|
Size: 6779
Comment:
|
Size: 6839
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| <<TableOfContents>> | == MEG somatosensory evoked responses == Acquisition on an ''CTF 275'' instrument for a left median nerve electric stimulation. |
| Line 4: | Line 5: |
| == MEG Auditory evoked potential == Acquisition system: ''Neuromag Vectorview306''. Description of the windows: * Main Brainstorm window * Timeseries of all the MEG sensors [-200ms, 500ms] * Magnetic field recorded by the magnetometers at t=106ms * Spatial view of the magnetometers time series [-200ms, 500ms] * Reconstruction of the cortical currents, based on the magnetometers, at t=106ms [[attachment:snap_1condition.jpg|{{attachment:snap_1condition_sm.jpg|attachment:snap_1condition.jpg}}]] |
[[attachment:snap_median.jpg|{{attachment:snap_median_sm.jpg|attachment:snap_median.jpg}}]] |
| Line 19: | Line 9: |
| == Database and right click == The tree in the main Brainstorm window represents the database for the selected study. This database has three levels of definition: Protocol (ie. study, selected in the toolbar), Subject, and Condition. Almost all the operations that can be performed on a file are accessible from the popup menu which is displayed by right clicking on the file. |
== Keeping your data organized and accessible == The tree in the main Brainstorm window represents the database for the selected study. This database has three levels of definition: Protocol (ie. study, selected in the toolbar), Subject, and Condition. Most of the operations that can be performed on a file are easily accessible from the popup menu revealed by right-clicking over the file. |
| Line 24: | Line 14: |
| * Anatomy: display the MRI and surfaces for each subject * Functional data (sorted by subject): sensors definition, recordings, sources, statistic results * Functional data (sorted by condition): idem, but sorted in a different way |
* Anatomy: display the MRI and surfaces for each participant in the study * Functional data (sorted by subject): sensor definitions, MEG and EEG data, source models, statistic and time-frequency maps * Functional data (sorted by condition): same as above, but sorted in a different way |
| Line 28: | Line 18: |
| The following example shows the MEG+EEG protocol "''Catching''", sorted by conditions. There are two experimental conditions, ''Catch ''and ''!NoCatch'', and seven subjects per condition. The popup menu shows all the actions that are available for the recordings of subject ''cc'', condition ''Catch''. | The following example features a MEG+EEG protocol called "''Catching''", sorted by conditions. There are two experimental conditions, ''Catch ''and ''NoCatch'', and 7 subjects per condition. The popup menu shows all the actions that are available for the recordings of subject ''cc'', condition ''Catch''. |
