|
Size: 7156
Comment:
|
Size: 5211
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Screenshots = == MEG auditory evoked responses == Acquisition on an ''Elekta'' ''Neuromag Vectorview306'' instrument. |
= Gallery = == Demo video == Tadel F & Baillet S, March 2015<<BR>>Brainstorm: Imaging neural activity at the speed of brain ([[http://neuroimage.usc.edu/videos/201503_brainstorm_demo.mp4|Download video]]) |
| Line 5: | Line 5: |
| Brief description of the following pictures: | <<HTML(<iframe id="ytplayer" type="text/html" width="853" height="480" src="http://www.youtube.com/embed/30eFJUrRcN4?autoplay=1&origin=http://http://neuroimage.usc.edu/brainstorm" frameborder="0"></iframe>)>> |
| Line 7: | Line 7: |
| * Main Brainstorm window * Time series of all MEG sensors [-200, 500] ms * Magnetic field topography, magnetometers at t=106ms * Topographical layout of magnetometers time series [-200, 500] ms * Modeling of cortical currents, obtained from magnetometers, at t=106ms |
<<HTML(<video width="853" height="480" controls><source src="http://neuroimage.usc.edu/videos/201503_brainstorm_demo.mp4" type="video/mp4"><source src="http://neuroimage.usc.edu/videos/201503_brainstorm_demo.ogg" type="video/ogg">Your browser does not support the video tag.</video>)>> |
| Line 13: | Line 9: |
| [[attachment:snap_1condition.jpg|{{attachment:snap_1condition_sm.jpg|attachment:snap_1condition.jpg}}]] | == MEG somatosensory evoked responses == Acquisition on a ''CTF 275'' instrument for a left median nerve electric stimulation. <<BR>>[[attachment:snap_median.jpg|{{attachment:snap_median_sm.jpg|attachment:snap_median.jpg}}]] |
| Line 15: | Line 12: |
| <<BR>> | == Baby auditory EEG responses == Acquisition system: ''EGI GSN - Baby 64 electrodes''. <<BR>> [[attachment:snap_3conditions.jpg|{{attachment:snap_3conditions_sm.jpg|attachment:snap_3conditions.jpg}}]] |
| Line 17: | Line 15: |
| == Keeping your data organized and accessible == The tree in the main Brainstorm window represents the database for the selected study. This database has three levels of definition: Protocol (ie. study, selected in the toolbar), Subject, and Condition. Most of the operations that can be performed on a file are easily accessible from the popup menu revealed by right-clicking over the file. |
== Continuous recordings and markers == Review recordings directly reading from the original files, edit markers, detect and correct artifacts. <<BR>> [[attachment:snap_raw.jpg|{{attachment:snap_raw_sm.jpg|attachment:snap_raw.jpg}}]] |
| Line 20: | Line 18: |
| The first three buttons in the toolbar allows the user to switch between different views of the same database: | == Artifact detection and correction with SSP == A fully automated cleaning pipeline for the most common artifacts (power lines, eye blinks, heartbeats) is available in Brainstorm. The ocular and cardiac artifacts are corrected using the [[Tutorials/TutRawSsp|Signal Space Projection]] approach. |
| Line 22: | Line 21: |
