|
Size: 3013
Comment:
|
Size: 7155
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| Half-day introduction to SEEG/ECOG analysis, as part of the [[https://cuttingeeg.org/|CutingEEG symposium]]. | Half-day introduction to SEEG/ECOG analysis, as part of the [[https://cuttingeeg.org/|CuttingEEG symposium]]. |
| Line 11: | Line 11: |
| <<HTML(<TR><TD>)>>'''Instructors'''<<HTML(</TD><TD>)>>[[AboutUs/FrancoisTadel|Francois Tadel]] (Grenoble Institute of Neuroscience), Anne-Sophie Dubarry (Aix-Marseille University)<<HTML(</TD></TR>)>> | <<HTML(<TR><TD>)>>'''Instructors'''<<HTML(</TD><TD>)>>[[AboutUs/FrancoisTadel|Francois Tadel]] (Grenoble Institute of Neuroscience)<<BR>>[[http://www.neurotrack.fr/contact/|Anne-Sophie Dubarry]] (Aix-Marseille University)<<HTML(</TD></TR>)>> |
| Line 13: | Line 13: |
| <<HTML(<TR><TD>)>>'''Audience'''<<HTML(</TD><TD>)>>Users interested in analyzing EEG/sEEG/ECoG/MEG recordings using Brainstorm.<<BR>>Some experience with EEG/MEG is recommended. Teaching in English. <<HTML(</TD></TR>)>> | <<HTML(<TR><TD>)>>'''Audience'''<<HTML(</TD><TD>)>>Users interested in analyzing sEEG/EEG/ECoG/MEG recordings using Brainstorm.<<BR>>Teaching in English. <<HTML(</TD></TR>)>> |
| Line 15: | Line 15: |
| <<HTML(<TR><TD>)>>'''Slides'''<<HTML(</TD><TD>)>>[[http://neuroimage.usc.edu/brainstorm/WorkshopCambridge2017?action=AttachFile&do=get&target=cambridge2017_slides_tadel.pdf|Introduction]] <<HTML(</TD></TR>)>> | <<HTML(<!-- TR><TD>)>>'''Slides'''<<HTML(</TD><TD>)>>[[http://neuroimage.usc.edu/brainstorm/WorkshopCambridge2017?action=AttachFile&do=get&target=cambridge2017_slides_tadel.pdf|Introduction]] <<HTML(</TD></TR -->)>> |
| Line 19: | Line 19: |
| == Requirements == The participants are required to <<HTML(<FONT color=red>)>> '''bring a laptop''' <<HTML(</FONT>)>> (an external mouse will add to your comfort). In order to make the session as efficient as possible, we ask all the attendees to <<HTML(<FONT color=red>)>>'''download, install and test'''<<HTML(</FONT>)>> the software and sample dataset on their laptops prior to the workshop. |
== Program == This session is based on a simplified version of the [[https://neuroimage.usc.edu/brainstorm/Tutorials/Epileptogenicity|SEEG/Epileptogenicity tutorial]]. |
| Line 22: | Line 22: |
| Please read carefully the following instructions:<<BR>>[[WorkshopParis2018Install|How to prepare your laptop for the training]] == Program: Day 1 == |
|
| Line 29: | Line 26: |
| <<HTML(<TR><TD>)>>9:15-9:45<<HTML(</TD><TD>)>>'''Lecture: Brainstorm overview (Tadel''') | <<HTML(<TR><TD>)>>9:15-9:45<<HTML(</TD><TD>)>>'''Lecture: Brainstorm overview<<BR>>''' |
| Line 33: | Line 30: |
| <<HTML(<TR><TD>)>>10:00-10:45<<HTML(</TD><TD>)>>'''Practice''' | <<HTML(<TR><TD>)>>9:45-10:30<<HTML(</TD><TD>)>>'''Anatomy''' |
| Line 37: | Line 34: |
| MRI volumes, surfaces and anatomical atlases<<HTML(</TD></TR>)>> | MRI volumes, surfaces |
| Line 39: | Line 36: |
| <<HTML(<TR><TD>)>>10:45-11:00<<HTML(</TD><TD>)>>''Coffee break''<<HTML(</TD></TR>)>> | Coregistration of pre- and post-implantation images<<HTML(</TD></TR>)>> |
| Line 41: | Line 38: |
| <<HTML(<TR><TD>)>>11:00-12:30<<HTML(</TD><TD>)>>'''Practice''' | <<HTML(<TR><TD>)>>10:30-11:30<<HTML(</TD><TD>)>>'''SEEG recordings''' |
| Line 43: | Line 40: |
| Registration EEG/MRI Review of continuous EEG recordings |
Reviewing continuous SEEG recordings |
| Line 49: | Line 44: |
| Frequency filters | Marking SEEG contacts on post-implantation image |
| Line 51: | Line 46: |
| <<HTML(</TD></TR>)>> | Projecting SEEG recordings on MRI and cortex surface |
| Line 53: | Line 48: |