| Line 34: | Line 24: |
| == Multiple conditions: Baby auditory EEG == | == Multiple conditions: Baby auditory EEG responses == |
| Line 48: | Line 38: |
| Brainstorm offers the possibility to reconstruct the cortical activity either on the individual subject anatomy, or on a default anatomy (MNI / Colin27). For this purpose, many interactive tools are available to view, register and process the MR images and the corresponding meshes. However, the cortex segmentation must be performed by an external program of your choice ([[Links|list here]]). | Brainstorm features the possibility to model MEG and EEG neural generators either from the individual subject anatomy, or by using a template anatomy (MNI / Colin27) that can be warped to the individual scalp surface. Multiple interactive tools are available to view, register and process the MR images and the corresponding tessellated envelopes. However, tissue segmentation must be performed using another software; multiple options exist today in the academic community ([[Links|listed here]]). |
| Line 50: | Line 40: |
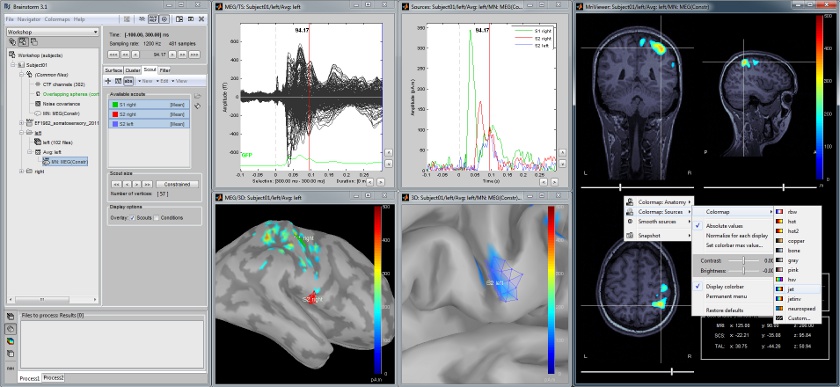
| Here are a few examples of the views you can obtain with a simple click on a MRI or a surface. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surfaces panel" and with their popup menus. The MRI slices can be moved with a simple mouse operation: right-click and mouse drag. | We provide a few examples of the views you can easily obtain with Brainstorm. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surface panel" and contextual popup menus. The MRI slices can be browsed with a simple mouse operation: right-click and mouse drag. |
| Line 57: | Line 47: |
| All the figures displayed with Brainstorm are always linked in time; and if they represent the same dataset, the sensors selection is also the same for all the views. The selection of a channel is done by clicking on it, in a time series or a 3D view. Selected channels can be displayed separately, marked as "bad", or deleted. | All the figures displayed by Brainstorm are linked in time. If they feature the same dataset, the sensor selection is also the same for all views. The selection of a channel subset can be easily perfomed by clicking on the corresponding channels in a time series display or a 3D view. Selected channels can be displayed separately, marked as "bad", or deleted. |
| Line 70: | Line 60: |
| == Cortical region of interest: Scout == | == Defining cortical region of interest: Scout == |
| Line 73: | Line 63: |
