|
Size: 871
Comment:
|
Size: 2288
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| == MEG Auditory evoked potential == Acquisition system: ''Neuromag Vectorview306''. Description of the windows: * Main !BrainStorm window * Timeseries of all the MEG sensors [-200ms, 500ms] * Magnetic field recorded by the magnetometers at t=106ms * Spatial view of the magnetometers time series [-200ms, 500ms] * Reconstruction of the cortical currents, based on the magnetometers, at t=106ms [[attachment:snap_1condition.jpg|{{attachment:snap_1condition_sm.jpg|attachment:snap_1condition.jpg}}]] |
|
| Line 5: | Line 18: |
| || {{attachment:snap_1condition.jpg||height="373",width="534"}} ||<style="vertical-align: top;">'''MEG auditory evoked potential'''<<BR>> (Neuromag Vectorview306)<<BR>> <<BR>> Description of the windows (left to right): <<MiniPage[[MiniPage(* Main BrainStorm window * Timeseries of all the MEG sensors [-200ms, 500ms] * Magnetic field recorded by the magnetometers at t=106ms * Spatial view of the magnetometers time series [-200ms, 500ms] * Reconstruction of the cortical currents, based on the magnetometers, at t=106ms)|(* Main BrainStorm window * Timeseries of all the MEG sensors [-200ms, 500ms] * Magnetic field recorded by the magnetometers at t=106ms * Spatial view of the magnetometers time series [-200ms, 500ms] * Reconstruction of the cortical currents, based on the magnetometers, at t=106ms)]]>> || | |
| Line 7: | Line 19: |
| == Multiple conditions: Baby auditory EEG == Acquisition system: ''EGI GSN - 64 electrodes'' |
|
| Line 8: | Line 22: |
| Description: | |
| Line 9: | Line 24: |
| * One subject: "001" * Three conditions: "GM", "GMM", "VM" * Two views: overlaid electrodes time series, and estimated cortical sources at t=376ms |
|
| Line 10: | Line 28: |
| ---- | [[attachment:snap_3conditions.jpg|{{attachment:snap_3conditions_sm.jpg|attachment:snap_3conditions.jpg}}]] |
| Line 12: | Line 31: |
== Subject anatomy: MRI and surfaces == BrainStorm offers the possibility to reconstruct the cortical activity either on the individual subject anatomy, or on a default anatomy (MNI / Colin27). For this purpose, many interactive tools are available to view, register and process the MR images and the corresponding meshes. However, the cortex segmentation must be performed by external program of your choice ([[Links|list here]]). Here are a few examples of the views you can obtain with a simple click on the subject's MRI or surface. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surfaces panel" and with their popup menu. The MRI slices can be moved with a simple mouse operation: right-click and mouse drag. [[attachment:snap_anatomy.jpg|{{attachment:snap_anatomy_sm.jpg|attachment:snap_anatomy.jpg}}]] <<BR>> == Channel selection == All the figures displayed with BrainStorm are always linked in time; and if they represent the same datasets, the sensors selection is also the same for all the views. Selection of a channel of data is done by clicking on it, in a time series or a 3D view. Selected sensors can be displayed separately, marked as "bad", or deleted. [[attachment:snap_channel.jpg|{{attachment:snap_channel_sm.jpg|attachment:snap_channel.jpg}}]] <<BR>> |
Screenshots
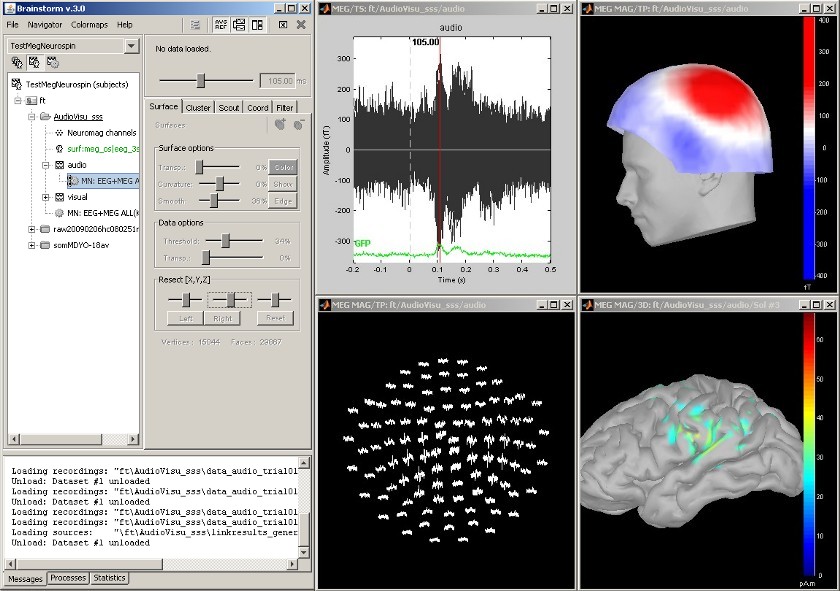
MEG Auditory evoked potential
Acquisition system: Neuromag Vectorview306.