| <<HTML(<TR><TD>)>>12:30-13:30<<HTML(</TD><TD>)>>''Lunch''<<HTML(</TD></TR>)>> | <<HTML(<TR><TD>)>>11:30-12:15<<HTML(</TD><TD>)>>'''Epileptogenicity maps''' |
| Line 55: | Line 50: |
| <<HTML(<TR><TD>)>>13:30-14:30<<HTML(</TD><TD>)>>'''Practice''' | Importing epochs of interest in the database |
| Line 57: | Line 52: |
| Artifact detection | Time-frequency analysis: Identification of ictal HFO frequency bands |
| Line 59: | Line 54: |
| Artifact correction with ICA | Computation of epileptogenicity maps: Localization of the seizure onset zone |
| Line 61: | Line 56: |
| Averaging<<HTML(</TD></TR>)>> <<HTML(<TR><TD>)>>14:30-15:00<<HTML(</TD><TD>)>>'''Lecture: Source modeling (Baillet)'''<<HTML(</TD></TR>)>> <<HTML(<TR><TD>)>>15:00-15:30<<HTML(</TD><TD>)>>''Coffee break''<<HTML(</TD></TR>)>> <<HTML(<TR><TD>)>>15:30-17:30<<HTML(</TD><TD>)>>'''Practice''' Head modeling, cortical source reconstruction Regions of interest (ROIs) |
Estimation of the propagation pathways |
| Line 76: | Line 61: |
<<BR>><<BR>> ---- = Prepare your laptop for the workshop (optional) = Computers will be available with Matlab and Brainstorm installed, however we recommend participants to bring a laptop. Setting up the software environment is an important part of the training and would be a good preparation step to this workshop. Follow the instructions below if you are insterested in following the hands-on session on your own laptop. == Important notes == * Working with MEG/EEG/SEEG recordings involves a lot of computational resources and large display windows. Therefore we recommend that you bring a laptop with a decent processing capacity, '''4Gb RAM''', a '''64bit''' operating system, and a '''screen larger than 13"'''. * You would add to your comfort by bringing an '''external mouse'''. Most of the manipulations are done with the mouse, and some involve an intensive use of the mouse wheel/scrolling. * Don't forget your '''power adapter'''! == Installation instructions == Before coming to the workshop, you need to download the software and the tutorial dataset from the Brainstorm website (110Mb). Additionally, the epileptogenicity analysis is based on SPM12 (100Mb). To streamline troubleshooting during the session, please save all the downloaded files '''__on your Desktop__'''. 1. From the Brainstorm [[http://neuroimage.usc.edu/bst/download.php|Download]] page, log in or create a Brainstorm account 1. Download the following files to '''your Desktop''' '''folder''': * '''Brainstorm software''': <<HTML(<FONT color=red>)>>brainstorm_YYMMDD.zip<<HTML(</FONT>)>> (70 Mb) * '''Tutorial dataset''': <<HTML(<FONT color=red>)>>workshop_cuttingeeg.zip<<HTML(</FONT>)>> (41 Mb) 1. If you already have Brainstorm on your laptop, make sure to update it before coming. 1. Download SPM12 from the UCL website, free registration required:<<BR>>http://www.fil.ion.ucl.ac.uk/spm/software/download/ 1. Unzip the three downloaded files on your desktop, then delete them 1. Create a folder "'''brainstorm_db'''" on your '''Desktop ''' 1. Final check: you should have now 4 folders on your desktop: * '''brainstorm3''': Program folder, with the source code and the compiled executable * '''brainstorm_db''': Brainstorm database (empty) * '''workshop_cuttingeeg''': Example dataset used during the training session * '''spm12''': SPM toolbox, required for the epileptogenicity analysis 1. Run Brainstorm, make sure it works correctly on your computer (read the next section) 1. Tell Brainstorm where you unzipped SPM12: Menu File > Edit preferences > SPM toolbox. == Running Brainstorm for the first time == ==== With Matlab ==== 1. Matlab versions >= 2014b are faster and produce nicer graphics: consider upgrading