| * Anatomy: display the MRI and surfaces for each participant in the study * Functional data (sorted by subject): sensor definitions, MEG and EEG data, source models, statistic and time-frequency maps * Functional data (sorted by condition): same as above, but sorted in a different way |
=== Example: Heartbeat === {{attachment:ssp_ecg.jpg}} |
| Line 26: | Line 24: |
| The following example features a MEG+EEG protocol called "''Catching''", sorted by conditions. There are two experimental conditions, ''Catch ''and ''NoCatch'', and 7 subjects per condition. The popup menu shows all the actions that are available for the recordings of subject ''cc'', condition ''Catch''. [[attachment:snap_rightclick.jpg|{{attachment:snap_rightclick_sm.jpg|attachment:snap_rightclick.jpg}}]] <<BR>> == Multiple conditions: Baby auditory EEG responses == Acquisition system: ''EGI GSN - Baby 64 electrodes'' Description: * One subject: "001" * Three conditions: "GM", "GMM", "VM" * Two views: overlaid electrodes time series, and estimated cortical sources at t=376ms [[attachment:snap_3conditions.jpg|{{attachment:snap_3conditions_sm.jpg|attachment:snap_3conditions.jpg}}]] <<BR>> |
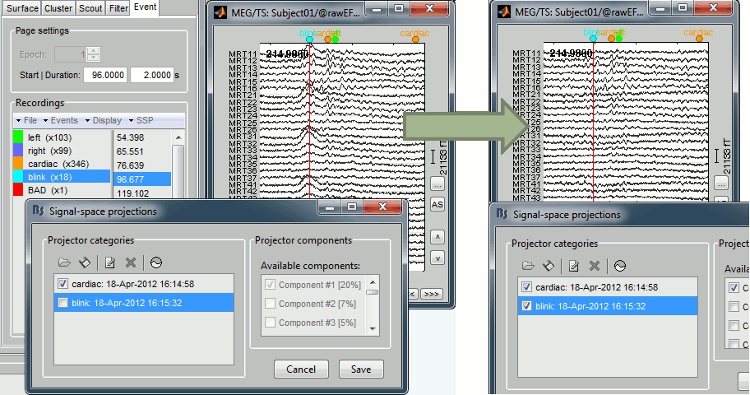
=== Example: Eye blink === [[attachment:sspExample.gif|{{attachment:sspExample_sm.jpg|attachment:sspExample.gif}}]] |
| Line 48: | Line 30: |
| We provide a few examples of the views you can easily obtain with Brainstorm. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surface panel" and contextual popup menus. The MRI slices can be browsed with a simple mouse operation: right-click and mouse drag. [[attachment:snap_anatomy.jpg|{{attachment:snap_anatomy_sm.jpg|attachment:snap_anatomy.jpg}}]] <<BR>> |
We provide a few examples of the views you can easily obtain with Brainstorm. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surface panel" and contextual popup menus. The MRI slices can be browsed with a simple mouse operation: right-click and mouse drag.<<BR>>[[attachment:snap_anatomy.jpg|{{attachment:snap_anatomy_sm.jpg|attachment:snap_anatomy.jpg}}]] |
| Line 55: | Line 33: |
| All the figures displayed by Brainstorm are linked in time. If they feature the same dataset, the sensor selection is also the same for all views. The selection of a channel subset can be easily perfomed by clicking on the corresponding channels in a time series display or a 3D view. Selected channels can be displayed separately, marked as "bad", or deleted. | All the figures displayed by Brainstorm are linked in time. If they feature the same dataset, the sensor selection is also the same for all views. The selection of a channel subset can be easily perfomed by clicking on the corresponding channels in a time series display or a 3D view. Selected channels can be displayed separately, marked as "bad", or deleted.<<BR>>[[attachment:snap_channel.jpg|{{attachment:snap_channel_sm.jpg|attachment:snap_channel.jpg}}]] |
| Line 57: | Line 35: |
