Corticomuscular coherence (MEG)
[TUTORIAL UNDER DEVELOPMENT: NOT READY FOR PUBLIC USE]
Authors: Raymundo Cassani, Francois Tadel & Sylvain Baillet.
Corticomuscular coherence measures a degree of similarity between electrophysiological signals (MEG, EEG, ECoG sensor traces or source time series, especially over the contralateral motor cortex) and the EMG signal recorded from muscle activity during voluntary movement. This signal similarity is due mainly to the descending communication along corticospinal pathways between primary motor cortex (M1) and muscles. For consistency and reproducibility purposes across major software toolkits, the present tutorial replicates the processing pipeline "Analysis of corticomuscular coherence" by FieldTrip.
Contents
- Background
- Dataset description
- Download and installation
- Importing and processing anatomy data
- Review the MEG and EMG recordings
- Pre-process
- Importing data epochs
- Coherence estimation (sensor level)
- MEG source imaging
- Estimating source-wise coherence
- Brain-wise coherence estimation (no scouts)
- Script
- Additional documentation
Background
Coherence measures the linear relationship between two signals in the frequency domain. Previous studies (Conway et al., 1995, Kilner et al., 2000) have reported cortico-muscular coherence effects in the 15–30 Hz range during maintained voluntary contractions.
IMAGE OF EXPERIMENT, SIGNALS and COHERENCE
Dataset description
The dataset comprises recordings from MEG (151-channel CTF MEG system) and bipolar EMG (from left and right extensor carpi radialis longus muscles) from one participant who was tasked to lift their hand and exert a constant force against a lever for about 10 seconds. The force was monitored by strain gauges on the lever. The participant performed two blocks of 25 trials using either the left or right wrist. EOG signals were also recorded, which will be useful for detection and attenuation of ocular artifacts. We will analyze the data from the left-wrist trials in the present tutorial. Replicating the pipeline with right-wrist data is a good exercise to do next!
Download and installation
Requirements: Please make sure you have completed the get-started tutorials and that you have a working copy of Brainstorm installed on your computer. In addition, you need to install CAT12 plugin in Brainstorm. CAT12 will be used for MRI segmentation.
Download the dataset:
Download SubjectCMC.zip from FieldTrip's FTP server:
ftp://ftp.fieldtriptoolbox.org/pub/fieldtrip/tutorial/SubjectCMC.zip- Unzip the .zip in a folder not located in any of current Brainstorm's folders (the app per se or its database folder).
Brainstorm:
- Launch Brainstorm (via Matlab's command line or use Brainstorm's Matlab-free stand-alone version).
Select the menu File > Create new protocol. Name it TutorialCMC and select the options:
No, use individual anatomy,
No, use one channel file per acquisition run.
The next sections describe how to import the participant's anatomical data, review raw data, manage event markers, pre-process EMG and MEG signals, epoch and import recordings for further analyzes, with a focus on computing coherence at the sensor (scalp) and brain map (sources) levels.
Importing and processing anatomy data
Right-click on the newly created TutorialCMC node in your Brainstorm data tree then: New subject > Subject01.
Keep the default options defined for the study (aka "protocol" in Brainstorm's jargon).Switch to the Anatomy view of the protocol.
Right-click on the Subject01 node then Import MRI:
Select the adequate file format from the pull-down menu: All MRI file (subject space)
Select the file: SubjectCMC/SubjectCMC.mri
This will open the MRI viewer showing the coronal, sagittal and axial views of the MRI. In addition, three anatomical fiducials: left and right pre-auricular points (LPA and RPA), and nasion (NAS) are automatically identifed. These fiducials are located near the left/right ears and just above the nose respectively. Click on Save.
We then need to segment the head tissues to obtain the surfaces required to derive a realistic MEG head model (aka "forward model"). Here, we will perform MRI segmentation with CAT12, this process takes between 30 to 60 minutes.
Right-click on the SubjectCMC MRI node, then MRI segmentation > CAT12: Cortex, atlases, tissues. This will prompt a series of windows to set the parameters for the MRI segmentation, use the following parameters:
Number of vertices on the cortex surface use 15000
Compute anatomical parcellations? select Yes
Compute cortical maps select Yes
Select all the tissues (scalp, skull, csf, gray and white).
Click OK.
For the option Generate surface meshes select No.
After the segmentation is complete, a tissues node will be shown in the tree.
Rick-click on the tissues node and select Generate triangular meshes.
- Select the 5 layers to mesh.
- Use the default parameters:
number of vertices: 10,000
erode factor: 0
fill holes factor: 2
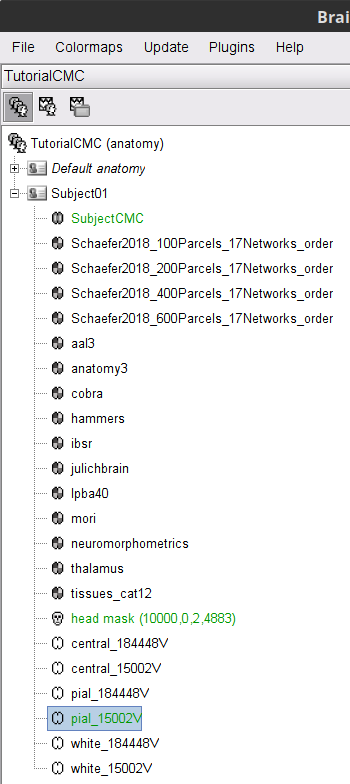
A set of (head and brain) surface files are now available for further head modelling (see below).
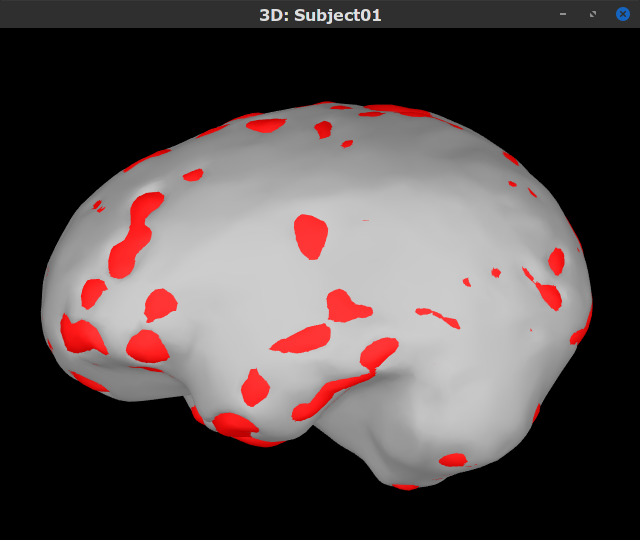
You can display the surfaces by double-clicking on these new nodes. There are a couple of issues with the structural data available from this tutorial. Note how the cortex (shown in red) overlaps with the innerskull surface (shown in gray). For this reason, the BEM forward model cannot be derived accurately with OpenMEEG. We will use an analytical approximation using the overlapping-spheres method, which in MEG has been shown to be adequately accurate for most studies. Note also how the cortex and white surfaces obtained do not register accurately with the cortical surface. We will therefore use a volume-based source estimation approach based on a volumic grid of elementary MEG source across the cerebrum (not a surface-constrained source model). We encourage users to try CAT12 or FreeSurfer, also directely from Brainstorm, to obtain surface segmentations of higher quality before running BEM modelling (as an extra exercise).
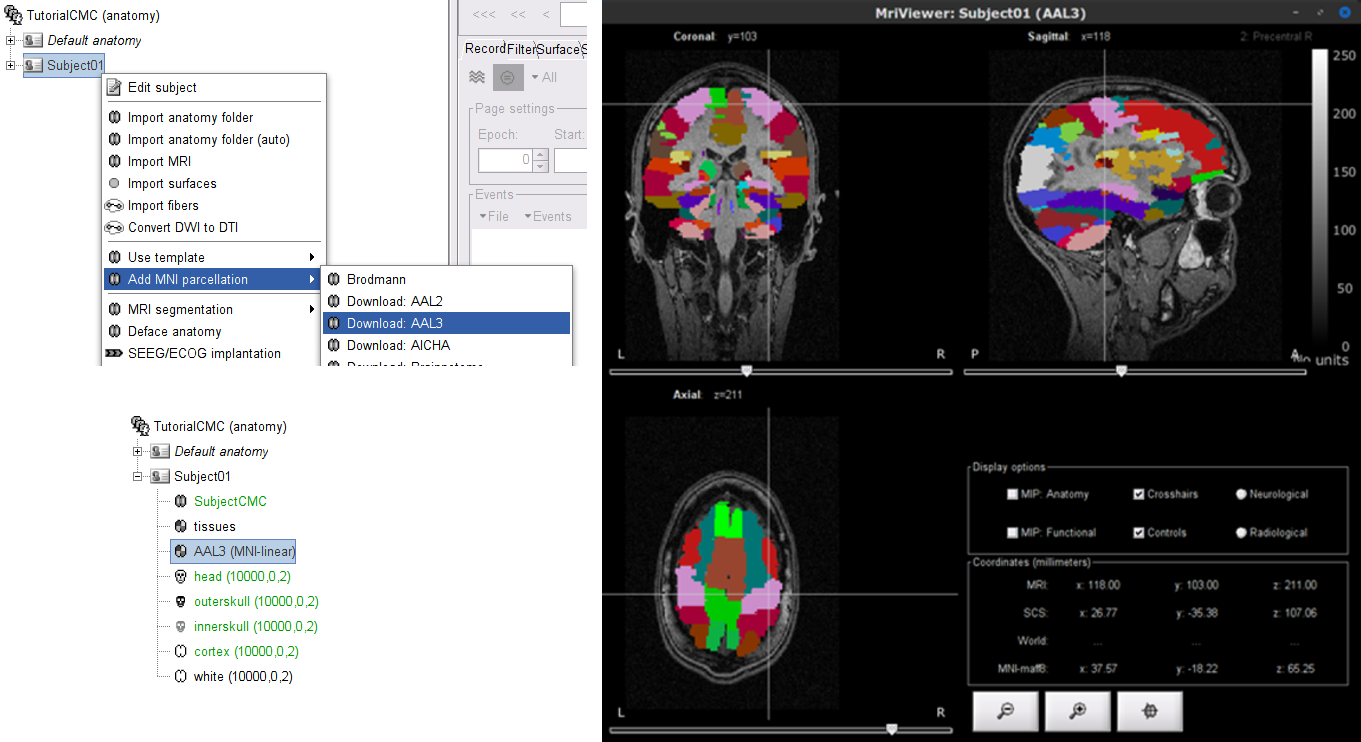
As the imported anatomy data is normalized in the MNI space, it is possible to apply use MNI parcellation templates to define anatomical regions of the brain of the subject. These anatomical regions can be used to create volume and surface scouts, which are convenient when performing the coherence analysis in the source level. Let's add the AAL3 parcellation to the imported data.
Right-click on Subject01 then go to the menu Add MNI parcellation > AAL3. The menu will appear as Download: AAL3 if the atlas is not in your system. Once the MNI atlas is downloaded, an atlas node (ICON) appears in the database explorer and the atlas is displayed in the the MRI viewer.
Review the MEG and EMG recordings
Link the recordings to Brainstorm's database
Switch now to the Functional data view (X button).
Right-click on the Subject01 node then Review raw file:
Select the file format of current data from the pulldown menu options: MEG/EEG: CTF(*.ds; *.meg4; *.res4)
Select the file: SubjectCMC.ds
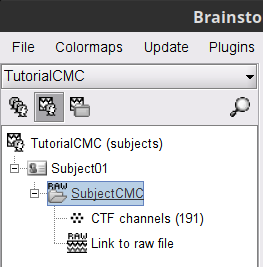
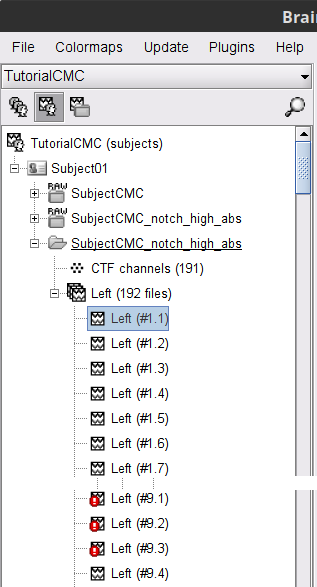
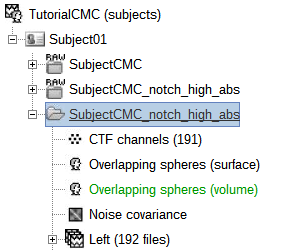
A new folder is now created in Brainstorm's database explorer and contains:
SubjectCMC: a folder that provides access to the MEG dataset. Note the "RAW" tag over the icon of the folder, indicating the files contain unprocessed, continuous data.