if possible 1. Start Matlab 1. Do '''NOT '''add brainstorm3 folder to your Matlab path: this will be done automatically 1. Go to the brainstorm3 folder 1. Type "brainstorm" in the command window 1. When asked for the Brainstorm database folder, pick the "brainstorm_db" you have just created 1. Tell Brainstorm where you unzipped SPM12: Menu File > Edit preferences > SPM toolbox. <<BR>>No need to add SPM to the Matlab path either, Brainstorm will do it when needed. ==== Without Matlab ==== . The Brainstorm distribution includes a stand-alone version, created with the Matlab Compiler, which can be used without any Matlab license. However, compiled code cannot execute Matlab scripts that are not compiled, which prevents the use of external toolboxes (SPM, FieldTrip).<<BR>><<BR>>The last part of this hands-on session will rely on functions from the SPM toolbox, therefore if you use the compiled version, you wouldn't be able to follow the session until the end on your laptop. If you still want to run the compiled version of Brainstorm on your laptop, please refer to the main [[https://neuroimage.usc.edu/brainstorm/Installation|installation page]]. == Make sure it works == In order to check if Brainstorm works properly or your computer, please follow these instructions: * Create a new protocol: Menu File > New protocol.<<BR>>Set the name to "Test", don't change the other default options. * In the folder "Default anatomy", double-click successively on the files: MRI, head, cortex_15002V * It should open the figures below, make sure they work correctly: * Click in the MRI Viewer to move the slices * Click+move your mouse in the 3D figure to rotate * Zoom in/out with the mouse wheel or the two-finger scroll on MacBook pads * The head surface should be transparent, with the brain visible through it {{attachment:test_bst.gif}} == Troubleshooting == For any technical problem, please contact Francois Tadel ( francois.tadel@mcgill.ca ) |
Paris, France: July 2, 2018
Half-day introduction to SEEG/ECOG analysis, as part of the CuttingEEG symposium.
General information
| Where | Telecom ParisTech |
| When | Monday July 2nd, 2018: 8:30-12:15 |
| Instructors | Francois Tadel (Grenoble Institute of Neuroscience) Anne-Sophie Dubarry (Aix-Marseille University) |
| Audience | Users interested in analyzing sEEG/EEG/ECoG/MEG recordings using Brainstorm. Teaching in English. |
Program
This session is based on a simplified version of the SEEG/Epileptogenicity tutorial.
| 8:30-9:15 | Onsite assistance in installing the material for the training session |
| 9:15-9:45 | Lecture: Brainstorm overview Software structure, typical data workflow |
| 9:45-10:30 | Anatomy Database explorer MRI volumes, surfaces Coregistration of pre- and post-implantation images |
| 10:30-11:30 | SEEG recordings Reviewing continuous SEEG recordings Montages and management of event markers Marking SEEG contacts on post-implantation image Projecting SEEG recordings on MRI and cortex surface |
| 11:30-12:15 | Epileptogenicity maps Importing epochs of interest in the database Time-frequency analysis: Identification of ictal HFO frequency bands Computation of epileptogenicity maps: Localization of the seizure onset zone Estimation of the propagation pathways |
Prepare your laptop for the workshop (optional)
Computers will be available with Matlab and Brainstorm installed, however we recommend participants to bring a laptop. Setting up the software environment is an important part of the training and would be a good preparation step to this workshop. Follow the instructions below if you are insterested in following the hands-on session on your own laptop.