| The ''scouts'' are subsets of the cortical vertices, defined graphically with the "''Scouts''" tab. They are useful to extract the time series of a the electrical activity of one or several brain regions. | Scouts are cortical regions of interest, defined graphically from the "Scouts" tab. They can be used to extract the time series of MEG and EEG generators within a single or mulitlple brain region. |
| Line 75: | Line 65: |
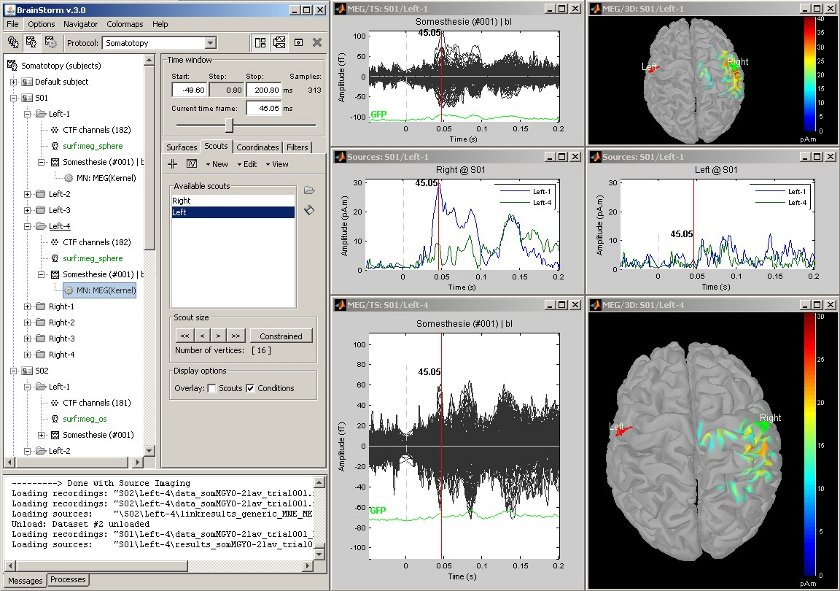
| This example shows the cortical response to an electric stimulation of the left index finger. With the two scouts ''Left ''and ''Right'' we can observe the electrical activity in the primary somatosensory cortex in both hemispheres. | The following example shows the cortical response to an electric stimulation of the left index finger. With the two scouts ''Left ''and ''Right,'' one can observe the electrical activity in the primary somatosensory cortex from each hemisphere. |
| Line 81: | Line 71: |
| == Scouts: multiple conditions == | == Scouts: browsing through multiple conditions == |
| Line 84: | Line 74: |
| Same experiment than the previous example, but showing at the same time the responses for condition ''Left-1'' (electric stimulation of the left index) and ''Left-4'' (left ring finger). | Same experiment than in the previous example, now showing the responses for two conditions: ''Left-1'' (electric stimulation of the left index finger) and ''Left-4'' (left ring finger). |
| Line 90: | Line 80: |
| == View cortical sources in MRI == | == From surface to volume: MRI integration == |
| Line 93: | Line 83: |
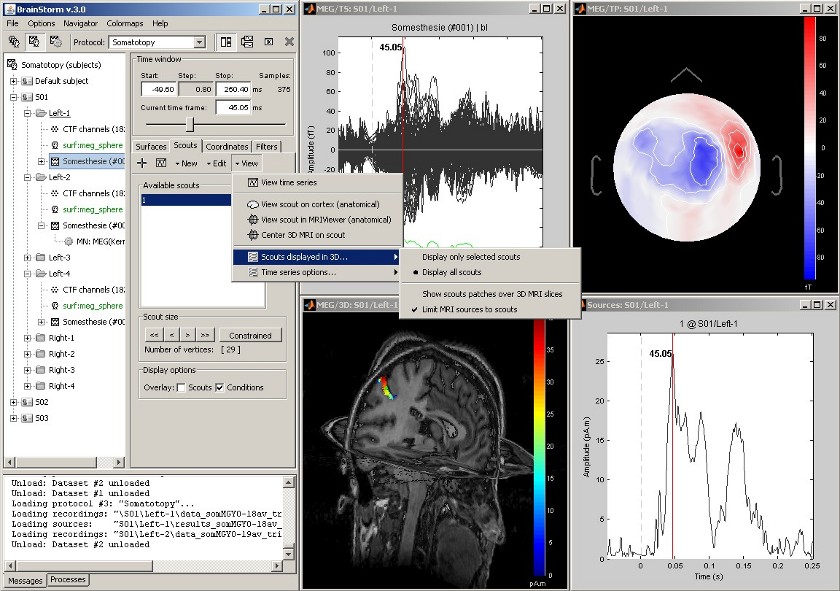
| Same experiment, with additional display of the scout activity on the 3D MRI slices. | Same experiment as above, with additional 3D display of the scout activity in MRI slices. |
| Line 99: | Line 89: |
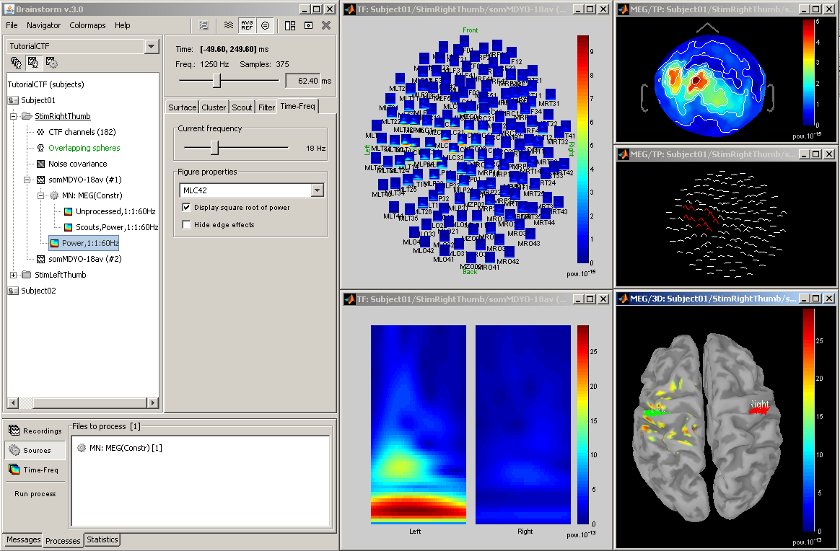
| == Statistical analysis: z-score == | == Time-frequency decompositions == Acquisition system: ''CTF MEG - 151 sensors'' Time-frequency decompositions of sensor data and source time series, extracted from cortical regions of interest. [[attachment:snap_timefreq.jpg|{{attachment:snap_timefreq_sm.jpg|attachment:snap_timefreq.jpg}}]] <<BR>> == Standardization: z-score == |
| Line 102: | Line 101: |
| The "''Processes''" tab in the main [[BrainStorm|Brainstorm]] window allows the user to apply several functions to a set of recordings of sources files. Drag and drop files from the database tree to the white box in the "Processes" tab, and click on "''Run''" to process them. | The "''Processes''" tab in the main Brainstorm window allows users to apply multiple processes to a set of data or source maps. Just drag and drop files from the database tree to the box in the "Processes" tab, and click on "''Run''" to start the processing. |
| Line 104: | Line 103: |