Description of the windows:
Main BrainStorm window
- Timeseries of all the MEG sensors [-200ms, 500ms]
- Magnetic field recorded by the magnetometers at t=106ms
- Spatial view of the magnetometers time series [-200ms, 500ms]
- Reconstruction of the cortical currents, based on the magnetometers, at t=106ms
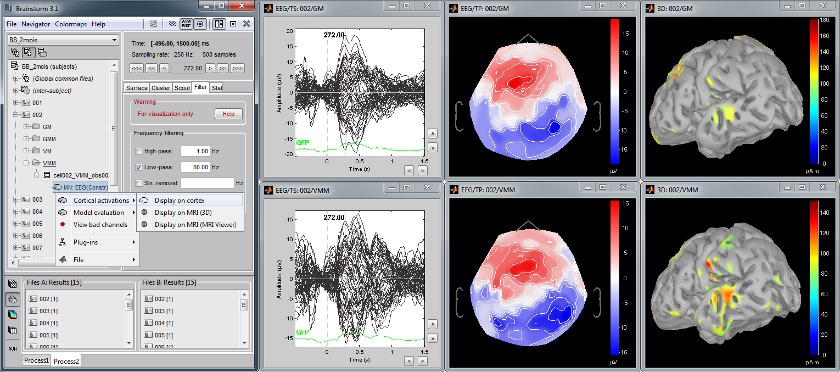
Multiple conditions: Baby auditory EEG
Acquisition system: EGI GSN - 64 electrodes
Description:
- One subject: "001"
- Three conditions: "GM", "GMM", "VM"
- Two views: overlaid electrodes time series, and estimated cortical sources at t=376ms
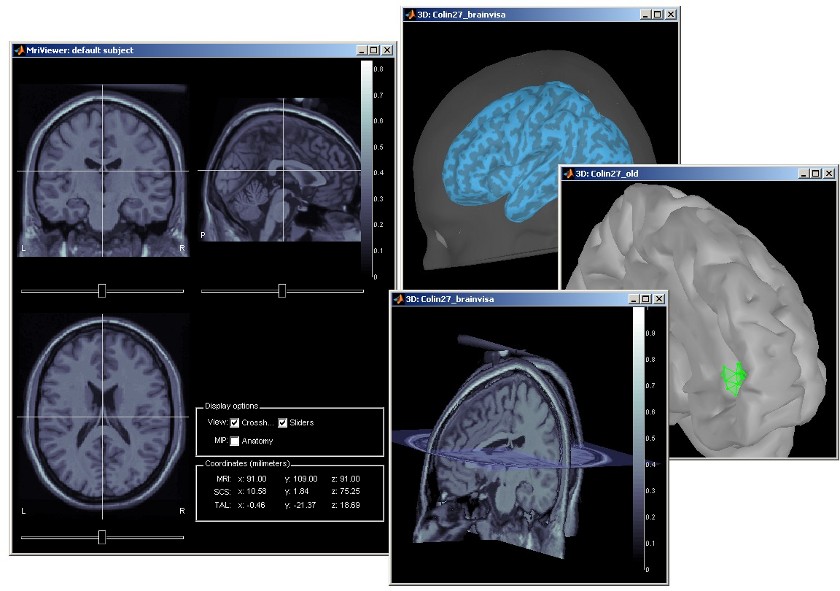
Subject anatomy: MRI and surfaces
BrainStorm offers the possibility to reconstruct the cortical activity either on the individual subject anatomy, or on a default anatomy (MNI / Colin27). For this purpose, many interactive tools are available to view, register and process the MR images and the corresponding meshes. However, the cortex segmentation must be performed by external program of your choice (?list here).
Here are a few examples of the views you can obtain with a simple click on the subject's MRI or surface. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surfaces panel" and with their popup menu. The MRI slices can be moved with a simple mouse operation: right-click and mouse drag.
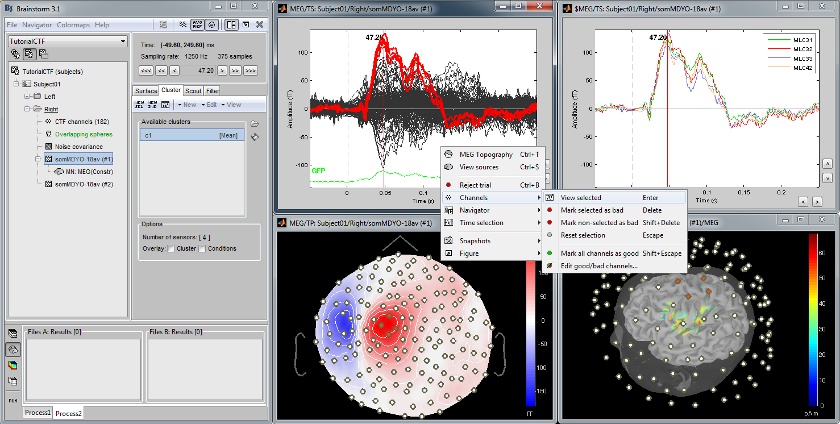
Channel selection
All the figures displayed with BrainStorm are always linked in time; and if they represent the same datasets, the sensors selection is also the same for all the views. Selection of a channel of data is done by clicking on it, in a time series or a 3D view. Selected sensors can be displayed separately, marked as "bad", or deleted.