CTF channels (191): a node containing channel information with all channel types, names locations, etc. The number of channels available (MEG, EMG, EOG etc.) is indicated between parentheses (here, 191).
Link to raw file provides access to to the original data file. All the relevant metadata was read from the dataset and copied inside the node itself (e.g., sampling rate, number of time samples, event markers). Note that Brainstorm's logic is not to import/duplicate the raw unprocessed data directly into the database. Instead, Brainstorm provides a link to that raw file for further review and data extraction (more information).
Display MEG helmet and sensors
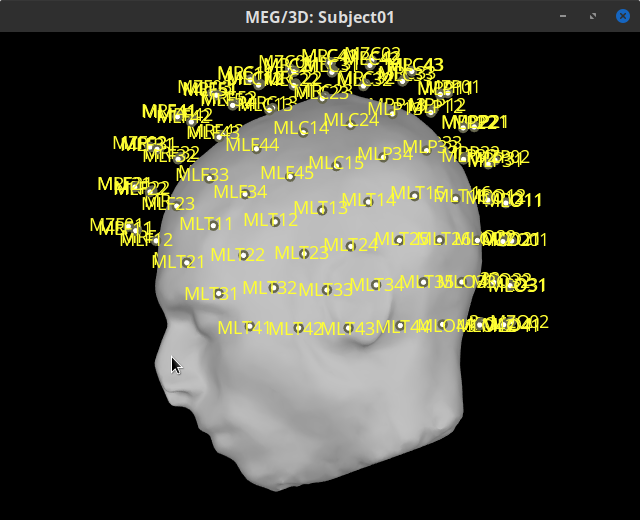
Right-click on the CTF channels (191) node, then select Display sensors > CTF helmet from the contextual menu and Display sensors > MEG. This will open a new display window showing the inner surface of the MEG helmet, and the lo MEG sensors respectively. Try additional display menus.
Reviewing continuous recordings
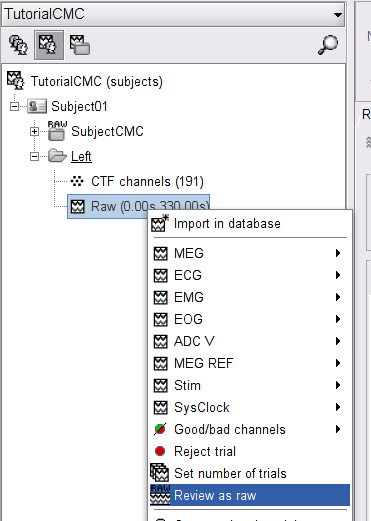
Right-click on the Link to raw file node, then Switch epoched/continuous to convert the file to continuous, a technical detail proper to CTF file formatting.
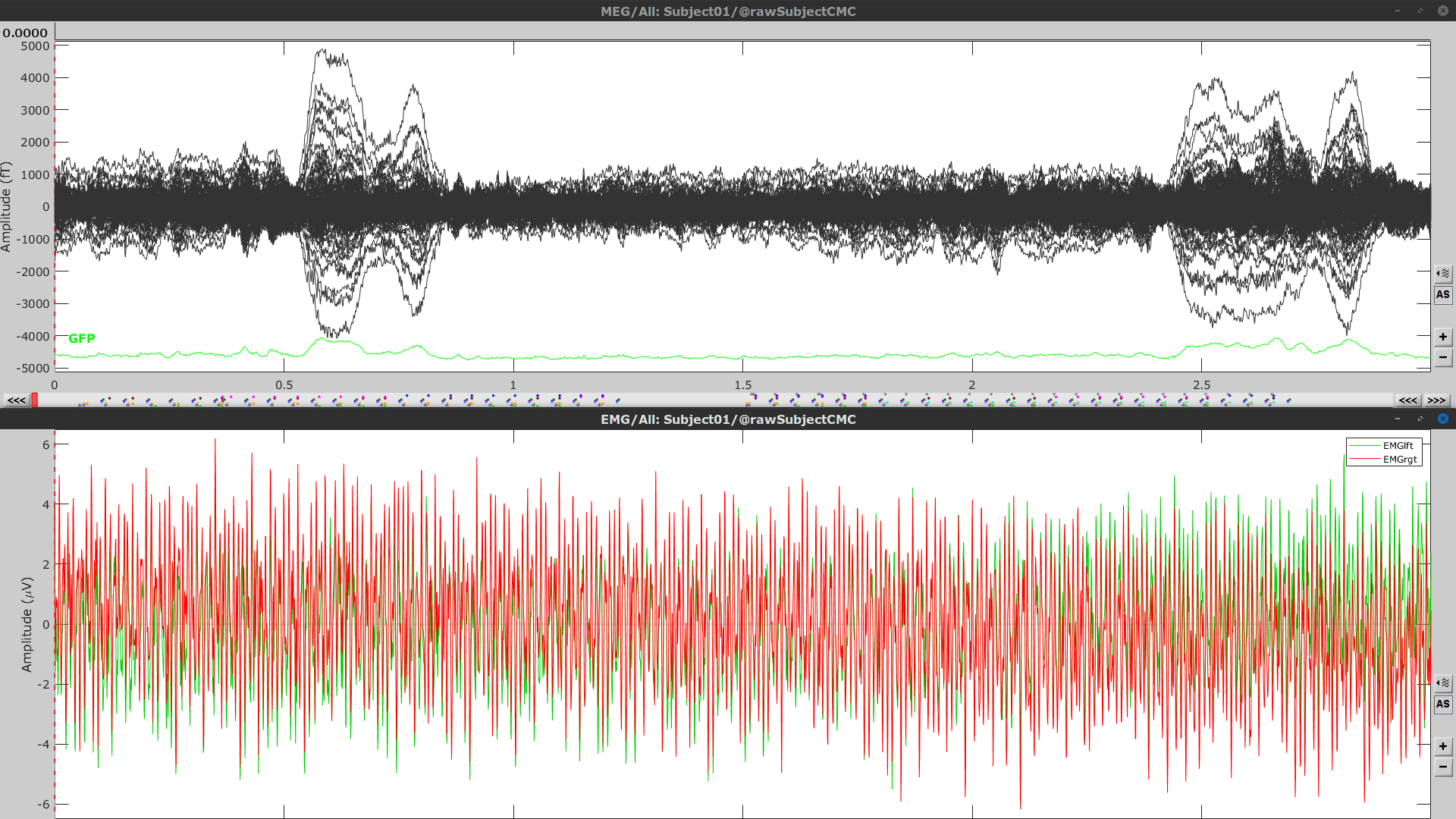
Right-click again on the Link to raw file node, then MEG > Display time series (or double-click on the node). This will open a new visualization window to explore data time series, also enabling the Time panel and the Record tab in the main Brainstorm window (see how to best use all controls in this panel and tab to explore data time series).
We will also display EMG traces by right-clicking on the Link to raw file node, then EMG > Display time series.
Event markers
The colored dots above the data time series indicate event markers (or triggers) saved with this dataset. The trial onset information of the left-wrist and right-wrist trials is saved in an auxiliary channel of the raw data named Stim. To add these markers, these events need to be decoded as follows:
While the time series figure is open, go to the Record tab and File > Read events from channel. From the options of the Read from channel process window, set Event channels = Stim, select Value, and click Run.
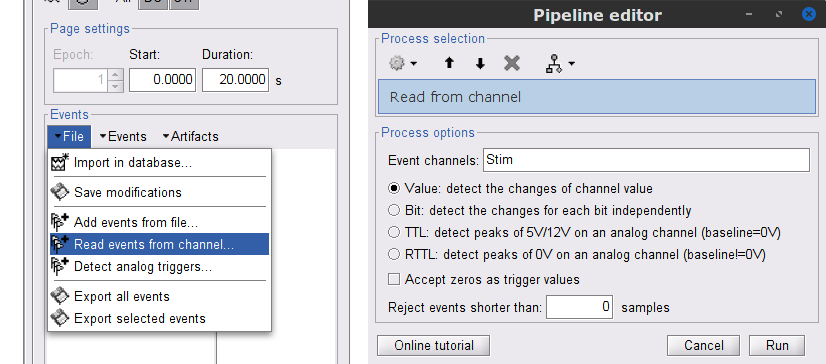
This procedure creates new event markers now shown in the Events section of the tab. along with previous event categories. In this tutorial, we will only use events U1 through U25, which correspond to how each of the 25 left-wrist trials had been encoded in the study. We will now delete other events of no interest, and merge the left trial events under a single event category, for convenience.
Delete other events: select the events to delete in the event box/list with Ctrl+click, then in the menu Events > Delete group and confirm. Alternatively, you can selected all events with Ctrl+A and deselect the U1 to U25 events by clicking on them.
To make sure we reproduce FieldTrip's tutorial, we need to reject trial #7: Select events U1 to U6 and U8 to U25, then from the Events menu, select Merge group and type in new label (Left_01) to describe this as the left-wrist condition.
These events correspond to the beginning of 10-s trials of left-wrist movements. We will compute coherence over 1-s epochs over the first 8 s of each trial. To that purpose, we will now create extra events to define these epochs.
Duplicate 7 times the Left events by selecting Duplicate group in the Events menu. The groups Left_02 to Left_08 will be created.
For each copy of the Left events, we will add a time offset of 1 s for Left02, 2 s for Left03, and so on. Select the Left event group to add a 1,000 ms time offset, by going to the menu Events > Add time offset, enter 1,000 in the text box. Repeat for each other group, entering 2,000, then 3,000 etc.
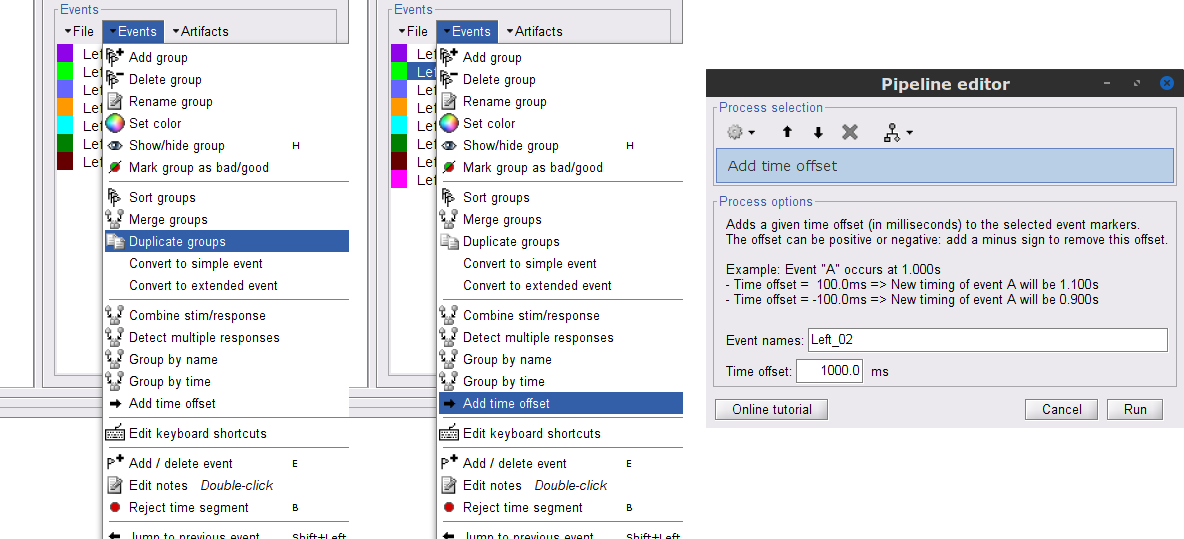
Once done for Left_08, merge all these Left* events into a single Left category, and select Save modifications in the File menu in the Record tab.