| [[attachment:snap_channel.jpg|{{attachment:snap_channel_sm.jpg|attachment:snap_channel.jpg}}]] | == Cortical regions of interest == Acquisition system: ''CTF MEG - 275 sensors'' |
| Line 59: | Line 38: |
| <<BR>> | Scouts are cortical regions of interest, defined graphically from the "Scout" tab. They can be used to extract the time series of MEG and EEG generators within a single or mulitlple brain region. |
| Line 61: | Line 40: |
| == Online bandpass filtering == Recordings and sources: 40Hz low-pass filtering with the "Filters" tab in main Brainstorm window. |
The following example shows the cortical response to an electric stimulation of the left median nerve. With the three scouts "S1 right", "S2 right" and "S2 left"'','' we can track the processing of the stimulus in the brain: 1) contralateral primary somatosensory cortex, 2) contralateral sencondary somatosensory, 3) ipsilateral secondary somatosensory.<<BR>>[[attachment:snap_1scout.jpg|{{attachment:snap_1scout_sm.jpg|attachment:snap_1scout.jpg}}]] |
| Line 64: | Line 42: |
| {{attachment:snap_online_filter.jpg|attachment:snap_online_filter.jpg}} | == Anatomical atlases == Integrated support for the individual surface-based anatomical atlases generated by FreeSurfer and BrainSuite.<<BR>><<BR>> {{attachment:atlases.jpg||height="358",width="750"}} |
| Line 66: | Line 45: |
| <<BR>> | == Scripting environment == Everything that can be done in the interface with mouse clicks can be converted automatically to Matlab scripts, using the Process1 and Process2 tabs.<<BR>>[[attachment:snap_scripting.jpg|{{attachment:snap_scripting_sm.jpg|attachment:snap_scripting.jpg}}]] |
| Line 68: | Line 48: |
| == Defining cortical region of interest: Scout == Acquisition system: ''CTF MEG - 151 sensors'' |
== Digitize and co-register == For accurate source localization in MEG and EEG with registration to anatomical MRI image volume and efficient head localization in MEG, a 3D digitization solution such as the Polhemus-Fastrak is required. Brainstorm features an integrated support for driving a Polhemus device that will help you save time and is efficient for error-checking while digitizing sensor positions and the subject's head shape.<<BR>>The digitized points are used to check and/or fix visually the alignment of the MRI/surfaces and the MEG/EEG sensors.<<BR>><<BR>> {{attachment:polhemus.jpg}} |
| Line 71: | Line 51: |
| Scouts are cortical regions of interest, defined graphically from the "Scouts" tab. They can be used to extract the time series of MEG and EEG generators within a single or mulitlple brain region. The following example shows the cortical response to an electric stimulation of the left index finger. With the two scouts ''Left ''and ''Right,'' one can observe the electrical activity in the primary somatosensory cortex from each hemisphere. [[attachment:snap_1scout.jpg|{{attachment:snap_1scout_sm.jpg|attachment:snap_1scout.jpg}}]] <<BR>> == Scouts: browsing through multiple conditions == Acquisition system: ''CTF MEG - 151 sensors'' Same experiment than in the previous example, now showing the responses for two conditions: ''Left-1'' (electric stimulation of the left index finger) and ''Left-4'' (left ring finger). [[attachment:snap_2scouts.jpg|{{attachment:snap_2scouts_sm.jpg|attachment:snap_2scouts.jpg}}]] <<BR>> == From surface to volume: MRI integration == Acquisition system: ''CTF MEG - 151 sensors'' Same experiment as above, with additional 3D display of the scout activity in MRI slices. [[attachment:snap_scout3D.jpg|{{attachment:snap_scout3D_sm.jpg|attachment:snap_scout3D.jpg}}]] <<BR>> == Time-frequency decompositions == Acquisition system: ''CTF MEG - 151 sensors'' Time-frequency decompositions of sensor data and source time series, extracted from cortical regions of interest. [[attachment:snap_timefreq.jpg|{{attachment:snap_timefreq_sm.jpg|attachment:snap_timefreq.jpg}}]] <<BR>> == Standardization: z-score == Acquisition system: ''EGI GSN - Baby 64 electrodes'' The "''Processes''" tab in the main Brainstorm window allows users to apply multiple processes to a set of data or source maps. Just drag and drop files from the database tree to the box in the "Processes" tab, and click on "''Run''" to start the processing. The function that was applied here is a simple z-score standardization. The top-right figure displays the initial sources estimate, and the bottom-left figure shows the corresponding z-score valued source map. [[attachment:snap_zscore.jpg|{{attachment:snap_zscore_sm.jpg|attachment:snap_zscore.jpg}}]] <<BR>> == Statistical inference: t-test maps == Acquisition system: ''EGI GSN - Baby 64 electrodes'' The "''Processes''" tab can also be used for running statistical tests e.g., to evaluate contrasts between two experimental conditions. The following example features the evaluation of the difference between conditions ''GM'' and ''GMM'', from the source maps of mulitple subjects (paired Student t-test, p<0.05). The top figure shows the significance of the difference at the sensor level across time. The bottom figure displays the thresholded t-values at the cortical level at 280ms. This image also illustrates the "''Coordinates''" tab: It is possible to pick any point from any surface by clicking on it, and immediately get its coordinates in all the coordinates systems used by Brainstorm (MRI, Subject's head and Talairach). [[attachment:snap_ttest.jpg|{{attachment:snap_ttest_sm.jpg|attachment:snap_ttest.jpg}}]] <<BR>> |
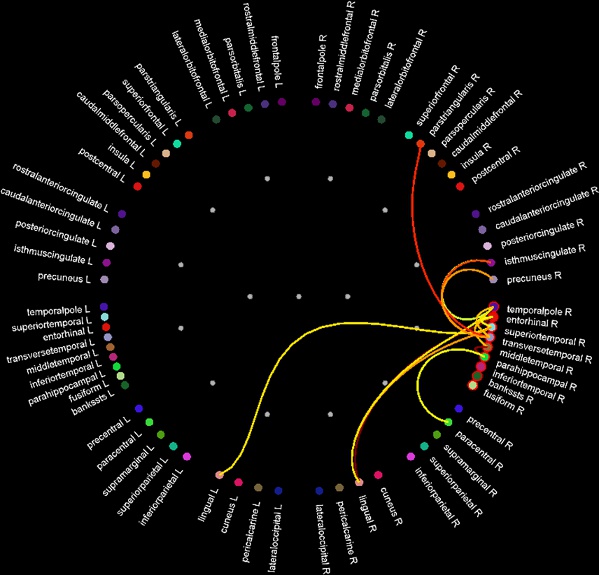
== Functional connectivity == {{attachment:connect_graph.jpg}} |
Gallery
Demo video
Tadel F & Baillet S, March 2015
Brainstorm: Imaging neural activity at the speed of brain (Download video)
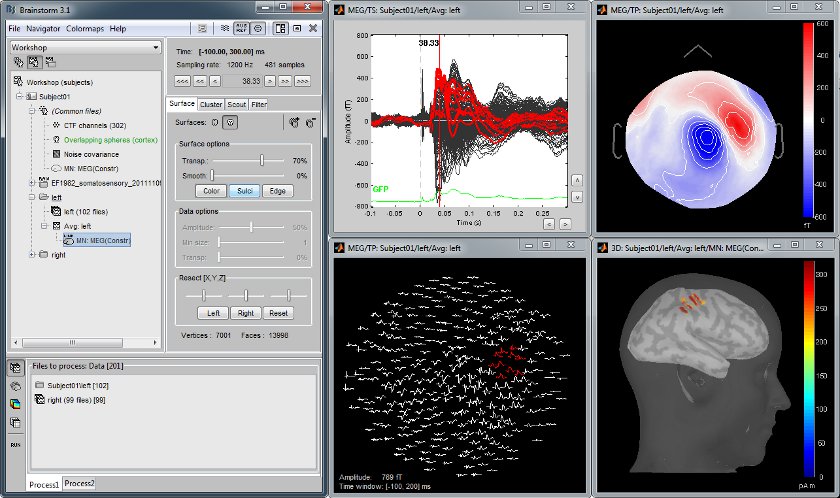
MEG somatosensory evoked responses
Acquisition on a CTF 275 instrument for a left median nerve electric stimulation.
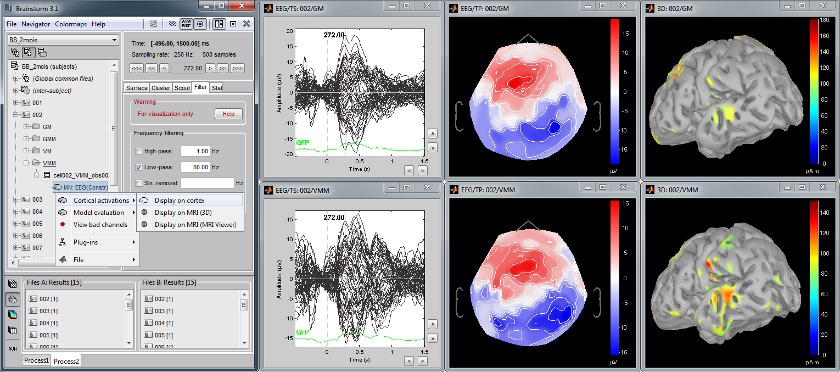
Baby auditory EEG responses
Acquisition system: EGI GSN - Baby 64 electrodes.
