|
Size: 11459
Comment:
|
Size: 11453
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 25: | Line 25: |
| '''''Nasiotis, K., Cousineau, M., Tadel, F., Peyrache, A., Leahy, R., Pack, C., and Baillet, S. (2019). Integrated Open-Source Software for Multiscale Electrophysiology. BioRxiv 584185.<<BR>>''''' |
'''Nasiotis, K., Cousineau, M., Tadel, F. ''et al.'' Integrated open-source software for multiscale electrophysiology. ''Sci Data'' 6, 231 (2019) doi:10.1038/s41597-019-0242-z''''''''<<BR>>''''' |
Brainstorm's Suite for Invasive Neurophysiology
Note: In order to follow this tutorial, you need the latest version of Brainstorm (at least version 01-Nov-2018).
Author: Konstantinos Nasiotis
Contents
Introduction
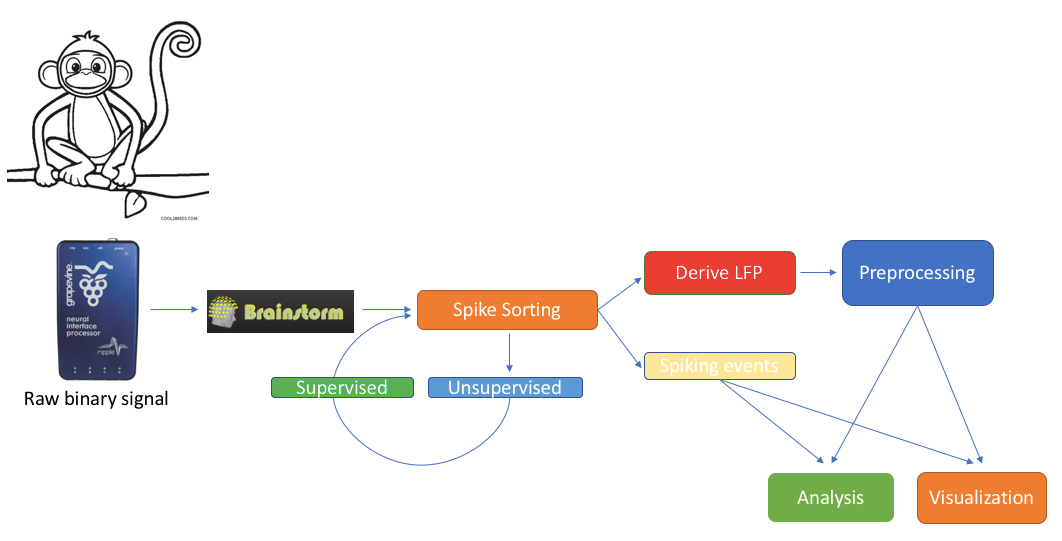
We feature a series of data analysis tools specifically dedicated to Invasive electrophysiology research (e-phys). We see this suite of expanded Brainstorm features as an opportunity for enabling a methodological continuum between (multi)cellular and systems level (with EEG and MEG) data analytics and facilitate the scientific dialogue and interactions between subdisciplines in the field.
Invasive electrophysiological recordings capture the spiking activity of individual or small groups of neurons. Since spiking events are transient phenomena (the typical duration of spikes is 1ms), the sampling rate of e-phys recording systems is typically much faster than for scalp EEG and MEG recordings. State-of-the-art acquisition systems can accommodate simultaneous recordings from hundreds of channels over several multi-electrode arrays positioned across the preparation.
We foresee that Brainstorm will contribute to the efficient data management and processing when there is a need for efficient, open and reproducible data management and utilization of large quantities of data. Users that have Matlab’s parallel processing toolbox, will greatly benefit from the utilization of its properties. Most functions of the electrophysiology toolbox have a checkbox where users can enable parallelization of the processes.
Additionally, it is highly recommended for users that need to analyze large files, to select a large number for memory utilization on any processes that allow users to adjust the memory used, since the speed of the processes (especially the spike sorting procedures that use demultiplexing of the raw signal) can be greatly improved by the extra memory used.