Pre-process
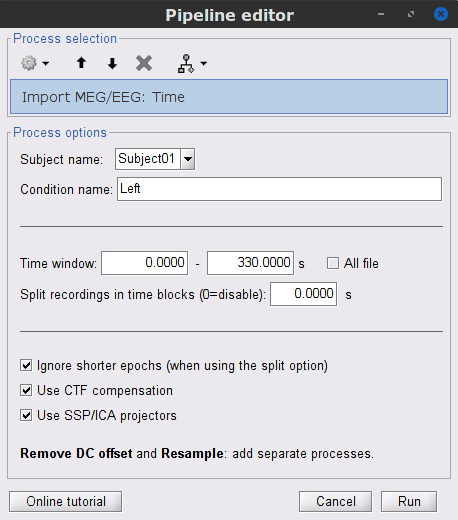
In this tutorial, we will analyze only the Left trials (left-wrist extensions). In the following sections, we will process only the first 330 s of the recordings, where the left-wrist trials were performed.
Another idiosyncrasy of the present dataset is that the CTF MEG data were saved without the desired 3-rd order gradient compensation for optimal denoising. We will now apply this compensation as follows:
In the Process1 box: Drag and drop the Link to raw file node.
Run process Artifacts > Apply SSP & CTF compensation:
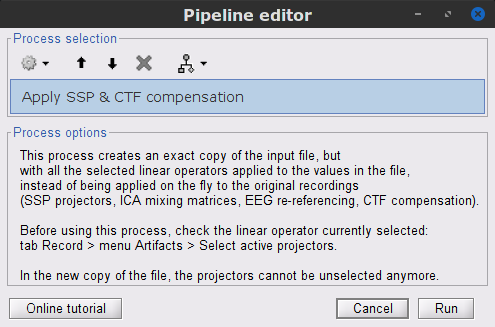
This process creates the SubjectCMC_clean folder that contains a copy of the channel file and a link to the raw file Raw | clean, which points to the original data and to the fact that the 3-rd order gradient compensation will be applied. Brainstorm does not create a physical copy of the actual, large dataset at this stage.
Removal of power line artifacts
We will start with identifying the spectral components of power line contamination of MEG and EMG recordings.
In the Process1 box: Drag and drop the Raw | clean node.
Run process Frequency > Power spectrum density (Welch):
Time window: 0 - 330 s
Window length=10 s
Overlap=50%
Sensor types=`MEG, EMG
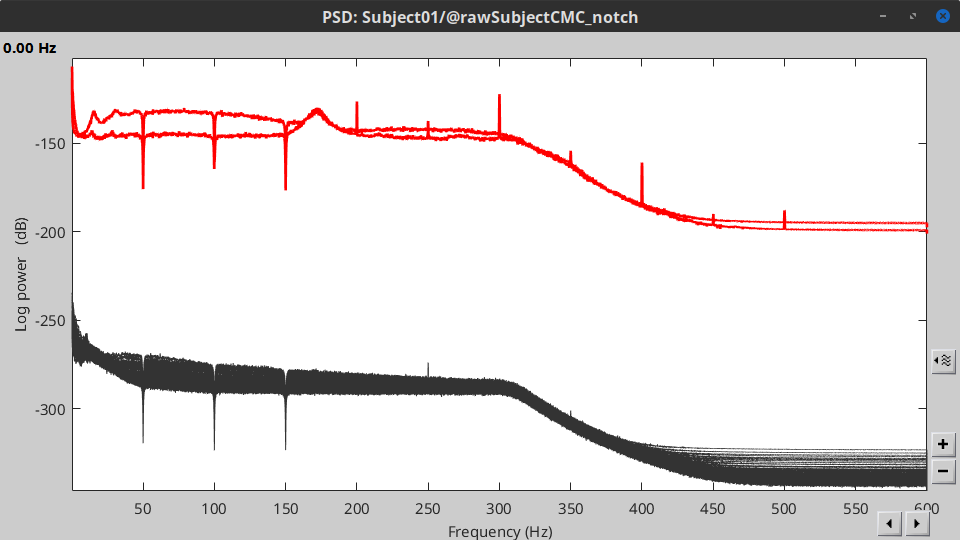
Double-click on the new PSD file to visualize the power spectrum density of the data.
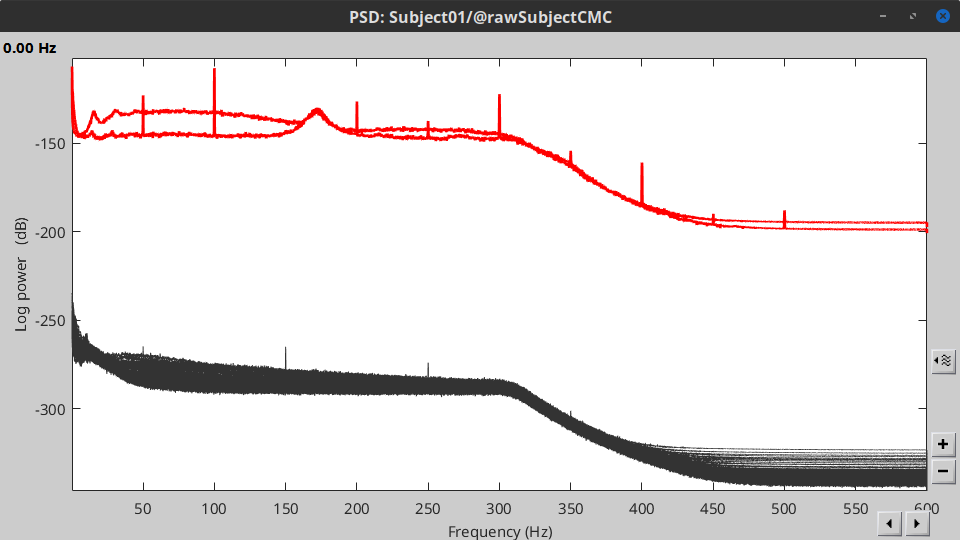
- The PSD plot shows two groups of sensors: EMG (highlighted in red above) and the MEG spectra below. Peaks at 50Hz and harmonics (150, 200Hz and above; European power line main and harmonics) are clearly visible. We will use notch filters to attenuate power line contaminants at 50, 150 and 200 Hz.
In the Process1 box: Drag and drop the Raw | clean node.
Run the process Pre-processing > Notch filter with:
Sensor types = MEG, EMG
Frequencies to remove (Hz) = 50, 100, 150
A new raw folder named SubjectCMC_clean_notch is created. Estimate the PSD of these signals to appreciate the effect of the notch filters applied. As above, please remember to indicate a Time window restricted from 0 to 330 s in the options of the PSD process.
EMG pre-processing
Two typical pre-processing steps for EMG consist in high-pass filtering and rectifying.
In the Process1 box: drag and drop the Raw | notch(50Hz 100Hz 150Hz) node.
Add the process Pre-process > Band-pass filter
Sensor types = EMG
Lower cutoff frequency = 10 Hz
Upper cutoff frequency = 0 Hz
Add the process Pre-process > Absolute values
Sensor types = EMG
- Run the pipeline
Two new folders SubjectCMC_clean_notch_high and SubjectCMC_clean_notch_high_abs are added to Brainstorm's database explorer. We can now safely delete folders that are not needed anymore:
Delete SubjectCMC_clean_notch and SubjectCMC_clean_notch_high by selecting both before pressing Delete (or right-click File > Delete).
MEG pre-processing
We need to remove more artifacts from the MEG traces via the:
Detection and removal of stereotypical artifacts with SSP
Detection of noisy (bad) data segments.
Detection and removal of artifacts with SSP (Signal Space Projection)
Stereotypical artifacts such eye blinks and heartbeats can be identified from their respective characteristic spatial distributions. Their contaminationn of MEG signals can then be attenuated specifically using Signal-Space Projections (SSPs). For more details, consult the dedicated tutorials about the detection and removal of artifacts with SSP. The present tutorial dataset features an EOG channel but no ECG. We will perform only the removal of eye blinks.
Display the MEG and EOG time series: Right-click on the pre-processed (for EMG) continuous file Raw | clean | notch(... (in the SubjectCMC_clean_notch_high_abs folder) then MEG > Display time series and EOG > Display time series.
In the Events section of the Record tab, select Artifacts > Detect eye blinks, and use the parameters:
Channel name= EOG
Time window = 0 - 330 s
Event name = blink
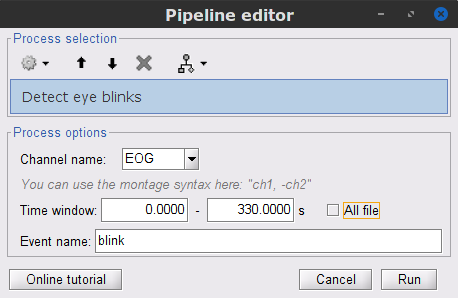
Three categories of blink events are created. Review the traces of EOG channels around a few of these events to ascertain they are related to eye blinks. In the present case, we note that the blink group contains genuine eye blinks, and that groups blink2 and blink3 capture saccade events.
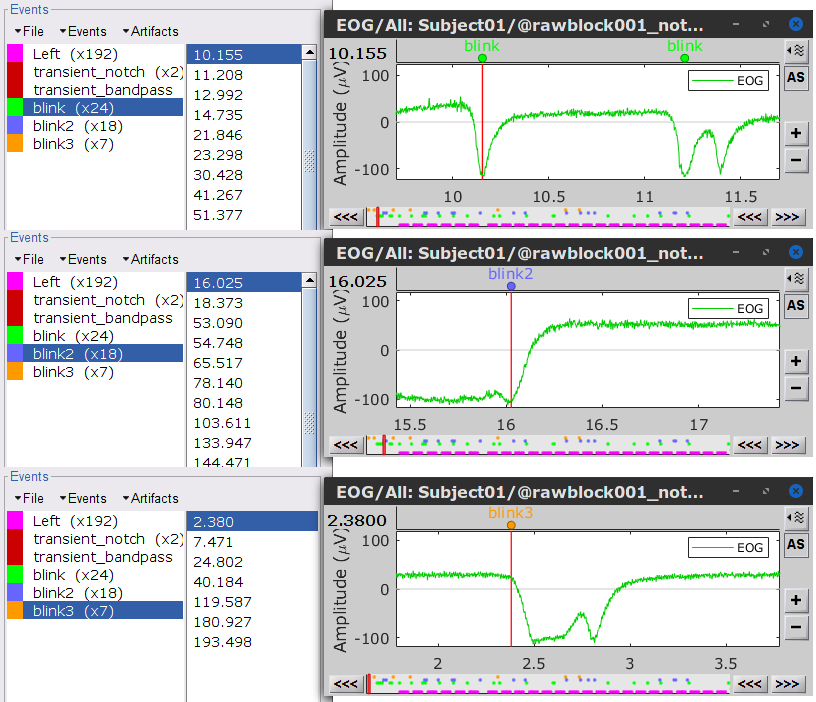
To remove blink artifacts with SSP go to Artifacts > SSP: Eye blinks, and use the parameters:
Event name=blink
Sensors=MEG
Check Compute using existing SSP/ICA projectors
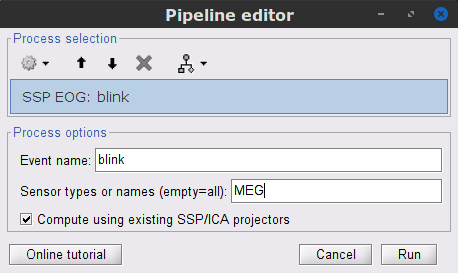
- Display the time series and topographies of the first two (dominant) SSP components identified. In the present case, only the first SSP component can be clearly related to blinks. Select only component #1 for removal.