Continuous recordings and markers
Review recordings directly reading from the original files, edit markers, detect and correct artifacts.
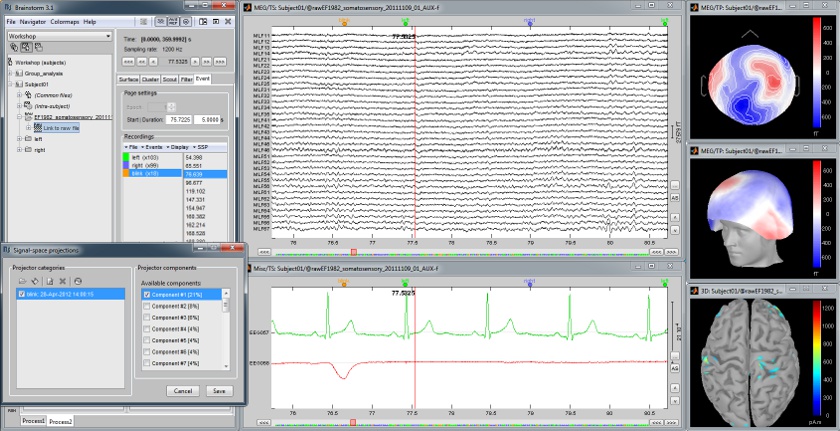
Artifact detection and correction with SSP
A fully automated cleaning pipeline for the most common artifacts (power lines, eye blinks, heartbeats) is available in Brainstorm. The ocular and cardiac artifacts are corrected using the ?Signal Space Projection approach.
Example: Heartbeat
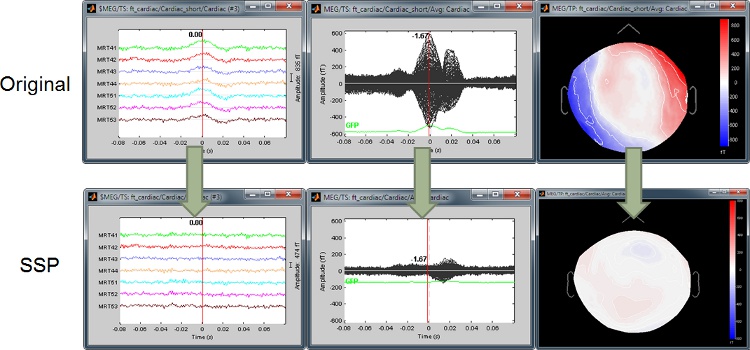
Example: Eye blink
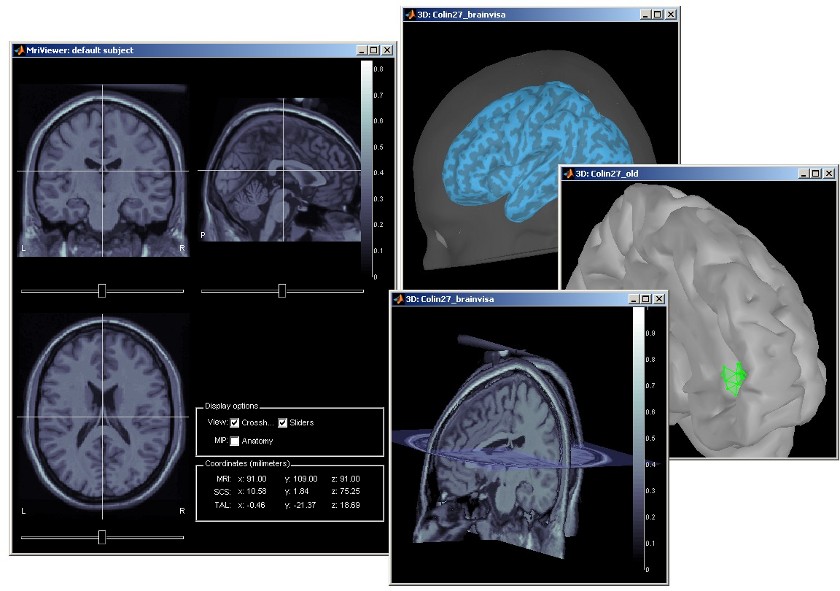
Subject anatomy: MRI and surfaces
Brainstorm features the possibility to model MEG and EEG neural generators either from the individual subject anatomy, or by using a template anatomy (MNI / Colin27) that can be warped to the individual scalp surface. Multiple interactive tools are available to view, register and process the MR images and the corresponding tessellated envelopes. However, tissue segmentation must be performed using another software; multiple options exist today in the academic community (?listed here).
