Hi all,
I have three questions regarding EEG channel location.
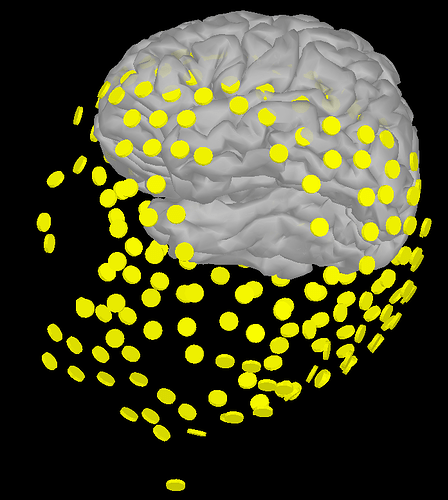
I am using preprocessed EEG fie in .mat and .set which was recorded with BioSemi 256 acticap. Each data set contains channel locations and names (according to Biosemi naming conventions) but .locs file was not available. I am assuming that I need .loc file to import the channel location to Brainstorm. So, I manually edited the channel locations using Channel Edit in Brainstorm. When I plot the modified channel location on cortex, it looks like this (below). So, my three questions are,
- Could anyone please let me know how I can fix this?
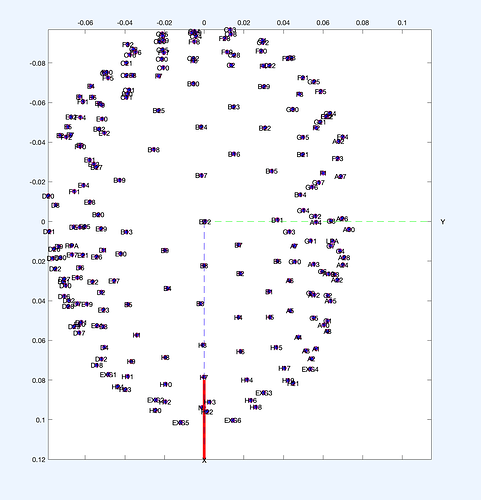
- Also, is there any way that I can export and convert the xyz coordinates I can find in .set file into .loc file?
- But I guess this will introduce other problem. As they are all preprocessed, each channel numbers are different across participants and when I compare xyz of each participants, they are slightly different.
- Then, do I have to edit it for each participant? Considering the number of channels and number of participants, I don't see this is very effective. So, would it be better to contact the author to get .loc file? I have googled it for days to no avail.
I would rally appreciate your help. Thank you so much.
Just for your information.