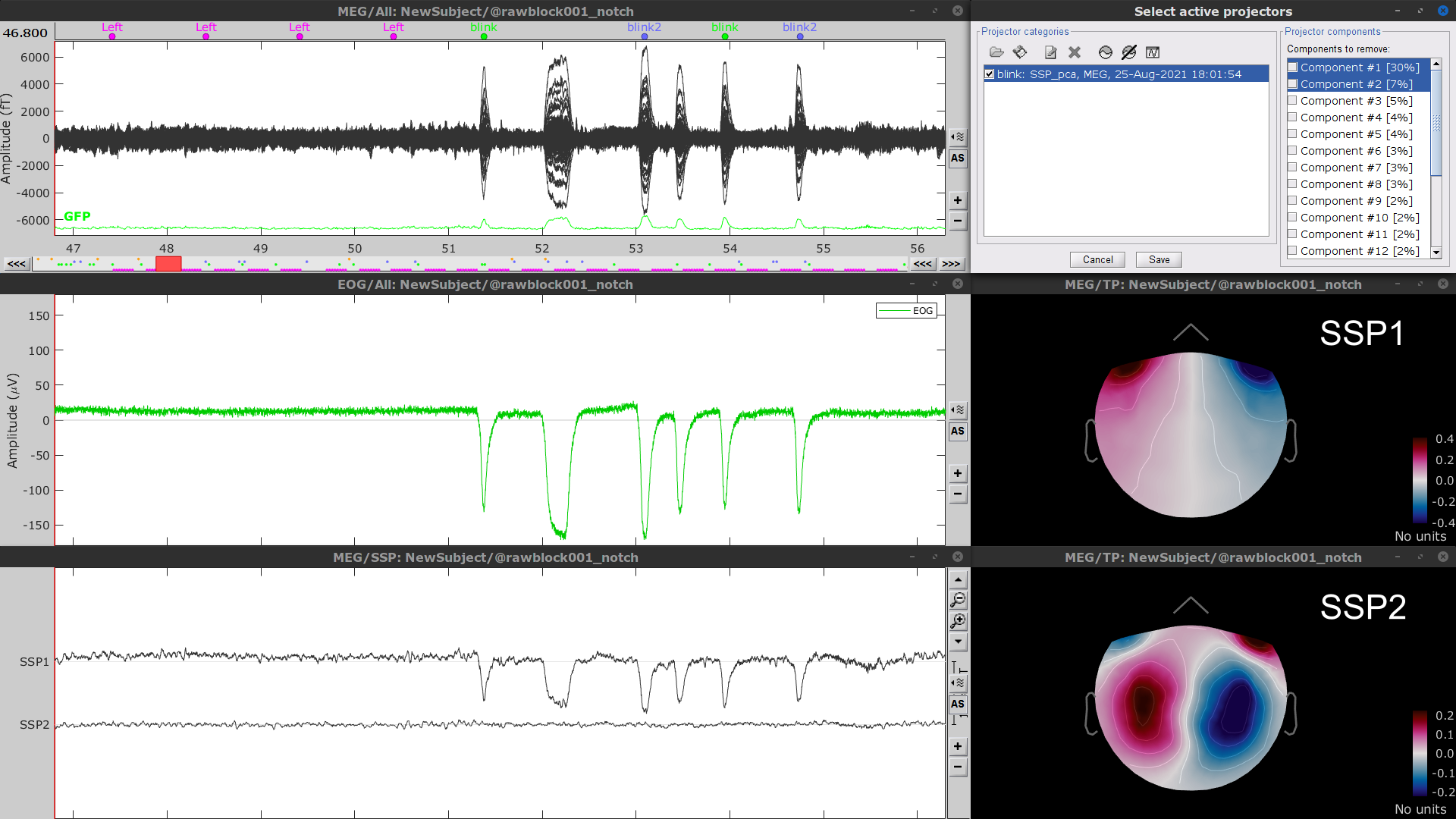
Follow the same procedure for the other blink events (blink2 and blink3). As mentioned above, none of the first two SSP components seem to be related to ocular artifacts. The figure below shows the visualization of the first two components for the blink2 group.
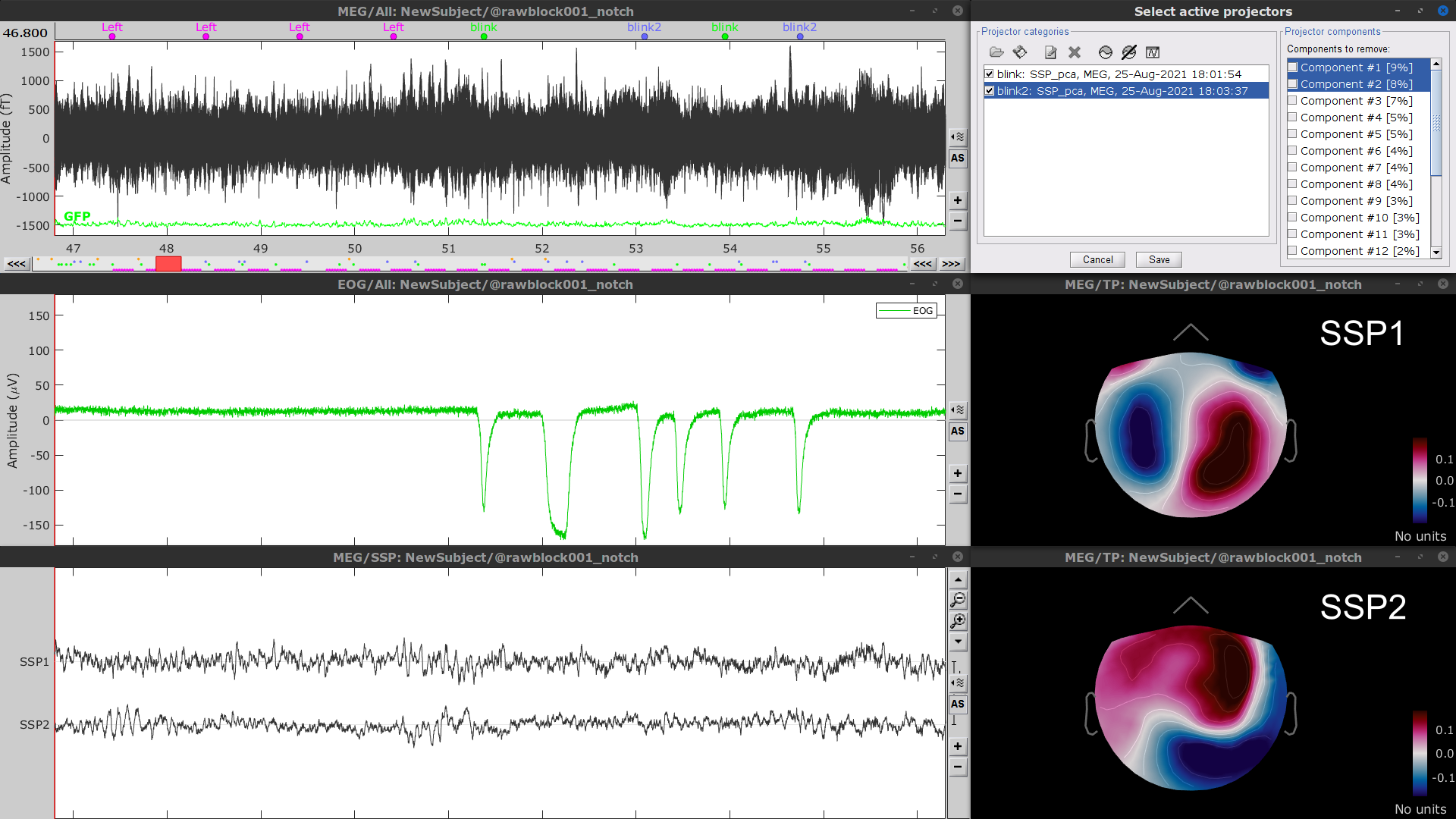
We therefore recommend to unselect the blink2 and blink3 groups from the Select Active Projectors panel (see below) rather than removing spatial components which nature remains ambiguous.
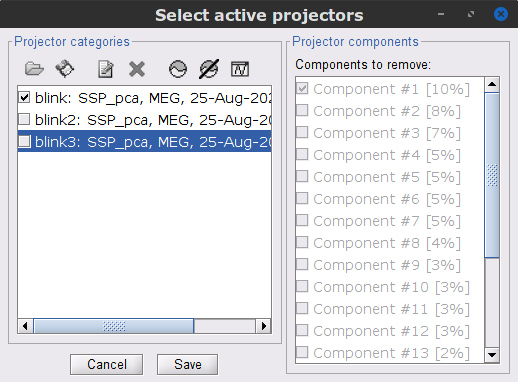
- Click on the large crosshair at the top right of the main Brainstorm window to close all visualization windows.
Detection of "bad" data segments:
Here we will use the automatic detection of artifacts to identify data segments contaminated by e.g., large eye and head movements and muscle contractions.
Display the MEG and EOG time series. In the Record tab, select Artifacts > Detect other artifacts and enter the following parameters:
Time window = 0 - 330 s
Sensor types=MEG
Sensitivity=3
Check both frequency bands 1-7 Hz and 40-240 Hz
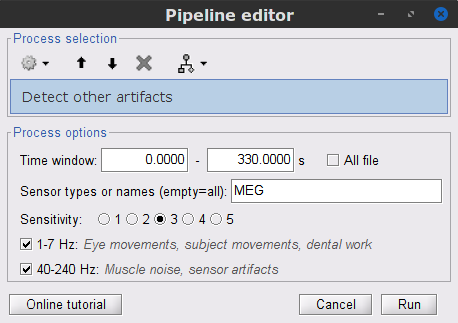
We encourage users to review and validate the segments marked using this procedure. In the present case, the segments detected as bad clearly point at contaminated MEG data segments, which we will now label these as "bad".
Select the 1-7Hz and 40-240Hz event groups and select Events > Mark group as bad from the contextual menu. Alternatively, you can also rename the events created above and append the bad_ prefix to their name: Brainstorm will automatically discard these data segments from further processing.
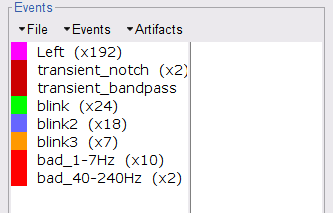
- Close all visualization windows and reply "Yes" to the save the modifications query.
Importing data epochs
At this point we are finished with the pre-processing of the EMG and MEG recordings. We will now extract and import specific data segments of interest into the Brainstorm database for further derivations. We refer to these segments as epochs or trials. As mentioned previously, we will focus on the Left (wrist) category of events.
Right-click on the filtered continuous file Raw | clean | notch(... (in the SubjectCMC_clean_notch_high_abs condition), then Import in database.
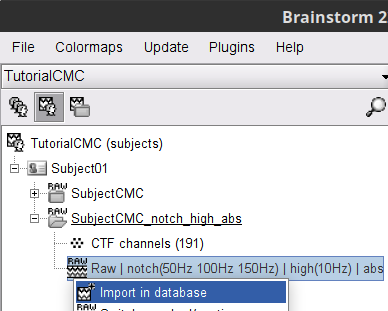
- Enter the following parameter values:
Time window = 0 - 330 s
Check Use events and highlight the Left(x192) event group
Epoch time = 0 - 1000 ms
Check Apply SSP/ICA projectors
Check Remove DC offset and select All recordings
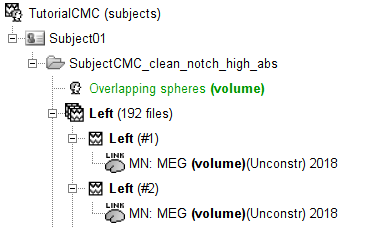
A new folder SubjectCMC_clean_notch_high_abs is created for Subject01. It contains a copy of the channel file from the original raw file, and individual trials tagged as Left in a new trial group. Expand the trial group and note there are trials marked with a question mark in a red circle (ICON). These indicate trials that occurred in the bad segments identified in the previous section. All the bad trials are automatically ignored for further processing, whenever dropped into the Process1 and Process2 tabs.
Coherence estimation (sensor level)
We will now compute the magnitude square coherence (MSC) between the left EMG signal and each of the MEG sensor data.
In the Process1 box, drag and drop the Left (192 files) trial group. Note that the number between square brackets is [185], as the 7 bad trials will be ignored by the MSC process.
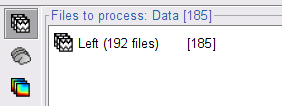
Run the process Connectivity > Coherence 1xN [2021] with the following parameters:
Time window = 0 - 1000 ms or check All file
Source channel = EMGlft
Do not check Include bad channels nor Remove evoke response
Magnitude squared coherence
Window length for PSD estimation = 0.5 s
Overlap for PSD estimation = 50%
Highest frequency of interest = 80 Hz
Average cross-spectra of input files (one output file)
More details on the Coherence process can be found in the ?connectivity tutorial.
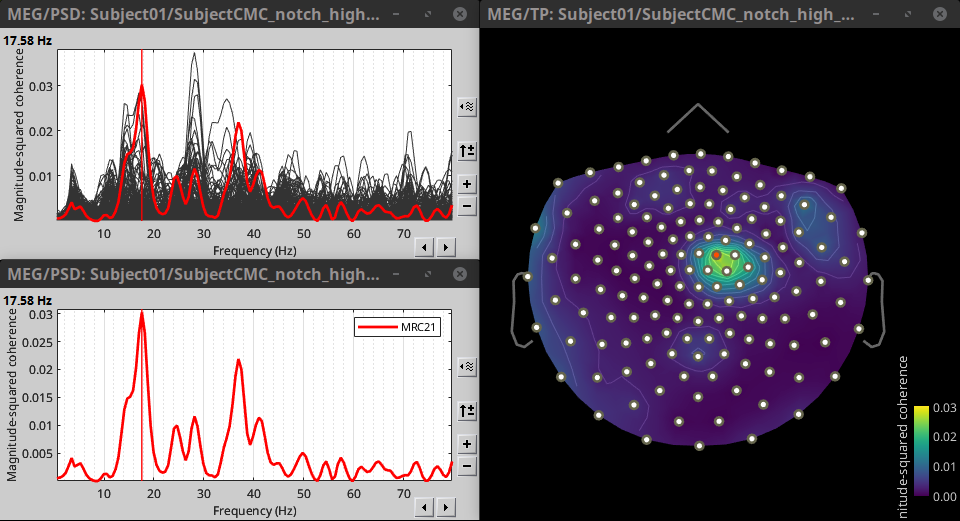
Double-click on the resulting node mscohere(0.6Hz,555win): EMGlft to display the MSC spectra. Click on the maximum peak in the 15 to 20 Hz range, and press Enter to plot it in a new figure. This spectrum corresponds to channel MRC21, and shows a large peak at 17.58 Hz. You can also use the frequency slider (under the Time panel) to explore the MSC output more precisely across frequencies.
* Right-click on the spectrum and select 2D Sensor cap for a topographical representation of the magnitude of the coherence results across the sensor array. You may also use the shortcut Ctrl-T. The sensor locations can be displayed with a right-click and by selecting Channels > Display sensors from the contextual menu (shortcut Ctrl-E).
We can now average magnitude of the MSC across a frequency band of interest (15-20 Hz):
In the Process1 box, drag-and-drop the mscohere(0.6Hz,555win): EMGlft node, and add the process Frequency > Group in time or frequency bands with the parameters:
Select Group by frequency
Type cmc_band / 15, 20 / mean in the text box.
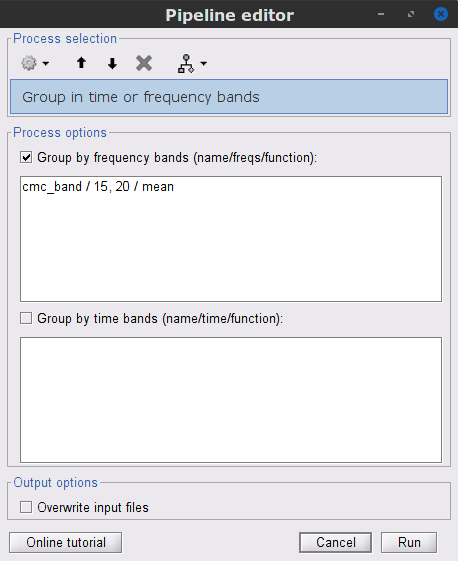
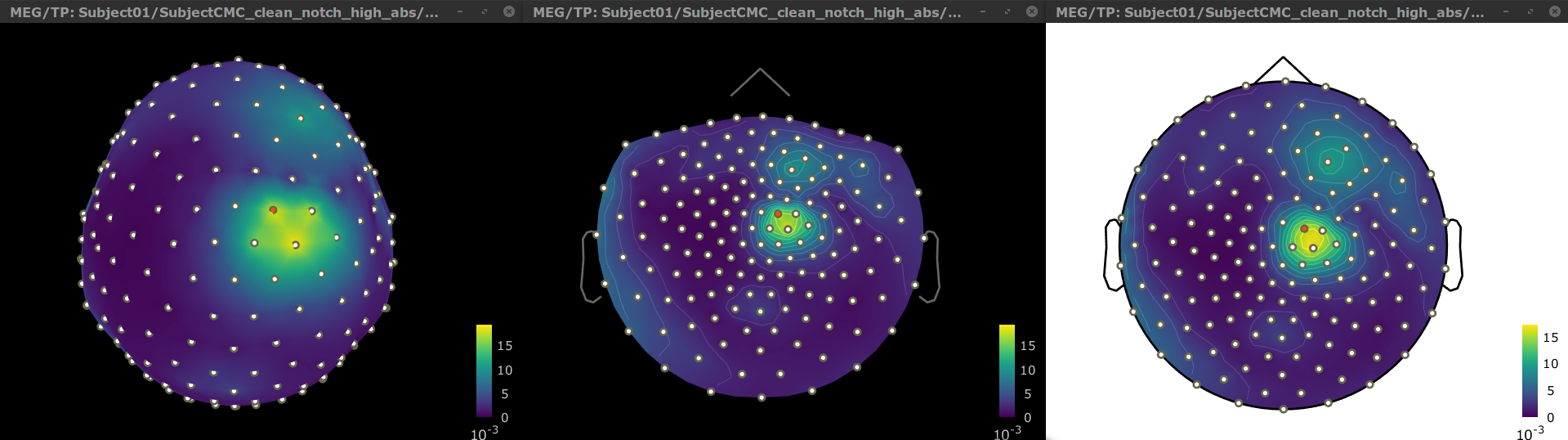
The resulting file mscohere(0.6Hz,555win): EMGlft | tfbands has only one MSC value for each sensor (the MSC average in the 15-20 Hz band). You may visualize the topography of this MSC statistics via 3 possible representations: 2D Sensor cap, 2D Sensor cap and 2D Disk, which are all accessible via a right-click over the MSC node. We clicked on sensor MRC21 below; it is shown in red.
We can observe higher MSC values between the EMG signal and MEG sensor signals over the contralateral set of central sensors in the beta band. In the next sections we will perform source estimation and compute coherence across the entire brain volume.
MEG source imaging
We will perform source modelling using a distributed model approach for two possible source maps:the cortex surface and the entire MRI volume.
SB: I thought we had said above the cortical surface was too crappy for cortical source modelling.
For the cortical source model, the locations of the sources are constrained to the cortical surface obtained from the participants anatomical MRI, as imported above. For MRI violume approach, elementary sources are uniformly distributed across the entire brain volume. Before estimating the brain sources, we need to derive a head model and estimate the noise covariance of the recordings. Note that a distinct head model is required for each source space tested here (cortical vs. volume).
Head modelling
The head model accounts for how neural electrical currents produce magnetic fields captured by sensors outside the head, considering head tissues electromagnetic properties and geometry, independently of actual empirical measurements. Please refer to the head model tutorial for more in-depth explanations.
Cortical surface
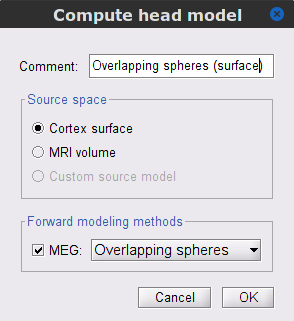
In the SubjectCMC_clean_notch_high_abs node, right-click over CTF channels (191) and select Compute head model from the contextual menu. Run the process with the default options as indicated below:
Comment = Overlapping spheres (surface)
Source space = Cortex surface
Forward model = Overlapping spheres.
The cortical head model will be derived from each of the 10,000 sources (surface vertices) as defined when when the subject anatomy was imported.
The (ICON) Overlapping spheres (surface) head model now appears in the database explorer.
MRI volume source model
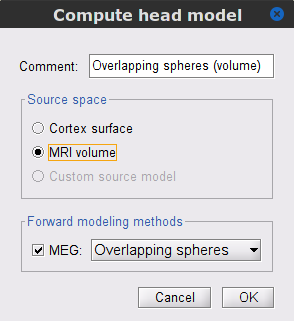
In the SubjectCMC_clean_notch_high_abs node, right-click over the CTF channels (191) node and select Compute head model. Keep all default options:
Comment = Overlapping spheres (volume)
Source space = MRI volume
Forward model = Overlapping spheres.
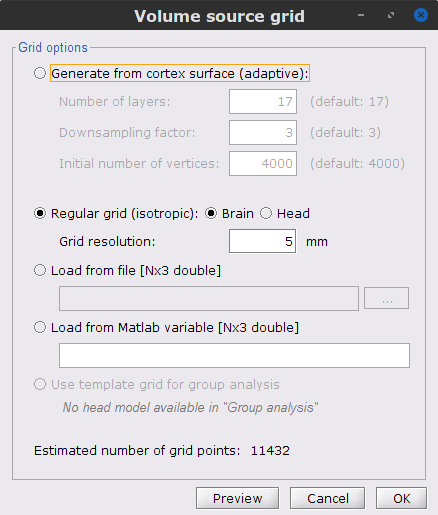
In the Volume source grid window, specify the following parameters that will produce about 12,200 elementary source grid points across the brain volume.
Select Regular grid and Brain
Grid resolution = 5 mm
The Overlapping spheres (volume) node is now added to the database explorer. The green color indicates this is the default head model for the current folder (this can be changed by simply double clicking over the surface node immediately above.)
Noise covariance
Basic noise statistics can be extracted (as recommended) from empty-room recordings. However, these latter are not available from the present tutorial data. However, we will assume that data segments prior to wrist extension can be used as proxies for MEG noise covariance. See the noise covariance tutorial for more details.
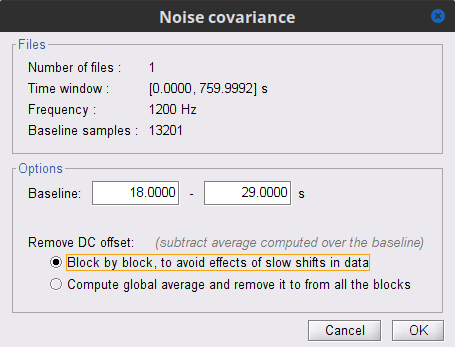
In the raw SubjectCMC_clean_notch_high_abs node, right-click over Raw | clean | notch(...and select Noise covariance > Compute from recordings. Please enter the following parameters:
Baseline: from 18 to 30 s
Select the Block by block option.
Lastly, copy the Noise covariance node to the SubjectCMC_clean_notch_high_abs folder along with the head model. This can be done using the shortcuts Ctrl-C and Ctrl-V.
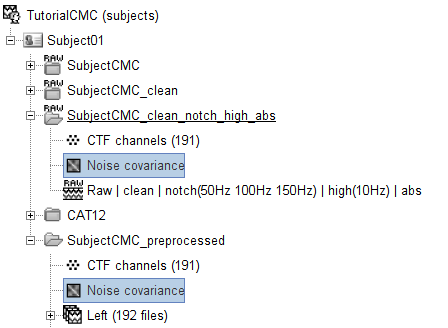
Source estimation
Now that the head model(s) and noise covariance are available, we will use the minimum-norm imaging approach to solve the so-called inverse problem of source estimation. This will produce a large matrix called the imaging kernel, that will be multiplied to the sensor data, to produce estimates of brain source time series. A different imaging kernel is derived for each of the head models we have produced above: surface and volume. See the source estimation tutorial for more details.
Cortical surface
To compute the corresponding imaging kernel, right-click over the Overlapping spheres (surface) head model and select Compute sources [2018]. Enter the following parameters:
Minimum norm imaging
Current density map
Unconstrained
Comment = MN: MEG (surface)
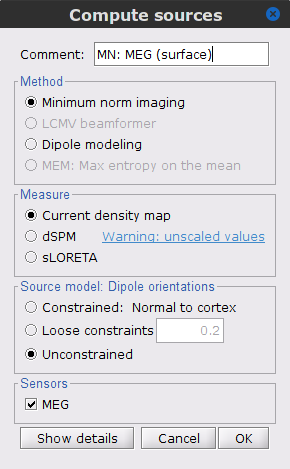
The inversion kernel (ICON) MN: MEG (surface)(Unconstr) 2018 is now available in the database explorer.
MRI volume
Compute the corresponding imaging kernel by right-clicking over the Overlapping spheres (volume) head model and select Compute sources [2018], with the following parameters:
Minimum norm imaging
Current density map
Unconstrained
Comment = MN: MEG (volume)
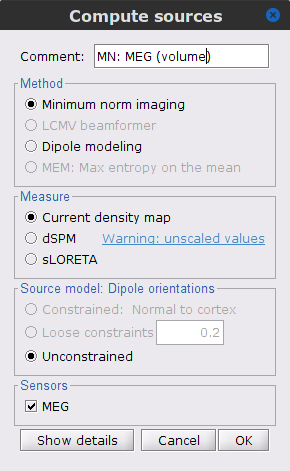
The imaging kernel (ICON) MN: MEG (volume)(Unconstr) 2018 is now also available in the database explorer.
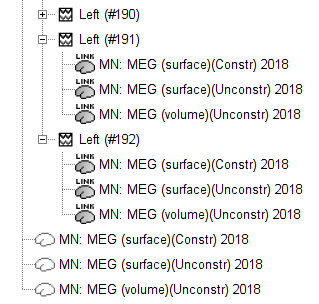
Note that now each trial is associated with two source link (ICON) nodes. One corresponds to the MN: MEG (surface)(Unconstr) 2018 kernel and the other to the MN: MEG (volume)(Unconstr) 2018 kernel.
Defining "scouts" (regions of interest)
To contain computational cost, we will now restrict the computation of coherence with the left EMG signal to selected MEG source activity. We will define regions of interest, whic are known as scouts in Brainstorm. They can be defined as surfaceor volume scouts.
Surface scouts
Richt click over MN: MEG (surface)(Unconstr) 2018 for one of the trials and select Cortical activations > Display on cortex. In the Surface tab, set the Amplitude slider to 100% to hide all the sources.
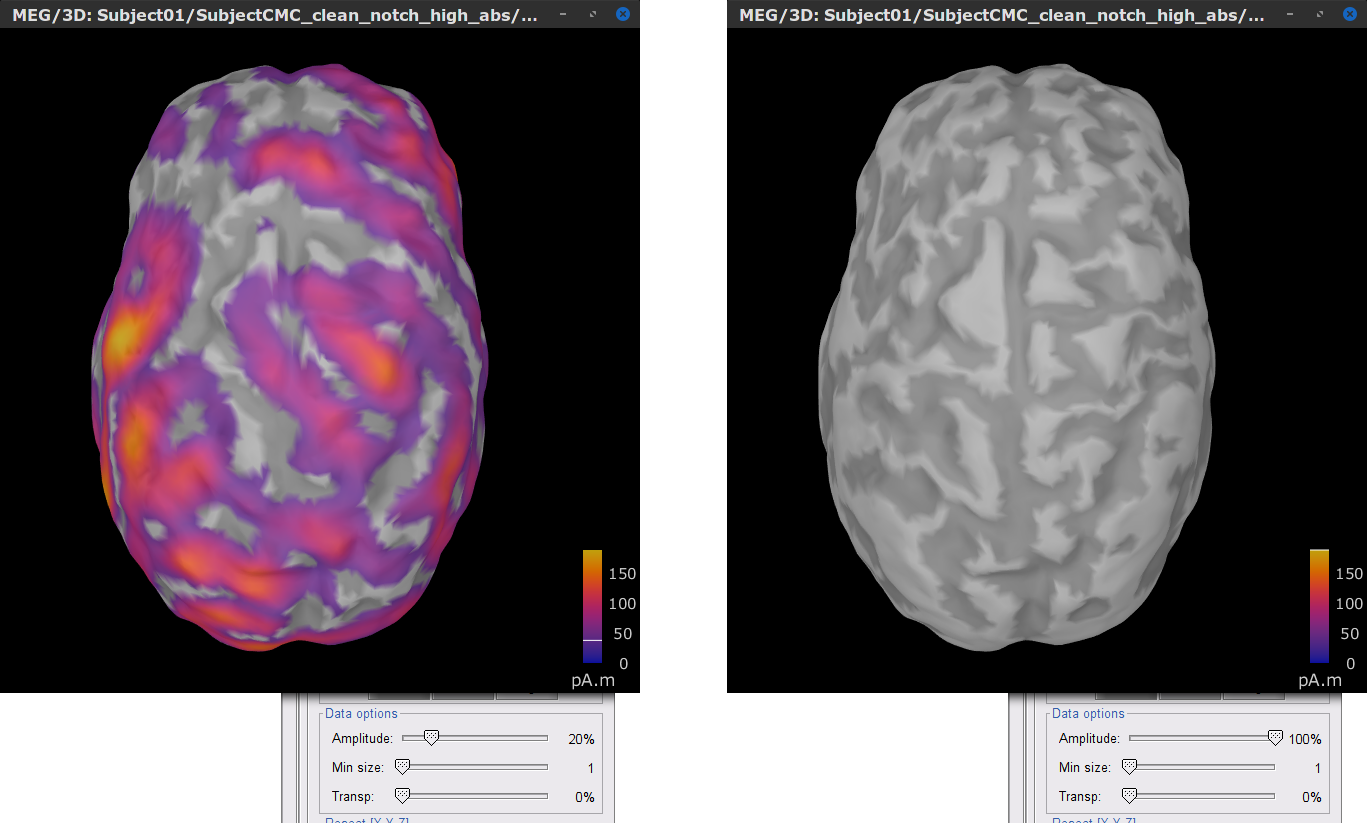
In the Scout tab, select from the menu Atlas > From subject anatomy > AAL3 (MNI-linear). This will create the From volume: AAL3 set of surface scouts. After you click on any scout label shown in the list, Brainstorm will show at the bottom the number of sources (vertices) within the region and an estimate of its surface area in cm2. Activate the (ICON) Show only the selected scouts option to shown only the scout(s) you have selected from the list.
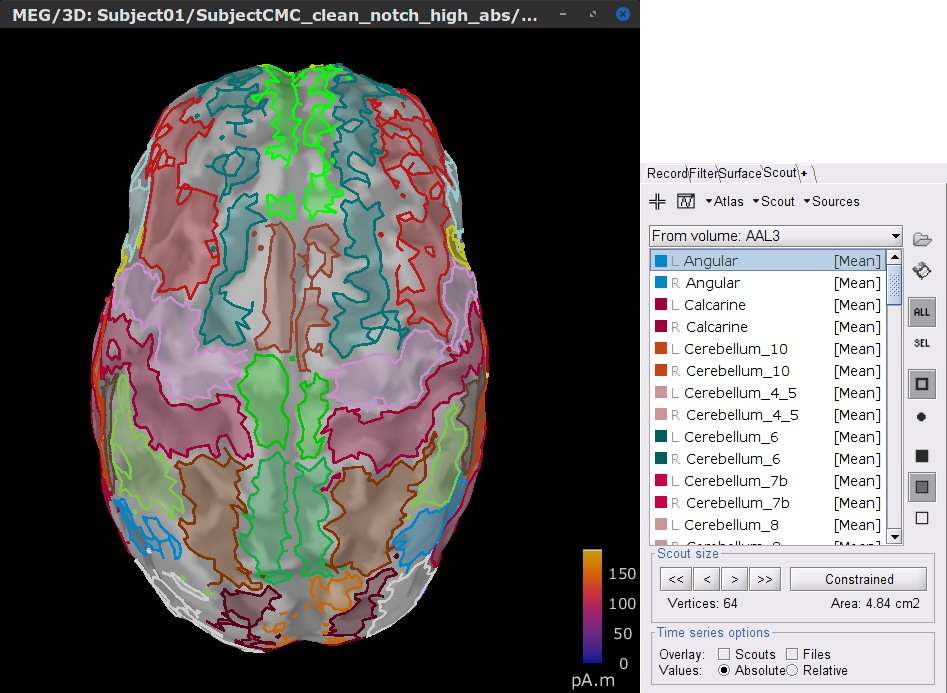
[TODO] A note, that the definition of scouts is far from perfect, but can give us a good idea of the surface projections of the MNI parcellations (described in the importing anatomy section).
- Close the figure.
Volume scouts
Right click over MN: MEG (volume)(Unconstr) 2018 for one of the trials and select Cortical activations > Display on MRI (3D): Subject CMC. In the Surface tab, set the Amplitude slider to 100% to hide all the source map.
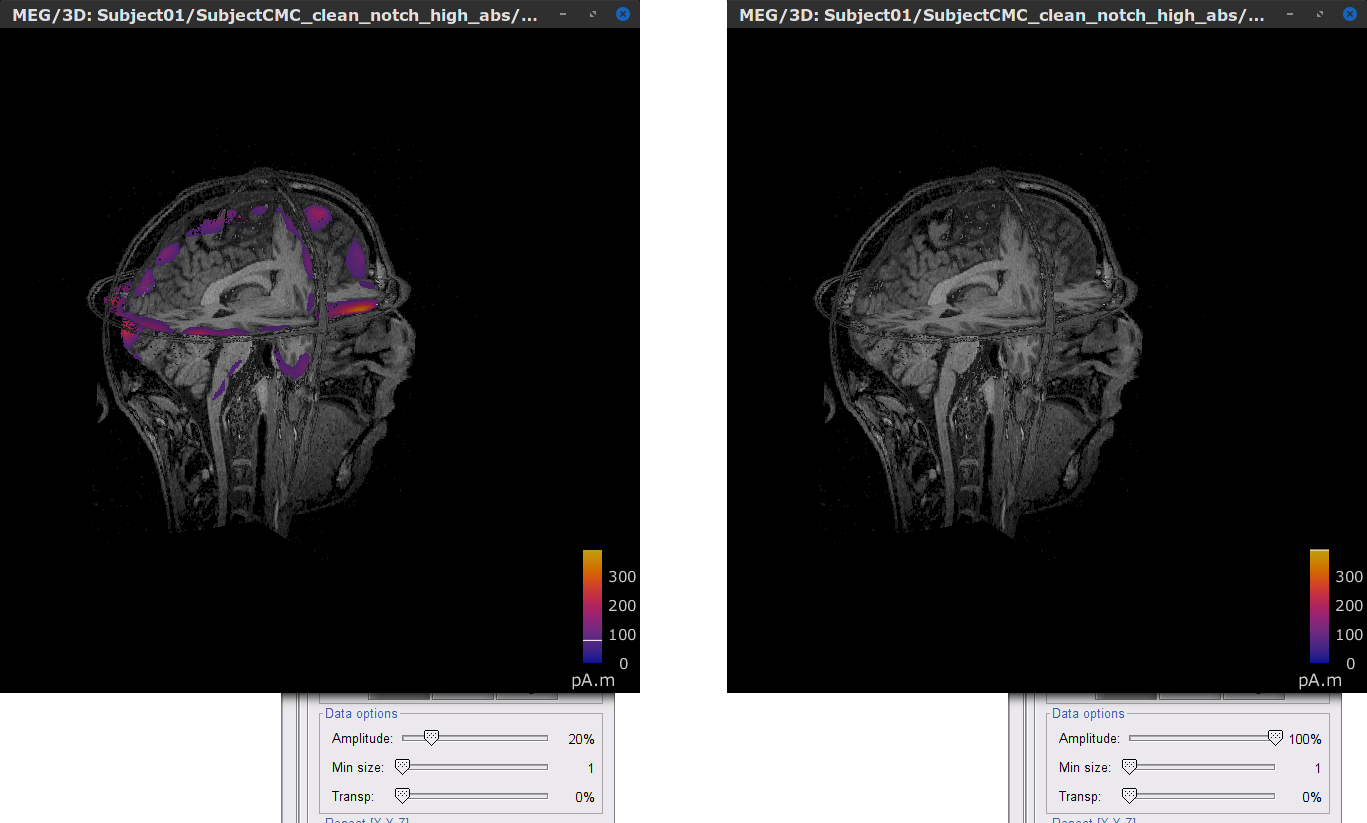
In the Scout tab, select the menu Atlas > From subject anatomy > AAL3 (MNI-linear). This will create a set of Volume 12203: AAL3 volumic scouts. As with the surface scouts above, clicking on a scout in the list will show the number of elementary sources withinand its volume in cm3. Activate the (ICON) Show only the selected scouts option to show only the selected scout(s).