| The function that was applied here is a z-score statistic. The algorithm is the following. | The function that was applied here is a simple z-score standardization. |
| Line 106: | Line 105: |
| For each channel: * Compute the mean ''m'' and the variance ''v'' for the baseline * For all the time samples: substract ''m'' and divide by ''v'' The top-right figure represents the initial sources estimate, and the bottom-left figure shows the z-score values for those sources. |
The top-right figure displays the initial sources estimate, and the bottom-left figure shows the corresponding z-score valued source map. |
| Line 117: | Line 111: |
| == Statistical analysis: t-test == | == Statistical inference: t-test maps == |
| Line 120: | Line 114: |
| The "''Processes''" tab can also be used for computing statistical tests, to evaluate the differences between two experimental conditions. | The "''Processes''" tab can also be used for running statistical tests e.g., to evaluate contrasts between two experimental conditions. |
| Line 122: | Line 116: |
| This example shows the evaluation of the difference between conditions ''GM'' and ''GMM'', using the results of many different subjects (paired Student t-test, p<0.05). The top figure represents the significance of the difference at the electrodes level across the time, and the bottom figure the thresholded t-values at the cortical level at 280ms. | The following example features the evaluation of the difference between conditions ''GM'' and ''GMM'', from the source maps of mulitple subjects (paired Student t-test, p<0.05). The top figure shows the significance of the difference at the sensor level across time. The bottom figure displays the thresholded t-values at the cortical level at 280ms. |
| Line 124: | Line 118: |
| This image also illustrates the "''Coordinates''" tab. It is possible to pick any point from any surface by clicking on it, and immediately get its coordinates in all the coordinates systems used by Brainstorm (MRI, Subject's head and Talairach). | This image also illustrates the "''Coordinates''" tab: It is possible to pick any point from any surface by clicking on it, and immediately get its coordinates in all the coordinates systems used by Brainstorm (MRI, Subject's head and Talairach). |
Screenshots
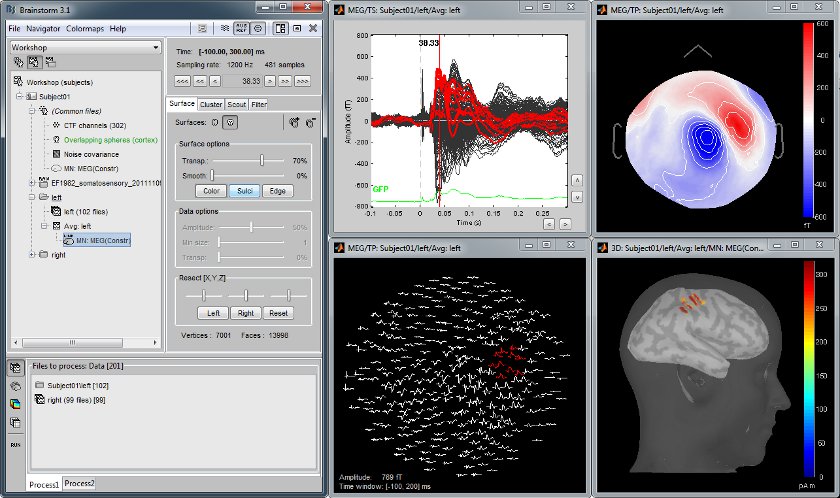
MEG somatosensory evoked responses
Acquisition on an CTF 275 instrument for a left median nerve electric stimulation.
Keeping your data organized and accessible
The tree in the main Brainstorm window represents the database for the selected study. This database has three levels of definition: Protocol (ie. study, selected in the toolbar), Subject, and Condition. Most of the operations that can be performed on a file are easily accessible from the popup menu revealed by right-clicking over the file.
The first three buttons in the toolbar allows the user to switch between different views of the same database:
- Anatomy: display the MRI and surfaces for each participant in the study
- Functional data (sorted by subject): sensor definitions, MEG and EEG data, source models, statistic and time-frequency maps
- Functional data (sorted by condition): same as above, but sorted in a different way
The following example features a MEG+EEG protocol called "Catching", sorted by conditions. There are two experimental conditions, Catch and NoCatch, and 7 subjects per condition. The popup menu shows all the actions that are available for the recordings of subject cc, condition Catch.
Multiple conditions: Baby auditory EEG responses
Acquisition system: EGI GSN - Baby 64 electrodes
Description:
- One subject: "001"
- Three conditions: "GM", "GMM", "VM"
- Two views: overlaid electrodes time series, and estimated cortical sources at t=376ms
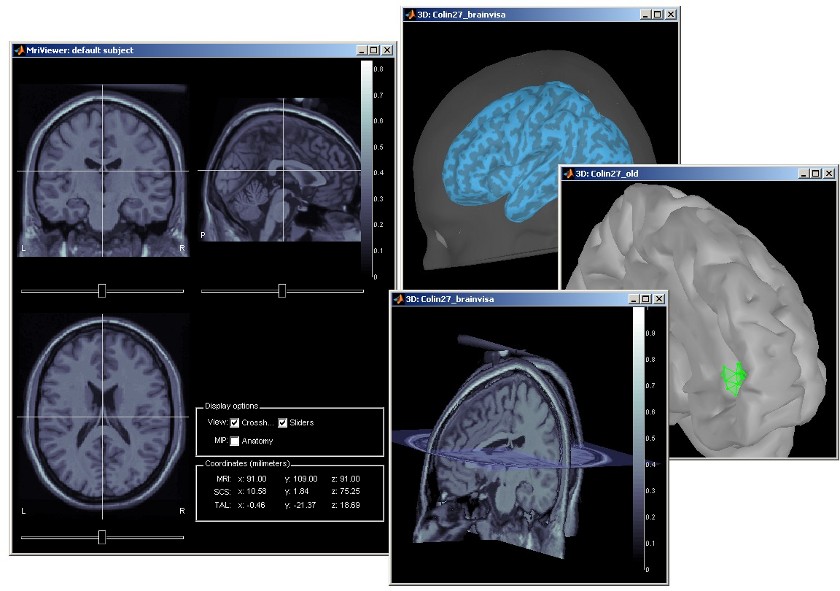
Subject anatomy: MRI and surfaces
Brainstorm features the possibility to model MEG and EEG neural generators either from the individual subject anatomy, or by using a template anatomy (MNI / Colin27) that can be warped to the individual scalp surface. Multiple interactive tools are available to view, register and process the MR images and the corresponding tessellated envelopes. However, tissue segmentation must be performed using another software; multiple options exist today in the academic community (?listed here).
We provide a few examples of the views you can easily obtain with Brainstorm. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surface panel" and contextual popup menus. The MRI slices can be browsed with a simple mouse operation: right-click and mouse drag.
Channel selection
All the figures displayed by Brainstorm are linked in time. If they feature the same dataset, the sensor selection is also the same for all views. The selection of a channel subset can be easily perfomed by clicking on the corresponding channels in a time series display or a 3D view. Selected channels can be displayed separately, marked as "bad", or deleted.
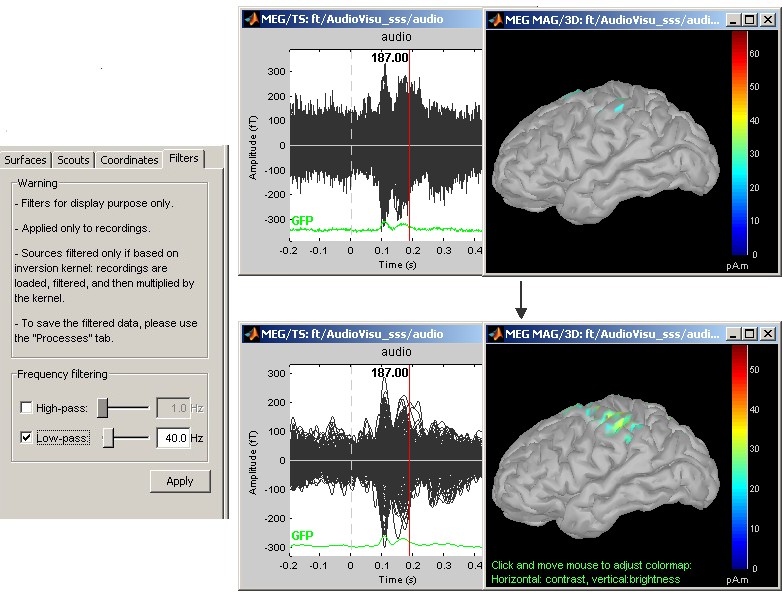
Online bandpass filtering
Recordings and sources: 40Hz low-pass filtering with the "Filters" tab in main Brainstorm window.