All the function that will be described on this toolbox, are located in the Electrophysiology processes tab.
Any toolboxes embedded represent the work of their respective authors and need to be appropriately cited. Links to the papers that the toolboxes correspond to are located on each section.
Users that utilize this toolbox can cite:
Nasiotis, K., Cousineau, M., Tadel, F. et al. Integrated open-source software for multiscale electrophysiology. Sci Data 6, 231 (2019) doi:10.1038/s41597-019-0242-z
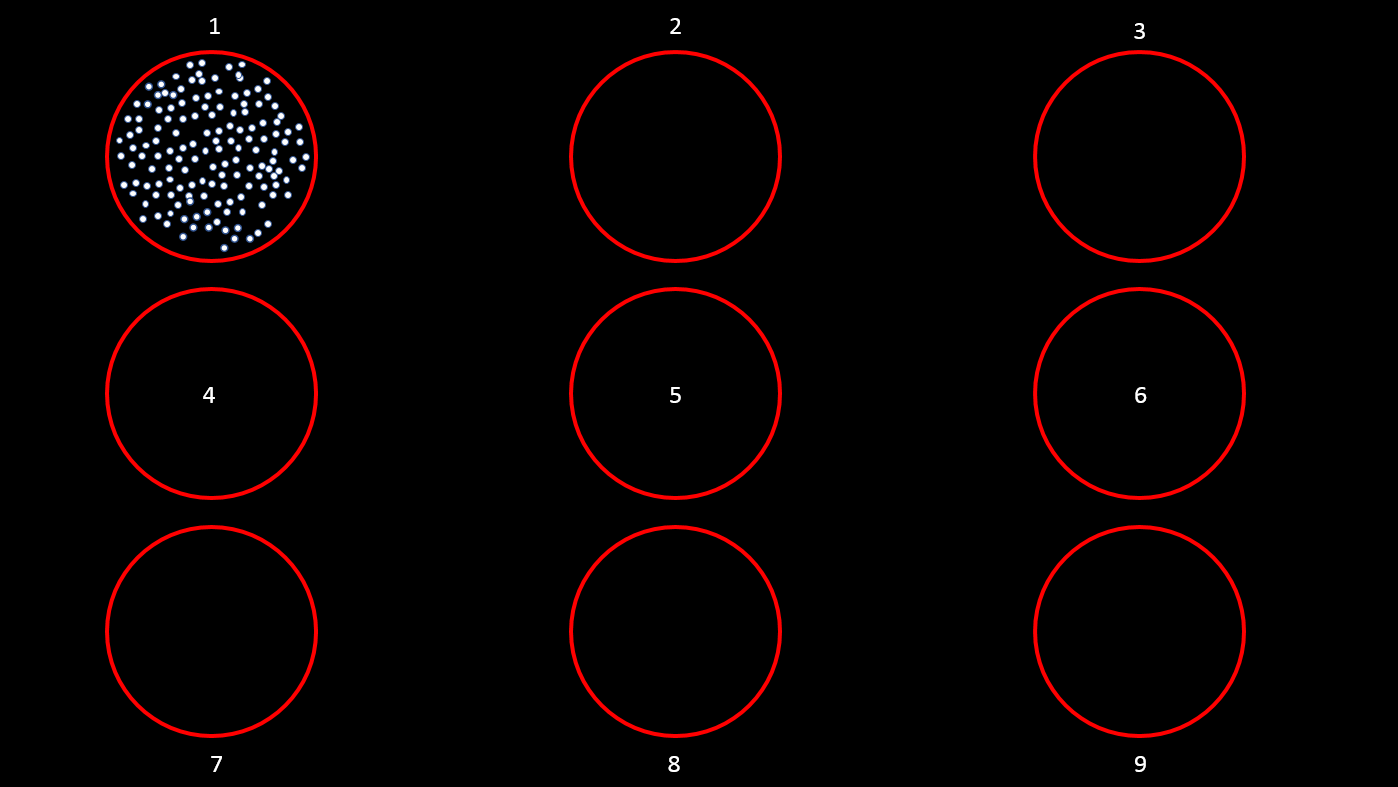
This tutorial uses a dataset collected from Nardin Nakhla in Dr. Christopher Pack’s lab (www.packlab.mcgill.ca), from a macaque monkey with a penetration chamber implanted in areas MT and MST. The recording probe consisted of 32 channels and 1 reference electrode. Users can download this tutorial data that will contain a converted Plexon acquisition system raw file (ytu288c-01.plx), an events file (converted to Brainstorm format, that labels the experimental conditions appropriately), a channel positioning file and an anatomy folder (Floyd MRI) that can all be imported to Brainstorm. The task for the animal was to maintain fixation on a target while a superflow stimulus (dots that rotate, expand or translate in a partially coherent mode) was presented in one of 9 possible visual locations (as shown in the figure below). The events from the stimulation system are saved along with pre-spike-sorted events.
Importing raw e-phys data
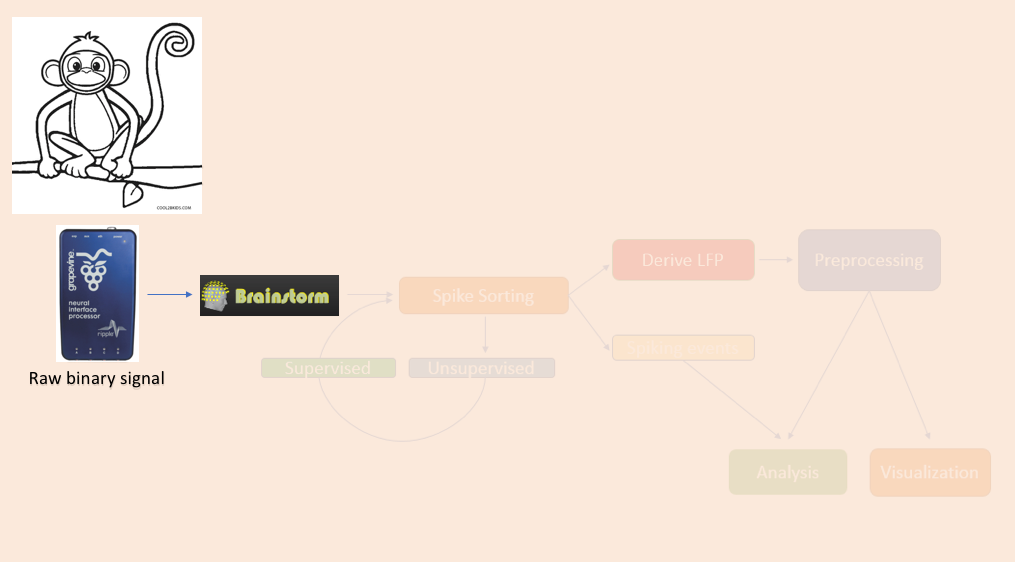
The importation of raw e-phys data into Brainstorm follows the same procedure as it does for any other raw recording. Users firstly assign a name for their protocol/study, followed by the name of the subject/monkey, and sequentially can review their raw file.
All these steps are briefly repeated here for the example e-Phys file for completeness.
Create protocol
Create a new protocol: File > New protocol.
Give the Protocol name: Tutorial_e-Phys.
Default anatomy: No, use individual anatomy.
Default channel File: No, use one channel file per acquisition run.
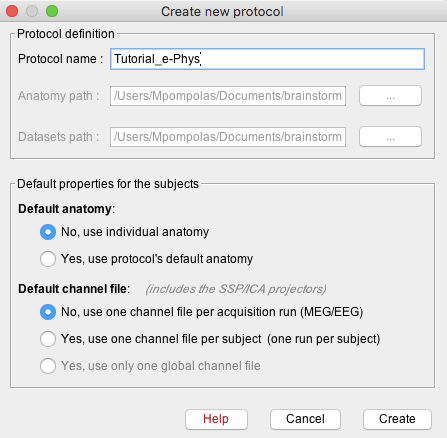
- Press create, and this will create the new protocol.
Add subject
The next step will be to set the subject/monkey:
- Right-click on the protocol created: Tutorial_e-Phys (subjects)
Select New subject
Make sure the tab Functional Data (sorted by subjects) is selected – select it under the Tutorial_e-Phys drop-down.
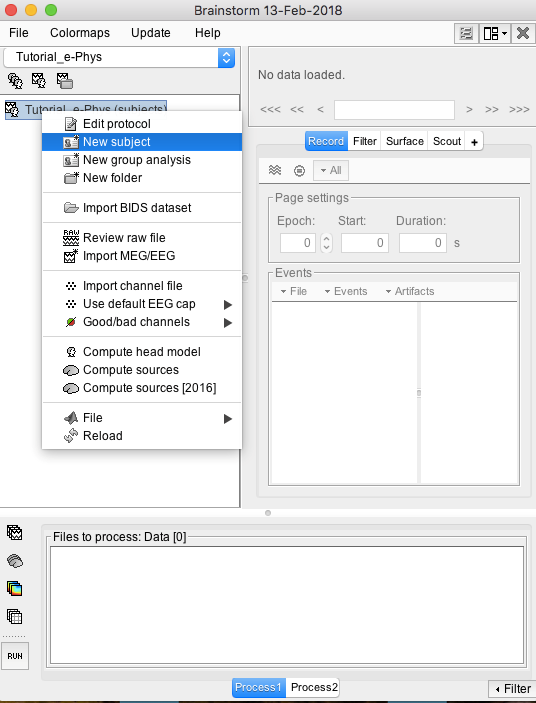
- A new window will appear that allows users to set the Subject name and to choose the individual anatomy and default channel file. Choose no for both default anatomy and default channel file.
Use subject Name: Floyd
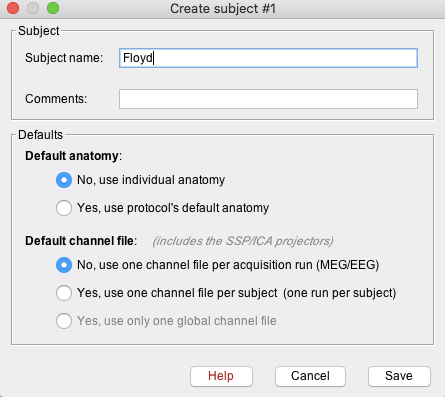
- Once Save is selected, a new subject (Floyd) will be created in the database.
Import anatomy (Optional)
Users can now import the MRI if needed. Instructions on how to import the MRI can be found here and just use the Floyd MRI folder as the Freesurfer/anatomy folder. However, the analysis described on later steps (besides a few visualization functions) does not require an anatomical MRI.
Import recordings
Raw files can be imported by selecting the acquisition system used, and sequentially selecting the raw file. Just right-click on the Subject (Floyd) and then click on Review raw file.
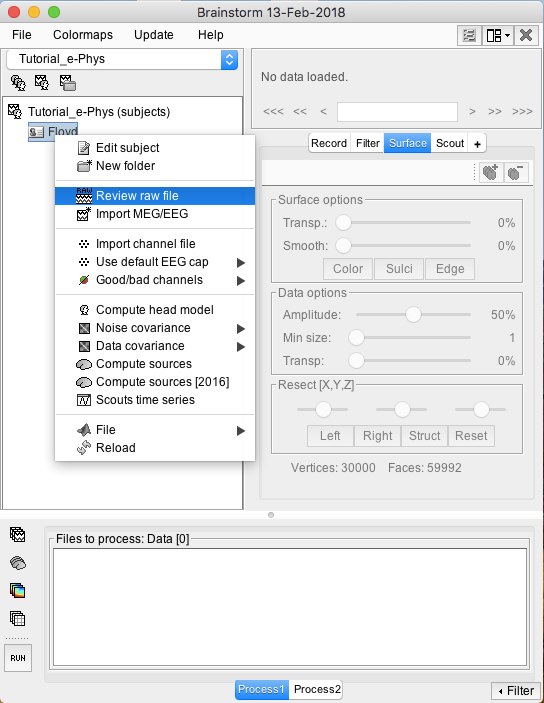
A new window will pop-up that allows users to select the acquisition system and the raw file. Select EEG:Plexon (*.plx) as the acquisition system and navigate to the folder the ytu288c-01.plx raw file is saved.
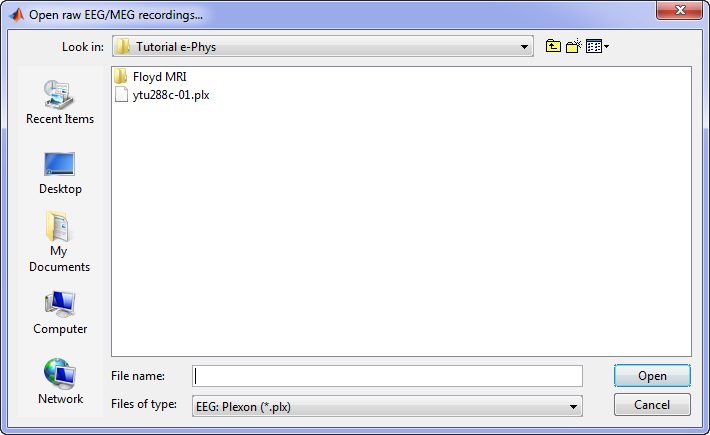
Once the procedure is complete, this will create a new folder and two new files in the database: ytu288c-01_converted which is the name of the raw file, and one channel file (Plexon channels (33)) and the Link to raw file. These two files contain all the information from the header of the acquisition system. It is important to note that no data (actual recordings) have been imported in Brainstorm yet.
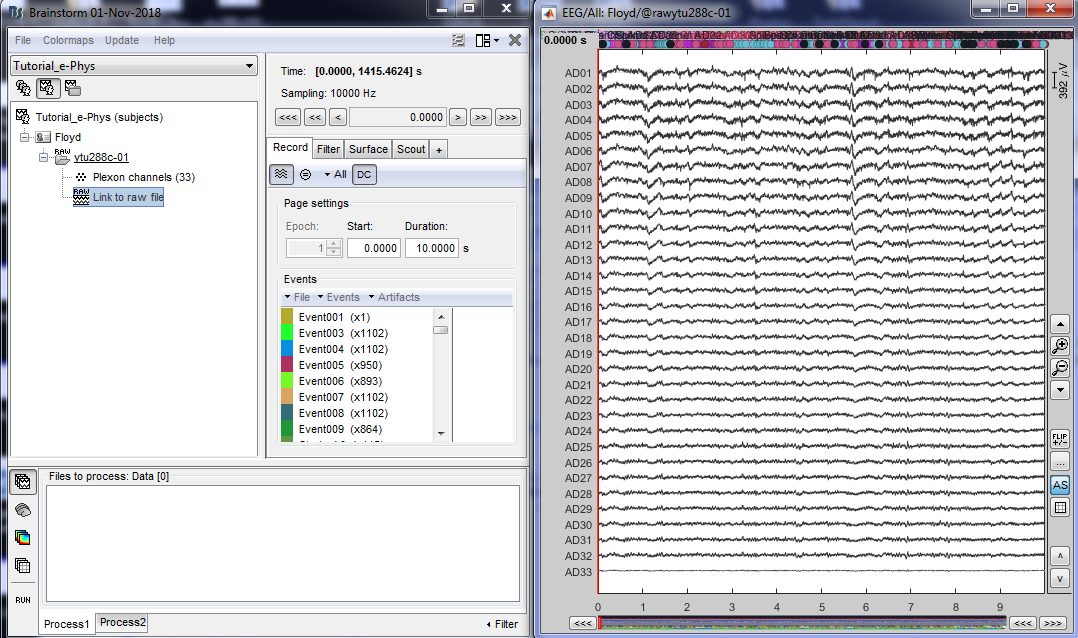
By double clicking on the Link to raw file, a viewer will open that allows the user to navigate through the recordings. This is a very easy way to inspect the recordings for artifacts or stimulation effects since only the selected part is loaded in memory on-the-fly, without importing anything in the database. Any events capured from the acquisition system are automatically loaded from the importer.