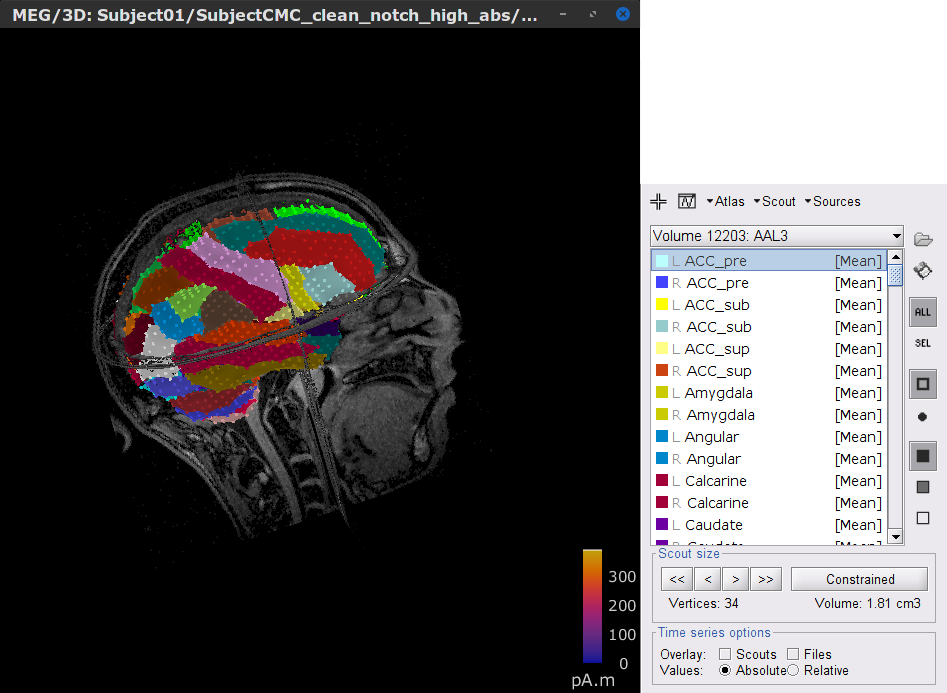
- Close the figure.
Estimating source-wise coherence
Surface scouts
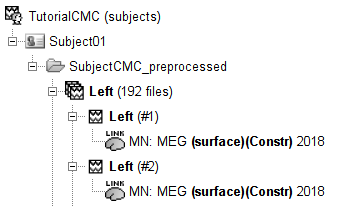
We will use the search features of the Brainstorm database to select the source maps we want to include in the EMG-cortex coherence estimation. We will first select files related to the MN: MEG (surface)(Unconstr) 2018 imaging kernel.
Click on the magnifying glass (ICON) above the database explorer to open the database search panel, and select New search.
- Set the search parameters as shown below:
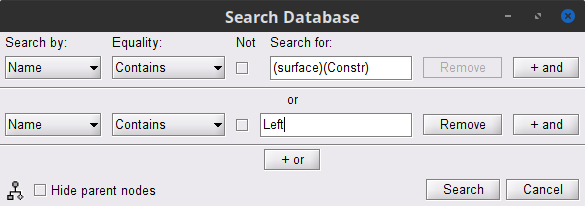
After pressing Search, a new tab called (surface) appears in the database explorer. This new tab contains the data filtered through the search query above.
Click the Process2 tab at the bottom of the main Brainstorm window and drag-and-drop the Left (192 files) trial group into the Files A box and repeat for the Files B box. Select Process recordings for Files A, and Process sources for Files B. The logic is that we will extract from the same files the EMG traces (Files A) and the source time series (Files
 and then compute coherence between these two sets. Note that blue labels over the Files A and the Files B boxes indicate that there are 185 "good trial" files per box.
and then compute coherence between these two sets. Note that blue labels over the Files A and the Files B boxes indicate that there are 185 "good trial" files per box. 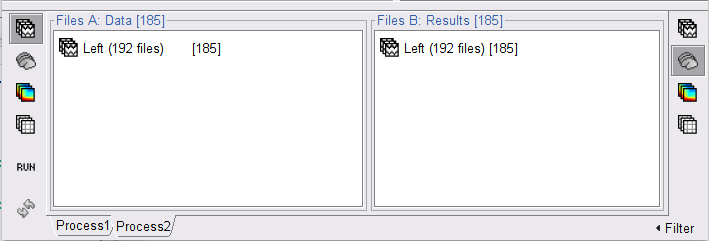
Open the Pipeline editor:
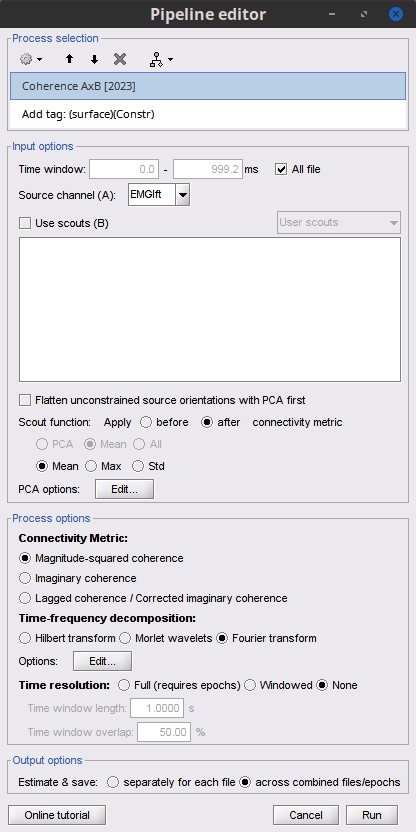
Add the process Connectivity > Coherence AxB [2021] with the following parameters:
Time window = 0 - 1000 ms or check All file
Source channel (A) = EMGlft
Check Use scouts (B)
Select From volume: AAL3 in the drop-down list (these are surface scouts)
Select all the scouts (shortcut Ctrl-A)
Scout function = Mean
When to apply = Before
Do not Remove evoked responses from each trial
Magnitude squared coherence, Window length = 0.5 s
Overlap = 50%
Highest frequency = 80 Hz
Average cross-spectra.
Add the process File > Add tag with the following parameters:
Tag to add = (surface)
Select Add to file name
- Run the pipeline
Double-click on the resulting node mscohere(0.6Hz,555win): Left (#1) | (surface) to display the coherence spectra. Open also the result node as an image with Display image from its contextual menu.
To visualize the location of the scouts on the cortex, double-click one of the (surface) source links for any of the trials. In the Surface tab, set the Amplitude threshold to 100% to hide the cortical activations. Lastly, in the Scouts tab, select the From volume: AAL3 atlas in the drop-down list, select Show only the selected scouts and the Show/hide the scout labels. Note that all visualization windows are linked depending on which scout is selected in the image representation of the coherence results.
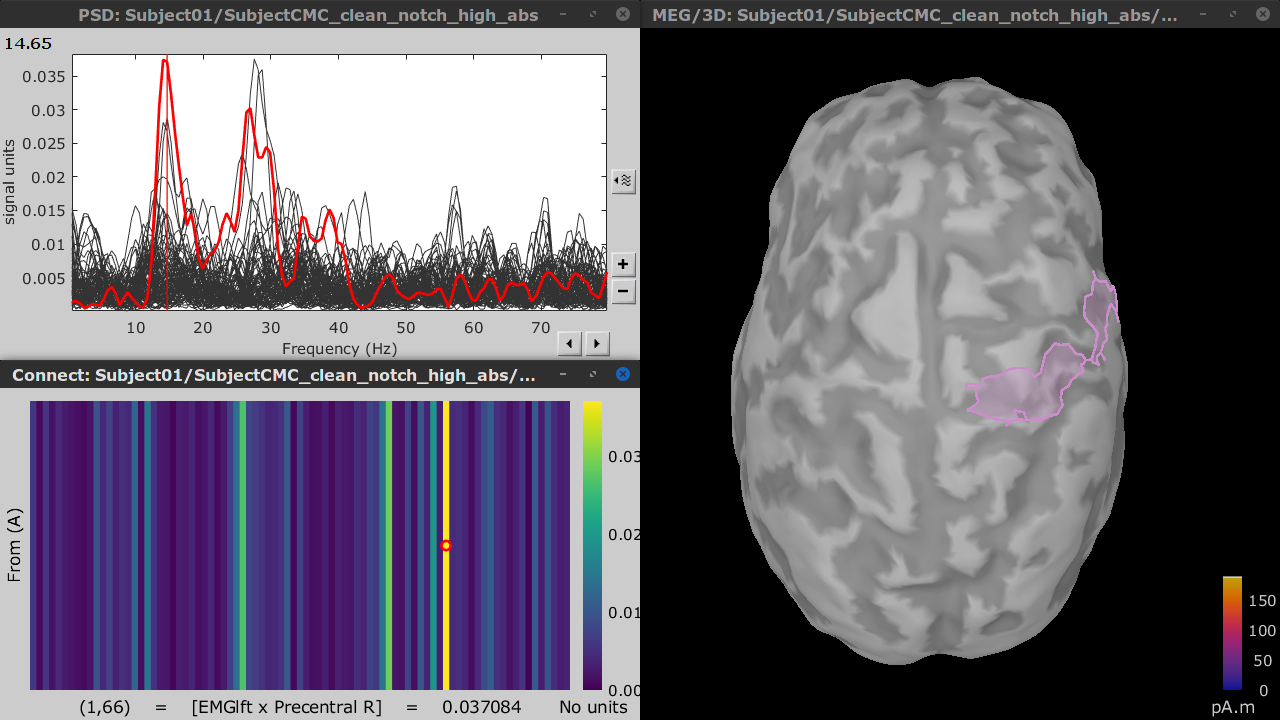
The large peak at 14.65 Hz in the coherence spectra corresponds to the Precentral R scout, which encompasses the right primary motor cortex, as expected.
Volume scouts
We will repeat the same approach Similar for volume scouts. Iniate a file selection query related to the MN: MEG (volume)(Unconstr) 2018 imaging kernel.
- Click on the magnifying glass (ICON) above the database explorer to open the search panel, with the following specifications:
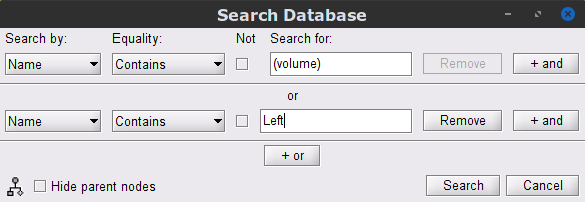
In the Process2 tab, drag and drop the Left (192 files) trial group into the Files A box and repeat for the Files B box. Select the Process recordings icon for Files A, and Process sources for Files B.

Open the Pipeline editor:
Add the process Connectivity > Coherence AxB [2021] with the following parameters:
Time window = 0 - 1000 ms or check All file
Source channel (A) = EMGlft
Check Use scouts (B)
Select Volume 12203: AAL3 in the drop-down list (these are volume scouts)
Select all the scouts (shortcut Ctrl-A)
Scout function = Mean
When to apply = Before
Do not Remove evoked responses from each trial
Magnitude squared coherence, Window length = 0.5 s
Overlap = 50%
Highest frequency = 80 Hz
Average cross-spectra.
Add the process File > Add tag with the following parameters:
Tag to add = (surface)
Select Add to file name
- Run the pipeline
Double-click on the resulting node mscohere(0.6Hz,555win): Left (#1) | (volume) to display the coherence spectra. Open also the result node by selecting Display as image from its contextual menu.
Open one of the (volume) source links for any of the trials by selecting from the contextual menu Display cortical activations > Display on MRI (3D): SubjectCMC . In the Surface tab, set the Amplitude threshold to 100% to hide all the cortical activations. Lastly, in the Scouts tab, select the Volume 12203: AAL3 atlas in the drop-down list, select the Show only the selected scouts and the Show/hide the scout labels.