We provide a few examples of the views you can easily obtain with Brainstorm. All the 3D views can be rotated freely with the mouse, zoomed with the wheel, edited with the "Surface panel" and contextual popup menus. The MRI slices can be browsed with a simple mouse operation: right-click and mouse drag.
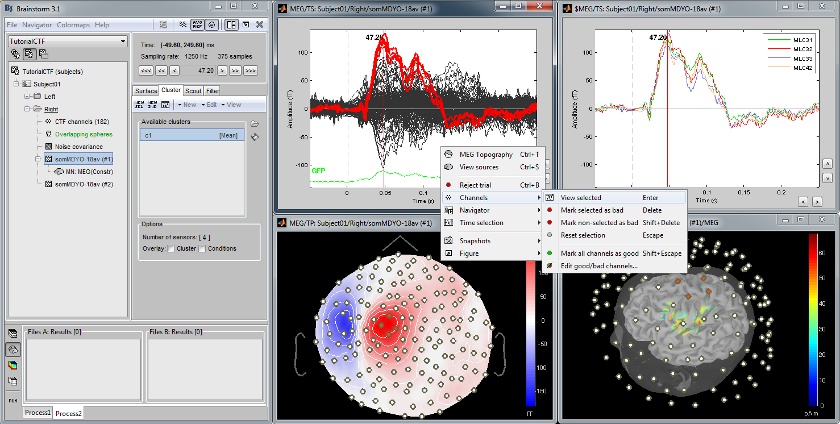
Channel selection
All the figures displayed by Brainstorm are linked in time. If they feature the same dataset, the sensor selection is also the same for all views. The selection of a channel subset can be easily perfomed by clicking on the corresponding channels in a time series display or a 3D view. Selected channels can be displayed separately, marked as "bad", or deleted.
Cortical regions of interest
Acquisition system: CTF MEG - 275 sensors
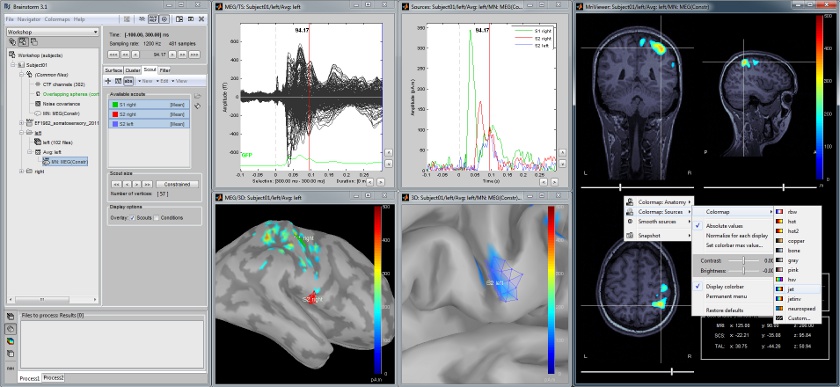
Scouts are cortical regions of interest, defined graphically from the "Scout" tab. They can be used to extract the time series of MEG and EEG generators within a single or mulitlple brain region.
The following example shows the cortical response to an electric stimulation of the left median nerve. With the three scouts "S1 right", "S2 right" and "S2 left", we can track the processing of the stimulus in the brain: 1) contralateral primary somatosensory cortex, 2) contralateral sencondary somatosensory, 3) ipsilateral secondary somatosensory.
