxfun: After 2016b, we can use directly the scalar operators (./ .* ...) instead of bsxfun. Update bst_bsxfun to skip the use of bsxfun when possible.
Interface scaling: Rewrite class IconLoader to scale only once the icons at startup instead of at each request of an icon (might improve the speed of the rendering of the tree)
- Hide Java panels instead of deleting them
- Processes with "radio" and "radio_line" options: Replace with "radio_label" and "radio_linelabel"
- Interpolations: Use scatteredInterpolant, griddedInterpolant, triangulation.nearestNeighbor (2014b)
- bst_warp and channel_project: Use tess_parametrize_new instead of tess_parametrize
- Shared kernels: "get bad channels" operation in a different way (reading all the files is too slow)
- Optimize bst_get:
- Now study and subject have necessarily the same folder name
- Replace big switch with separate functions
- Fix all the 'todo' blocks in the code