Important notes
Working with MEG/EEG/SEEG recordings involves a lot of computational resources and large display windows. Therefore we recommend that you bring a laptop with a decent processing capacity, 4Gb RAM, a 64bit operating system, and a screen larger than 13".
You would add to your comfort by bringing an external mouse. Most of the manipulations are done with the mouse, and some involve an intensive use of the mouse wheel/scrolling.
Don't forget your power adapter!
Installation instructions
Before coming to the workshop, you need to download the software and the tutorial dataset from the Brainstorm website (110Mb). Additionally, the epileptogenicity analysis is based on SPM12 (100Mb). To streamline troubleshooting during the session, please save all the downloaded files on your Desktop.
From the Brainstorm Download page, log in or create a Brainstorm account
Download the following files to your Desktop folder:
Brainstorm software: brainstorm_YYMMDD.zip (70 Mb)
Tutorial dataset: workshop_cuttingeeg.zip (41 Mb)
- If you already have Brainstorm on your laptop, make sure to update it before coming.
Download SPM12 from the UCL website, free registration required:
http://www.fil.ion.ucl.ac.uk/spm/software/download/- Unzip the three downloaded files on your desktop, then delete them
Create a folder "brainstorm_db" on your Desktop
- Final check: you should have now 4 folders on your desktop:
brainstorm3: Program folder, with the source code and the compiled executable
brainstorm_db: Brainstorm database (empty)
workshop_cuttingeeg: Example dataset used during the training session
spm12: SPM toolbox, required for the epileptogenicity analysis
- Run Brainstorm, make sure it works correctly on your computer (read the next section)
Tell Brainstorm where you unzipped SPM12: Menu File > Edit preferences > SPM toolbox.
Running Brainstorm for the first time
With Matlab
Matlab versions >= 2014b are faster and produce nicer graphics: consider upgrading if possible
- Start Matlab
Do NOT add brainstorm3 folder to your Matlab path: this will be done automatically
- Go to the brainstorm3 folder
- Type "brainstorm" in the command window
- When asked for the Brainstorm database folder, pick the "brainstorm_db" you have just created
Tell Brainstorm where you unzipped SPM12: Menu File > Edit preferences > SPM toolbox.
No need to add SPM to the Matlab path either, Brainstorm will do it when needed.
Without Matlab
The Brainstorm distribution includes a stand-alone version, created with the Matlab Compiler, which can be used without any Matlab license. However, compiled code cannot execute Matlab scripts that are not compiled, which prevents the use of external toolboxes (SPM, FieldTrip).
The last part of this hands-on session will rely on functions from the SPM toolbox, therefore if you use the compiled version, you wouldn't be able to follow the session until the end on your laptop. If you still want to run the compiled version of Brainstorm on your laptop, please refer to the main installation page.
Make sure it works
In order to check if Brainstorm works properly or your computer, please follow these instructions:
Create a new protocol: Menu File > New protocol.
Set the name to "Test", don't change the other default options.- In the folder "Default anatomy", double-click successively on the files: MRI, head, cortex_15002V
- It should open the figures below, make sure they work correctly:
- Click in the MRI Viewer to move the slices
- Click+move your mouse in the 3D figure to rotate
Zoom in/out with the mouse wheel or the two-finger scroll on MacBook pads
- The head surface should be transparent, with the brain visible through it
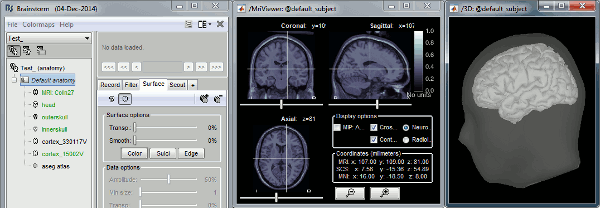
Troubleshooting
For any technical problem, please contact Francois Tadel ( francois.tadel@mcgill.ca )