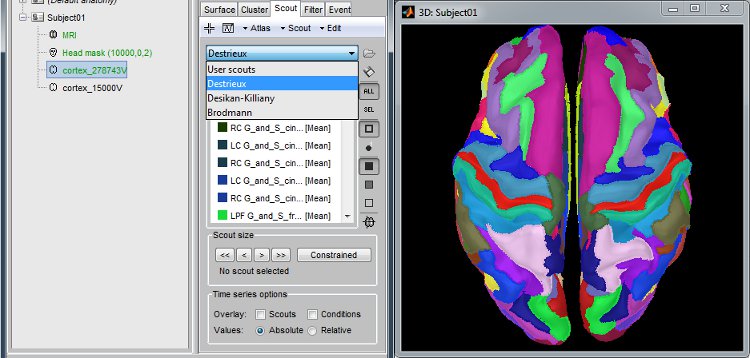
Anatomical atlases
Integrated support for the individual surface-based anatomical atlases generated by FreeSurfer and BrainSuite.
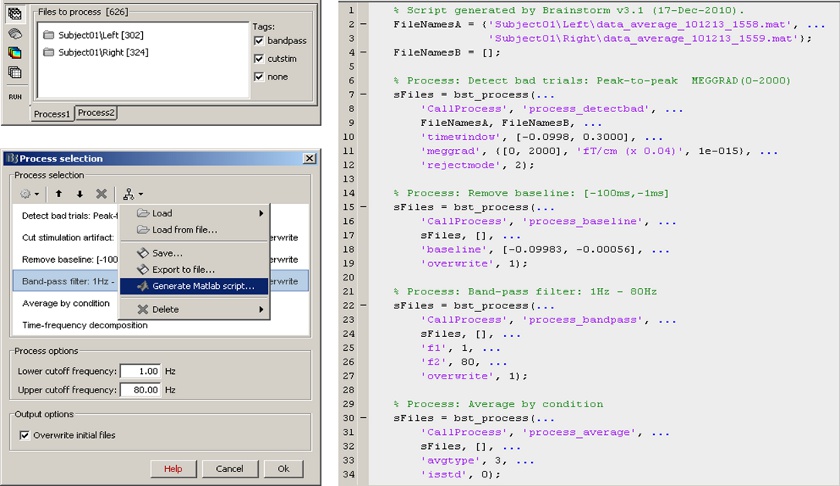
Scripting environment
Everything that can be done in the interface with mouse clicks can be converted automatically to Matlab scripts, using the Process1 and Process2 tabs.
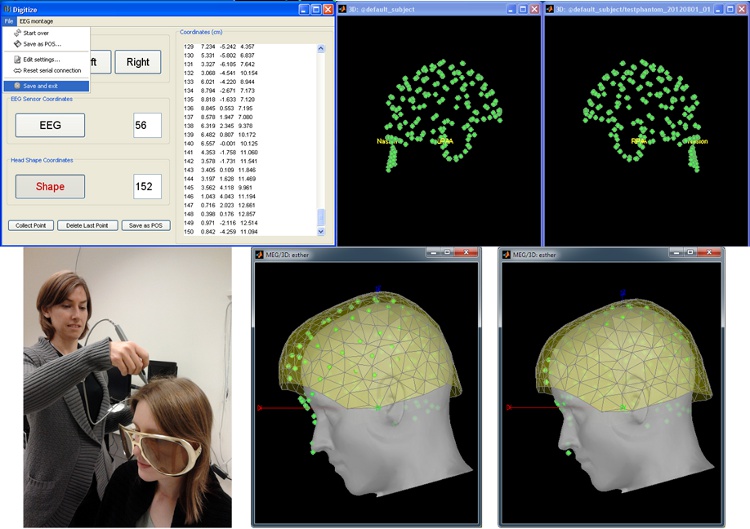
Digitize and co-register
For accurate source localization in MEG and EEG with registration to anatomical MRI image volume and efficient head localization in MEG, a 3D digitization solution such as the Polhemus-Fastrak is required. Brainstorm features an integrated support for driving a Polhemus device that will help you save time and is efficient for error-checking while digitizing sensor positions and the subject's head shape.
The digitized points are used to check and/or fix visually the alignment of the MRI/surfaces and the MEG/EEG sensors.
Functional connectivity