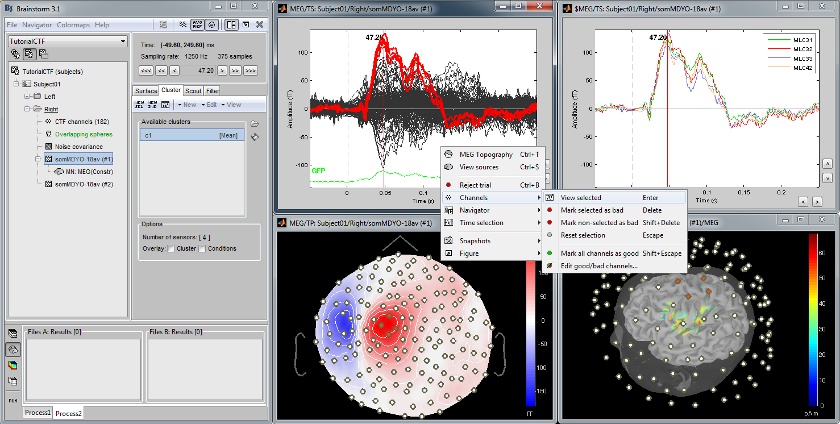
Defining cortical region of interest: Scout
Acquisition system: CTF MEG - 151 sensors
Scouts are cortical regions of interest, defined graphically from the "Scouts" tab. They can be used to extract the time series of MEG and EEG generators within a single or mulitlple brain region.
The following example shows the cortical response to an electric stimulation of the left index finger. With the two scouts Left and Right, one can observe the electrical activity in the primary somatosensory cortex from each hemisphere.
Scouts: browsing through multiple conditions
Acquisition system: CTF MEG - 151 sensors
Same experiment than in the previous example, now showing the responses for two conditions: Left-1 (electric stimulation of the left index finger) and Left-4 (left ring finger).
From surface to volume: MRI integration
Acquisition system: CTF MEG - 151 sensors
Same experiment as above, with additional 3D display of the scout activity in MRI slices.
Time-frequency decompositions
Acquisition system: CTF MEG - 151 sensors
Time-frequency decompositions of sensor data and source time series, extracted from cortical regions of interest.
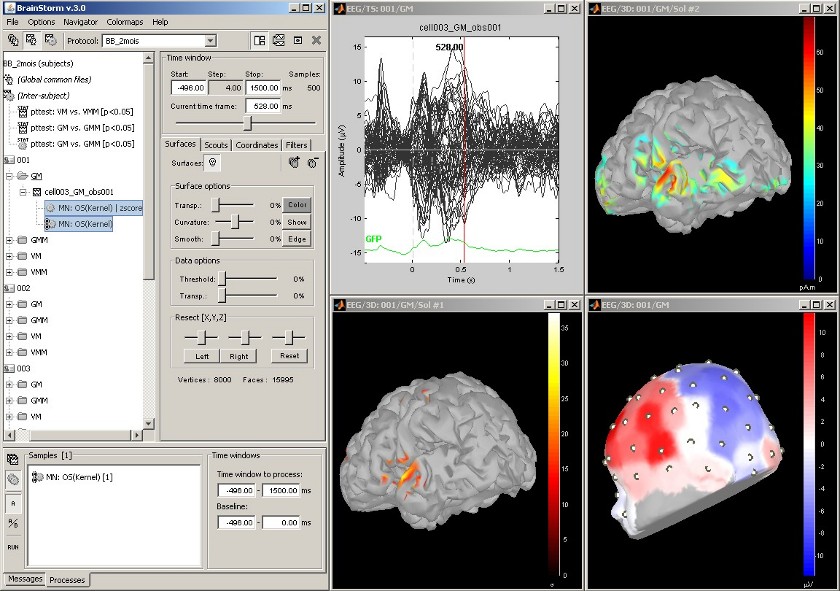
Standardization: z-score
Acquisition system: EGI GSN - Baby 64 electrodes
The "Processes" tab in the main Brainstorm window allows users to apply multiple processes to a set of data or source maps. Just drag and drop files from the database tree to the box in the "Processes" tab, and click on "Run" to start the processing.