It is important to note that some acquisition systems might have enabled their own automatic spike sorter. These events are automatically labeled with the convention that is used throughout the e-phys toolbox:
“Spikes Channel ChannelLabel”, where ChannelLabel is the Label of the electrode assigned by the acquisition system. If the acquisition system assigned the spikes’ waveforms into more than one cluster/neuron during this automatic clustering, the neuronal events would be (e.g. for an electrode that picked up spikes from 3 neurons): “Spikes Channel ChannelLabel |1|”, “Spikes Channel ChannelLabel |2|”, “Spikes Channel ChannelLabel |3|”.
For more information, users can visit this page.
The events are comprised of the signals that the stimulation system has sent (Stim On, Start, Stop etc.) and the pre-spike-sorted events that the Plexon acquisition system can perform (Spikes Channel AD01, Spike Channel AD02 etc.).
Add the electrode positions
(MRI should have already been imported on a previous step)
A modified channel-file has been included in the downloaded datasets, that contains the positioning of the electrodes on the cortical surface. Users can edit their own file by following the equivalent procedure that is featured on this link.
In order to load the modified channel file, users should:
1. Import Floyd_channel.mat file in Matlab's wokspace
2. Import the struct Floyd_Channels into the channels entry on the datatree (Right click on Plexon channels (33) icon -> File -> Import From Matlab)
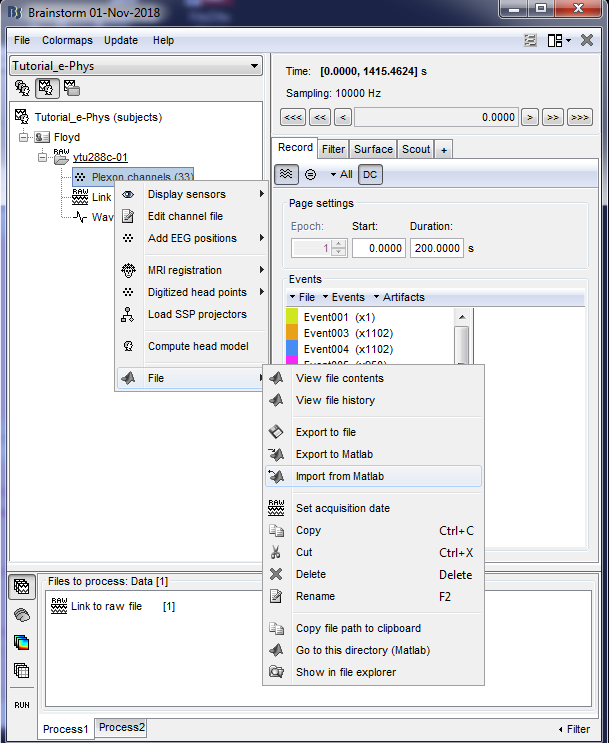
This procedure will be update the positioning of the electrodes. Users can double-click on the Plexon channels (33) icon to display the relative positioning on the cortical surface (It requires the MRI to be already imported).
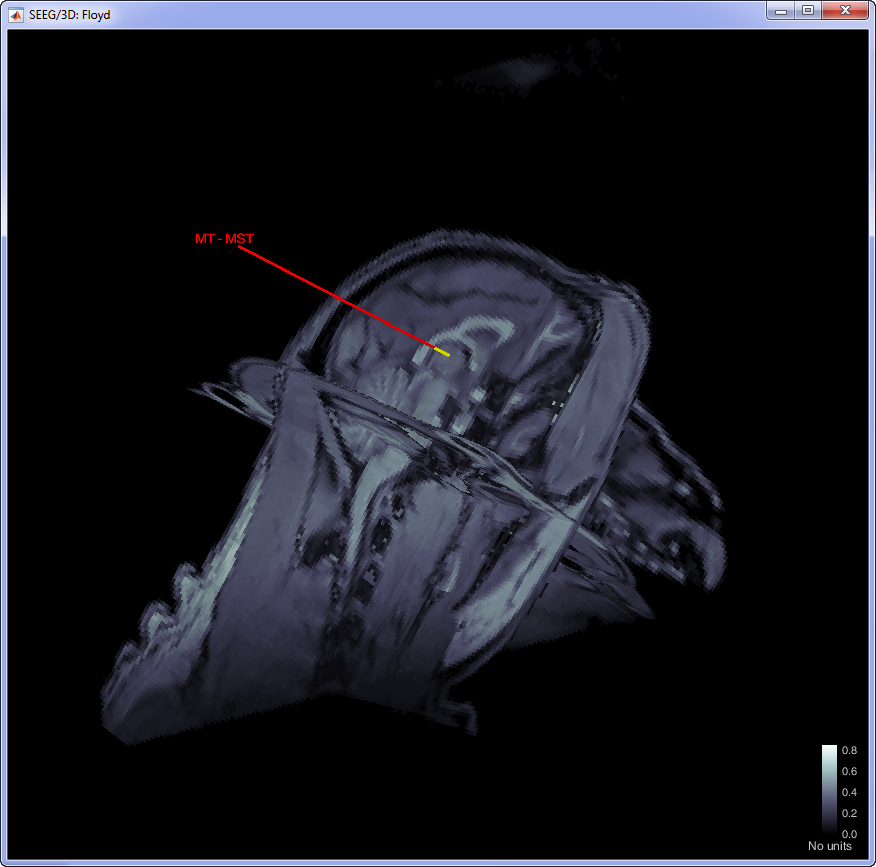
Add additional events
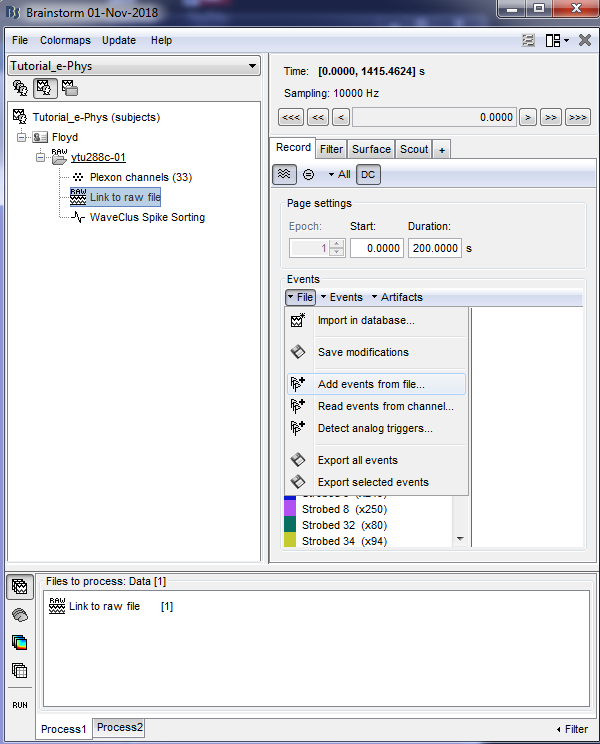
In case users need to input events from a separate file, this can be easily done by clicking on the events panel: File > Add events from file.
