|
Size: 3376
Comment:
|
Size: 2051
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| . . . {{http://brainsuite.usc.edu/images/brainsplash_sm.jpg}} |
= BrainSuite = {{attachment:BrainSuite13_650px.png}} |
| Line 5: | Line 4: |
| '''BrainSuite homepage has moved: '''[[http://www.loni.ucla.edu/Software/BrainSuite|click here]]'''.''' '''BrainSuite forum: '''[[http://forum.loni.ucla.edu/forumdisplay.php?f=9|click here]]'''.''' | [[http://brainsuite.org/|BrainSuite]] is a collection of image analysis tools designed to process magnetic resonance images (MRI) of the human head. !BrainSuite13 provides an automatic sequence to extract cortical surface mesh models from the MRI, tools to register these to a labeled atlas to define anatomical regions of interest, and tools for processing diffusion imaging data. !BrainSuite13 also contains visualization tools for exploring these data, and can produce interactive maps of regional connectivity. |
| Line 7: | Line 6: |
| . '''BrainSuite documentation: '''[[http://www.loni.ucla.edu/~shattuck/brainsuite/|click here]]'''.''' | BrainSuite is collaboratively developed between David Shattuck at the [[http://bmap.ucla.edu|Ahmanson-Lovelace Brain Mapping Center]] at UCLA and Richard Leahy in the [[http://neuroimage.usc.edu|Biomedical Imaging Group]], in collaboration with the [[http://www.loni.usc.edu/|Laboratory of Neuro Imaging]], at USC. Other contributors to the development of the methods embodied in this software include: Anand A. Joshi, Chitresh Bhushan, Justin P. Haldar, Belma Dogdas, Stephanie Sandor-Leahy, Bijan Timsari, and Paul M. Thompson. |
| Line 9: | Line 8: |
| == BrainSuite == BrainSuite is a magnetic resonance (MR) image analysis tool designed for identifying tissue types and surfaces in MR images of the human head. |
!BrainSuite features include: |
| Line 12: | Line 10: |
| BrainSuite was specifically designed to guide its users through the process of cortical surface extraction. We have written the software to require minimal user interaction and with the goal of completing the entire process of extracting a topologically spherical cortical surface from a raw MR volume within several minutes on a modern workstation. The individual components of BrainSuite may also be used for soft tissue, skull and scalp segmentation and for surface analysis and visualization. BrainSuite was written in Microsoft Visual C++ using the Microsoft Foundation Classes for its graphical user interface and the OpenGL library for rendering. BrainSuite runs under the Windows 2000 and Windows XP Professional operating systems. | * Automated cortical surface extraction from T1-weighted MRI. Tools include brain surface extraction, bias field correction, voxel classification, cerebrum labeling, and surface generation * Automated surface/volume registration software * Tools for correction of B0 effects in diffusion data and coregistration to structural MRI * Tools for processing of diffusion data including ODF and tensor fitting and tractography * Sophisticated tools for visualizing and exploring MRI data, diffusion data, tractography, and connectivity |
| Line 14: | Line 16: |
| BrainSuite has been written to be compatible with our [[Brainsuite/../brainstorm|BrainStorm Software]] package for the analysis of MEG and EEG data. Cortical surfaces extracted using BrainSuite can be used for part of a cortically constrained minimum norm imaging; brain, skull and scalp surfaces can be used to solve forward calculations using BEM or FEM functions in BrainStorm. BrainStorm also includes utilities to downsample surface tessellations and build FEM meshes from segmented volumes and surfaces. | The currently released version of the software (BrainSuite) includes graphical user interface versions for Mac OSX and Windows and command line versions for Mac OSX, Windows and Linux. More details can be found on following links: |
| Line 16: | Line 18: |
| BrainSuite was written by Dr. David W. Shattuck and is produced and distributed as a collaborative project between the [[http://www.loni.ucla.edu/index.shtml|Laboratory of Neuro Imaging]] at the University of California Los Angeles (Director: Dr. Arthur W. Toga) and the [[Brainsuite/../|Biomedical Imaging Research Group]] at the University of Southern California (Director: Dr. Richard M. Leahy). The work is supported by grants from the National Institute of Biomedical Imaging and Bioengineering (R01 EB002010, PI: R.M. Leahy) and from the National Center for Research Resources (P41 RR013642, PI: A.W. Toga). | * [[http://brainsuite.org/|Brainsuite Documentation]] * [[http://brainsuite.org/download/|BrainSuite Downloads]] |
| Line 18: | Line 21: |
| Contributors to the development of the methods embodied in this software include: David Shattuck, Anand Joshi, Belma Dogdas, Stephanie Sandor-Leahy, Bijan Timsari, Paul Thompson and Richard M. Leahy. BrainSuite '''features''' include: * Sophisticated visualization tools, such as MRI visualization in 3 orthogonal views (either seperately or in 3D view), and overlayed surface visualization of cortex, skull, and scalp * Cortical surface extraction, using a multi-stage user friendly approach. Tools include brain surface extraction, bias field correction, voxel classification, cerebellum removal, and surface generation * Topological correction of cortical surfaces, which uses a graph-based approach to remove topological defects (handles and holes) and ensure a tessellation with spherical topology * Parameterization of generated cortical surfaces, minimizing a harmonic energy functional in the p-norm * Skull and scalp surface extraction |
<<BR>><<BR>> |
BrainSuite
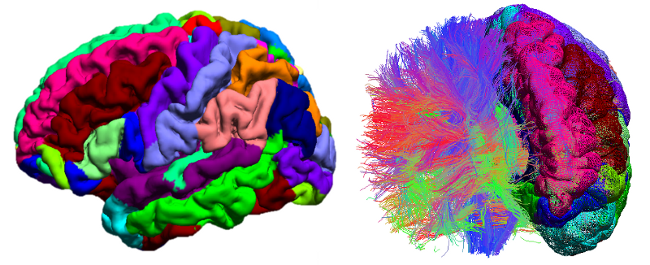
BrainSuite is a collection of image analysis tools designed to process magnetic resonance images (MRI) of the human head. BrainSuite13 provides an automatic sequence to extract cortical surface mesh models from the MRI, tools to register these to a labeled atlas to define anatomical regions of interest, and tools for processing diffusion imaging data. BrainSuite13 also contains visualization tools for exploring these data, and can produce interactive maps of regional connectivity.
BrainSuite is collaboratively developed between David Shattuck at the Ahmanson-Lovelace Brain Mapping Center at UCLA and Richard Leahy in the Biomedical Imaging Group, in collaboration with the Laboratory of Neuro Imaging, at USC. Other contributors to the development of the methods embodied in this software include: Anand A. Joshi, Chitresh Bhushan, Justin P. Haldar, Belma Dogdas, Stephanie Sandor-Leahy, Bijan Timsari, and Paul M. Thompson.
BrainSuite features include:
- Automated cortical surface extraction from T1-weighted MRI. Tools include brain surface extraction, bias field correction, voxel classification, cerebrum labeling, and surface generation
- Automated surface/volume registration software
- Tools for correction of B0 effects in diffusion data and coregistration to structural MRI
- Tools for processing of diffusion data including ODF and tensor fitting and tractography
- Sophisticated tools for visualizing and exploring MRI data, diffusion data, tractography, and connectivity
The currently released version of the software (BrainSuite) includes graphical user interface versions for Mac OSX and Windows and command line versions for Mac OSX, Windows and Linux. More details can be found on following links:
