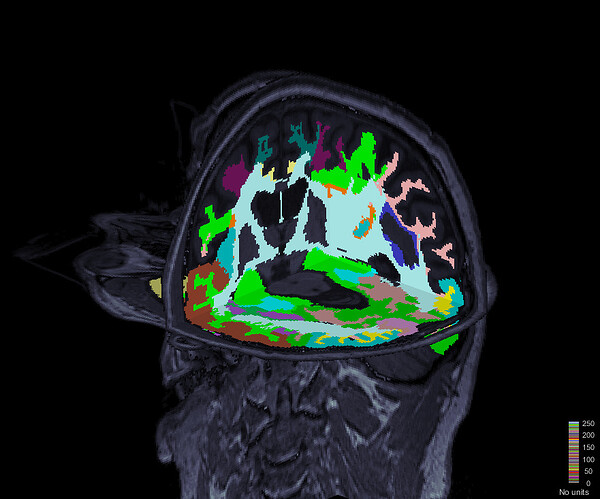
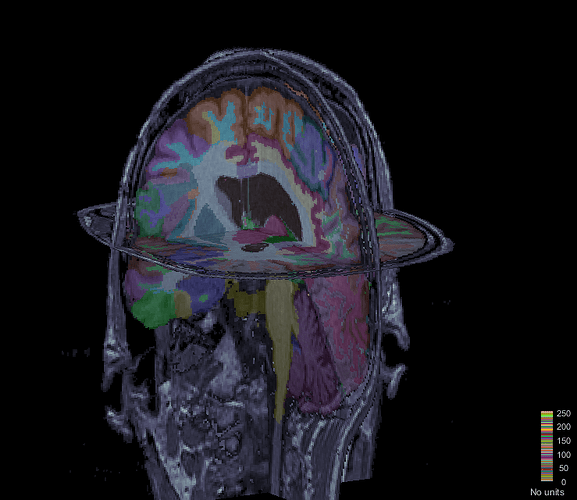
When I am loading a new atlas(e.g. Harvard-Oxford cortical Atlas), brainstorm loading process changing the labels and also not loading all the regions as scout. How I can load a new atlas correctly?
What kind of atlas is it? computed by which program?
The instruction at the end of the FreeSurfer tutorial show how to import a surface atlas (as scouts) or a volume atlas (as separate surfaces, for sub-cortical regions): http://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer
Note that you will not be able to load a volume atlas (labeled MRI file) as a set of scouts on the surface. You can only import atlases that are already defined on the surface.
Hi Francois,
So, Is there any way to load labeled MRI files? Not as scouts, but just for visualization in Anatomy section. Assume that we don’t want to use the defined structures in FreeSurfer.
Thanks
Hossein
Hi Hossein,
You can load a volume atlas with the menu “Import MRI”, like any other volume, but you won’t have named labels, just numbers. This is all mentioned here: http://neuroimage.usc.edu/brainstorm/News#Volume_atlases
Is there anything you need that you don’t manage to get with these menus?
Francois
Thanks.
So right now I can import labeled volume atlases in Brainstorm and display them on MRI. Since I use the labeled MRI from BrainSuite their labels have much wider range than [0 256] as it is in the atlas colormap so I cannot see many clusters.
I tried to define new colormaps from [0 1000], for instance, and changed the colormap range but it shows nothing in that case. In case I want to load a colormap it asks me an LUT file which I don't know how to create or format it based on Brainsuite colormap. Can You help me on this? Briefly, we need a mapping from Brainsuite to Brainstorm.
Also, we don't have cluster names in the scout tab. Is this because we haven't specified that text file you've mentioned in the tutorial?
Hossein
Hi Hossein,
The problem here is that Brainstorm forces the conversion of most loaded volumes (except for MNI atlases) to uint8 values, to make them faster to read/write/process. For the level of precision we need in anatomical images, this is precise enough, but of course, this is not adapted to atlases with more than 256 regions… We need to adapt this block of code to make this possible:
Suggestions? For now you can simply comment it out, and try to add custom colormap with more values too (256 by default). But we’ll need to discuss this and make a choice for the common distribution here.
Are you really hoping to identify and read 1000 different ROIs in an atlas?
What is the purpose of what you are currently doing?
Francois
PS: Can you send me your files, so I can look at them?
Thanks
Francois,
That problem has been solved. We don't have more than 256 clusters.I just needed to change the threshold value and see all clusters.
However, There is another issue. Right now in anatomy section, I can depict labeled MRIs as
I can change the transparency of the MRI and labels. However, In this section, I cannot show the SEEG electrodes. If I want to show it in "Functional data", instead of labels, the estimated sources will be projected on MRI slices and labels will be depicted as spheres and volumes, which is not what we want.
So briefly, Is there any way to have SEEG electrodes in this figure?
Thanks
Hossein
Hi Hossein,
Start by displaying the SEEG on the T1, then you can add a layer to the MRI display:
OverlayType = 'Anatomy';
OverlayFile = '/.../brainstorm_db\ProtocolName/anat/subjectname/subjectimage_atlas.mat';
isOk = panel_surface('SetSurfaceData', gcf, 1, OverlayType, OverlayFile, 0);
Cheers,
Francois
Thank you so much.
So, currently for imported labeled MRIs, we have a 3d matrix (Cube) that shows the label of each voxel in (Slice Coordinate probably or maybe SCS!). Also, for each label ID we have a name. Is there any possibility to show the label name (like “Insula (wm)”, “R. Temporal Lobe (wm)”,…) in the MriViewer online?
Right now we can show the coordinates when we click on a point and since we have those labels and their corresponding names, it should be possible? Isn’t it? I tried to modify the code for “figure_mri.m” to add this but it is in Java and near 3000 lines, so I gave up.
Also, I built the necessary .txt file for loading the label names with MRIs but it didn’t work. I’ll send you all files by email and if you have time please take a look at it. I really appreciate your help.
Hi Hossein,
Yes, this is doable. But this feature should be added properly:
- The volume atlas and the label names should be read automatically when importing a folder generated by BrainSuite (import_anatomy_bs.m)
- db_template.m: Add a field “AtlasLabels” in ‘mrimat’
- import_anatomy_bs.m: Load the atlas volume, load the atlas labels and add them in the MRI structure of the atlas
- tree_callbacks.m: Add a popup menu to load the atlas labels in (around line 900)
- figure_mri.m: Add a popup menu to load the atlas labels in DisplayFigurePopup()
- in_mri_bst.m: Add a test to add “AtlasLabels” if not available
- bst_memory.m > LoadMri(): Not sure if we have to explicitly add code to load AtlasLabel, maybe not
- figure_mri.m: Display the label of the current point in the MRI viewer (function UpdateCoordinates, line 1248)
- Edit the online tutorial to describe these new features: http://neuroimage.usc.edu/brainstorm/Tutorials/SegBrainSuite
You can get started, open a PR on github when you have code to submit and I will review it.
Cheers,
Francois
FYI: We improved significantly the tools available in Brainstorm for processing and visualizing SEEG and ECOG data, including new options for volume coregistration. They are now documented in a new tutorial:
http://neuroimage.usc.edu/brainstorm/Tutorials/Epileptogenicity