Specifically for the tutorial dataset, the stimulus presentation events were coded from a software that was not triggering events on the acquisition system, so they need to be manually imported. Users should first double click on the Link to raw file (this activates the Record tab) and then the events can be added by selecting the file: events_ytu288c-01_converted.mat
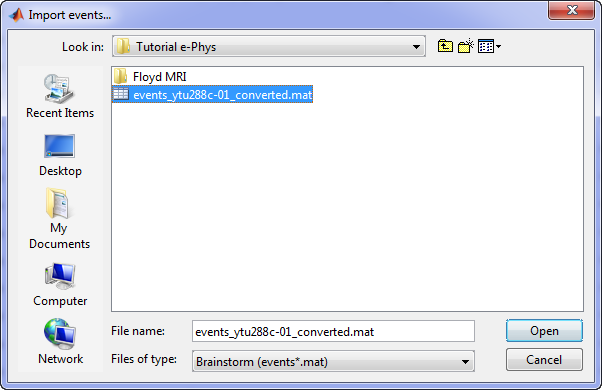
This procedure will add 9 new types of events at the end of the events list:
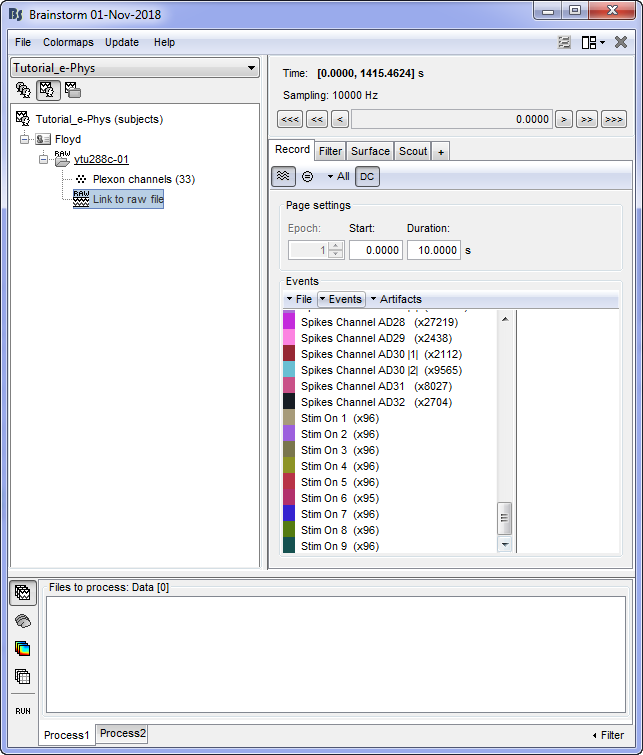
At this stage, users can start performing their analysis. This tutorial has covered all of the options that users will need to fully take advantage of the Invasive Neurophysiogy toolbox (importing datasets, adding the animal's MRI, adding events and channel locations).
Users that want to analyze their datasets without adding MRI scans and/or channel locations can skip those steps.
Importing PL2 Plexon files
To import newer .PL2 Plexon files, you need to download Plexon's public Matlab Offline Files SDK.
Go to Plexon download page: https://plexon.com/software-downloads/
Go to the SDKs tab.
Download OmniPlex and MAP Offline (For reading previously recorded data files) for C/C++ and MATLAB.
Extract the downloaded zip file: OmniPlex-and-MAP-Offline-SDK-Bundle_0.zip.
From the extracted files, find Matlab Offline Files SDK.zip and extract this second zip's content
Copy the extracted folder Matlab Offline Files SDK to a permanent location on your computer.
Add the folder to your MATLAB path. You can do that from the MATLAB command window with the following function, just change it to your proper path: addpath('C:\path\to\Matlab Offline Files SDK');
Once downloaded and added to your Matlab path, you can Review as raw PL2 files from Brainstorm by selecting the file type "EEG: Plexon (*.plx;.pl2)" from the dropdown.