The function that was applied here is a simple z-score standardization.
The top-right figure displays the initial sources estimate, and the bottom-left figure shows the corresponding z-score valued source map.
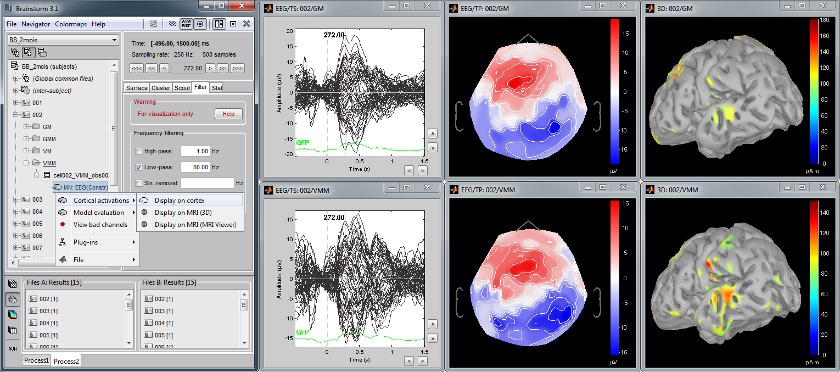
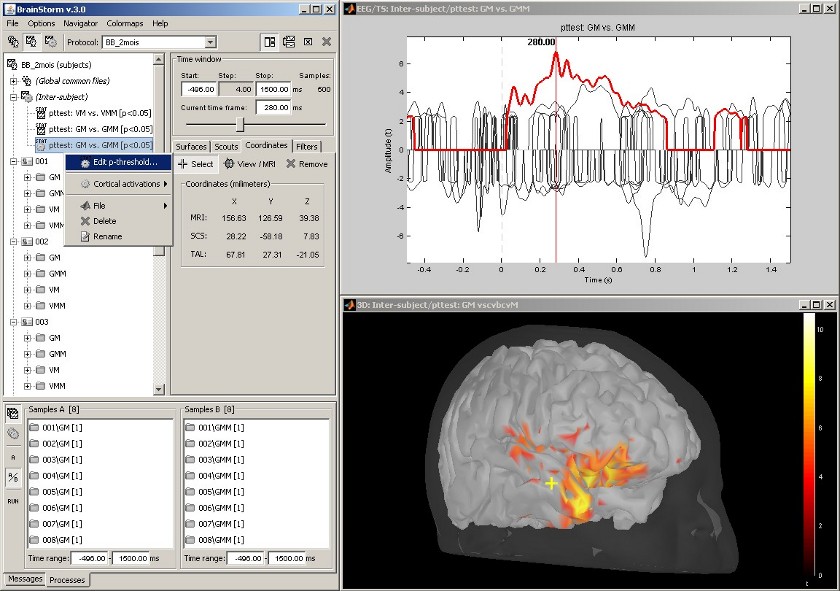
Statistical inference: t-test maps
Acquisition system: EGI GSN - Baby 64 electrodes
The "Processes" tab can also be used for running statistical tests e.g., to evaluate contrasts between two experimental conditions.
The following example features the evaluation of the difference between conditions GM and GMM, from the source maps of mulitple subjects (paired Student t-test, p<0.05). The top figure shows the significance of the difference at the sensor level across time. The bottom figure displays the thresholded t-values at the cortical level at 280ms.
This image also illustrates the "Coordinates" tab: It is possible to pick any point from any surface by clicking on it, and immediately get its coordinates in all the coordinates systems used by Brainstorm (MRI, Subject's head and Talairach).