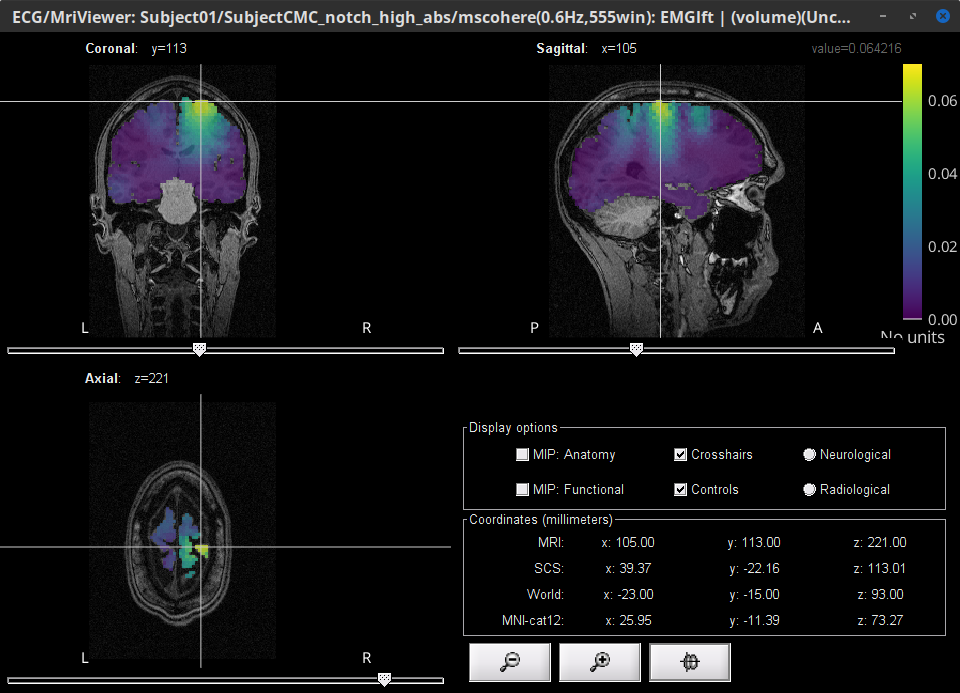
Here too, the coherence peak at 14.65 Hz corresponds to the Precentral R scout, which encompasses the right primary motor cortex, as expected.
Brain-wise coherence estimation (no scouts)
- We could downsample the surface and create a more sparse volume grid
- OR
- Refactor the coherence process to accumulate the auto- and cross-spectra outside of the function
- OR
Comparison of cortical surface with FieldTrip and CAT
[TO DISCUSS among authors] This image and GIF are just for reference. They were obtained with all the surface sources using ?FieldTrip and CAT derived surfaces.
Comparison for 14.65 Hz
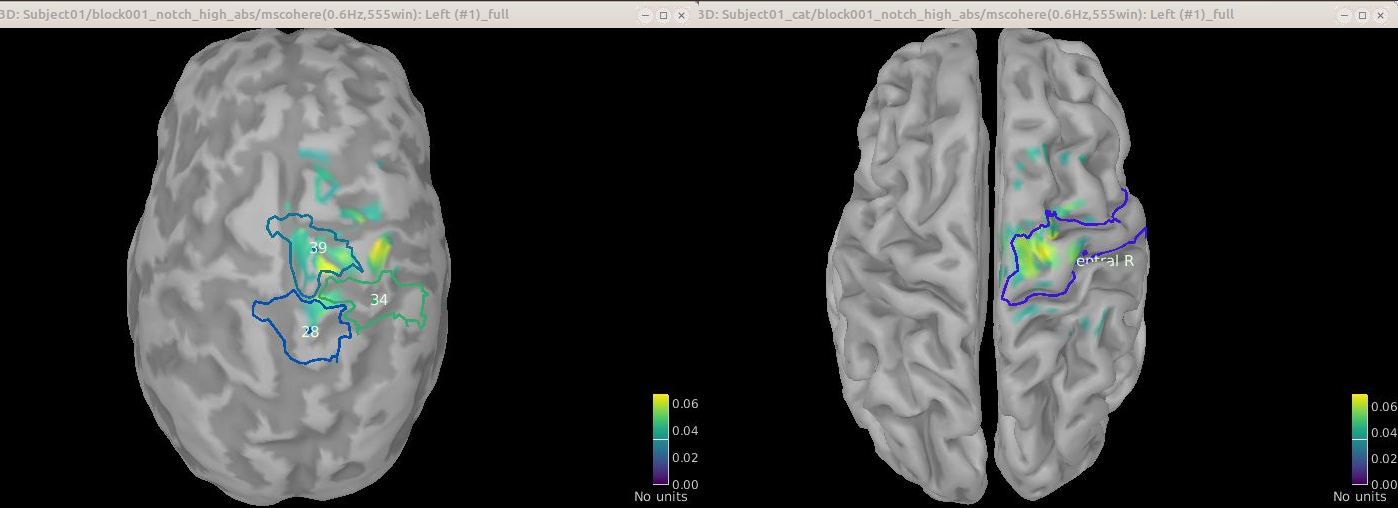
Sweeping from 0 to 80 Hz
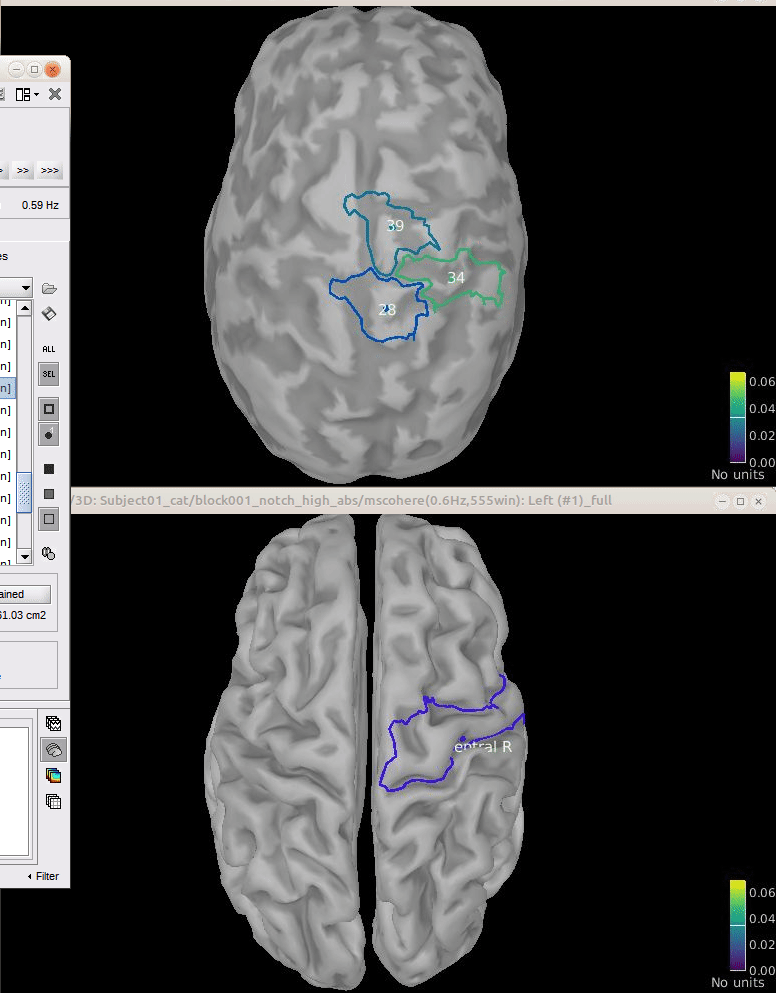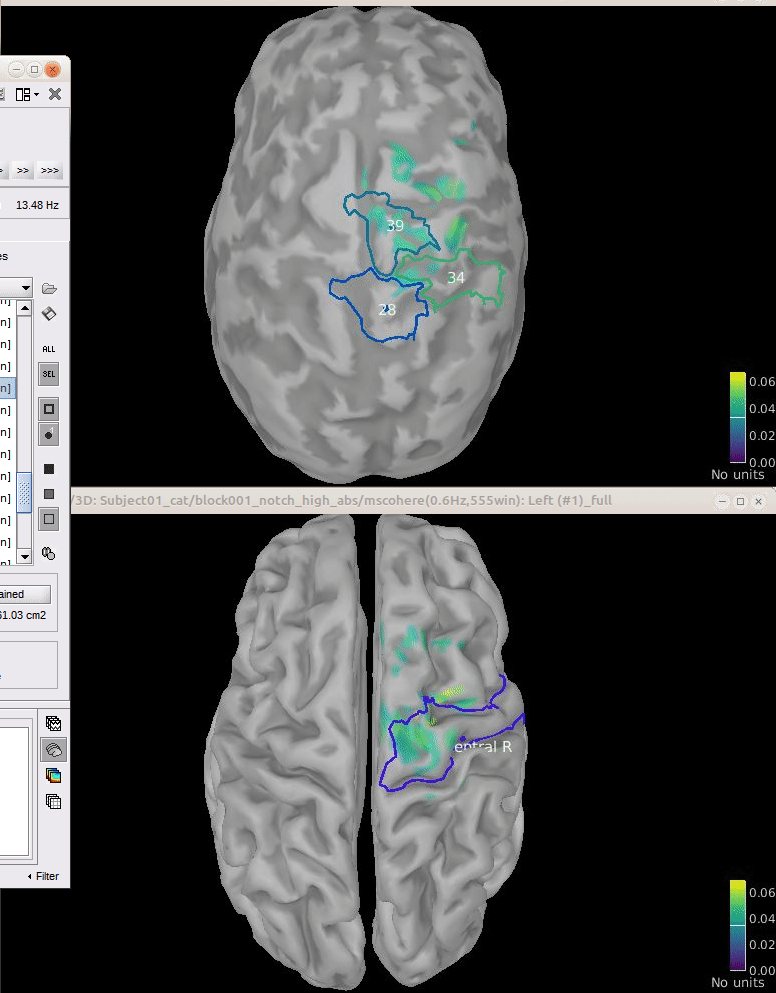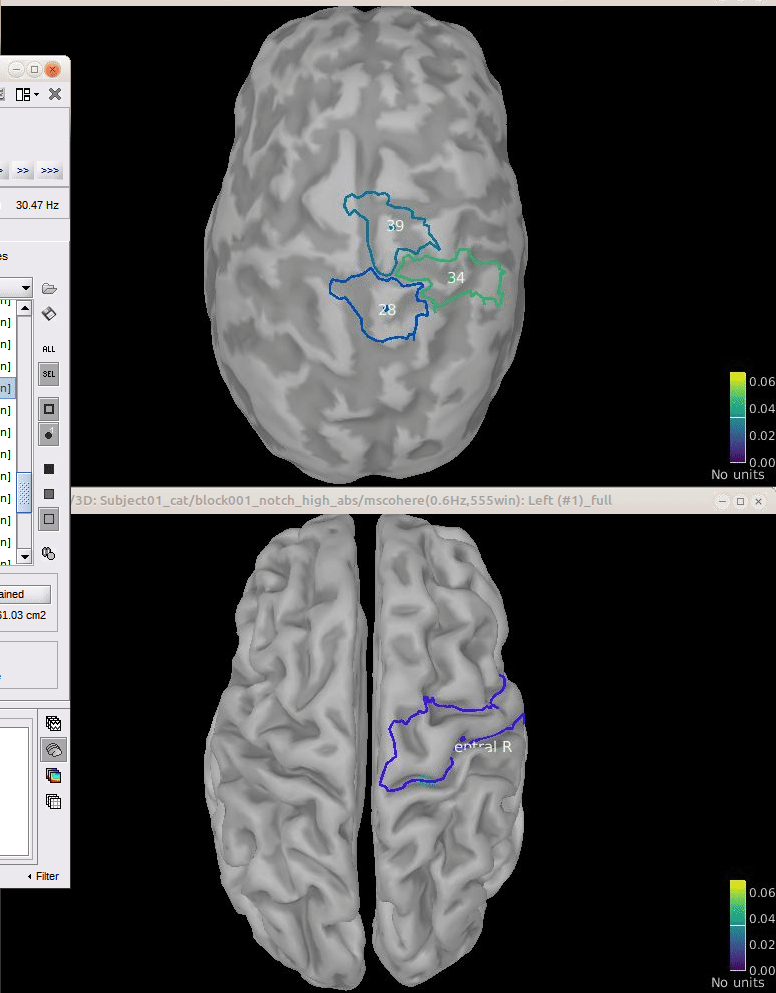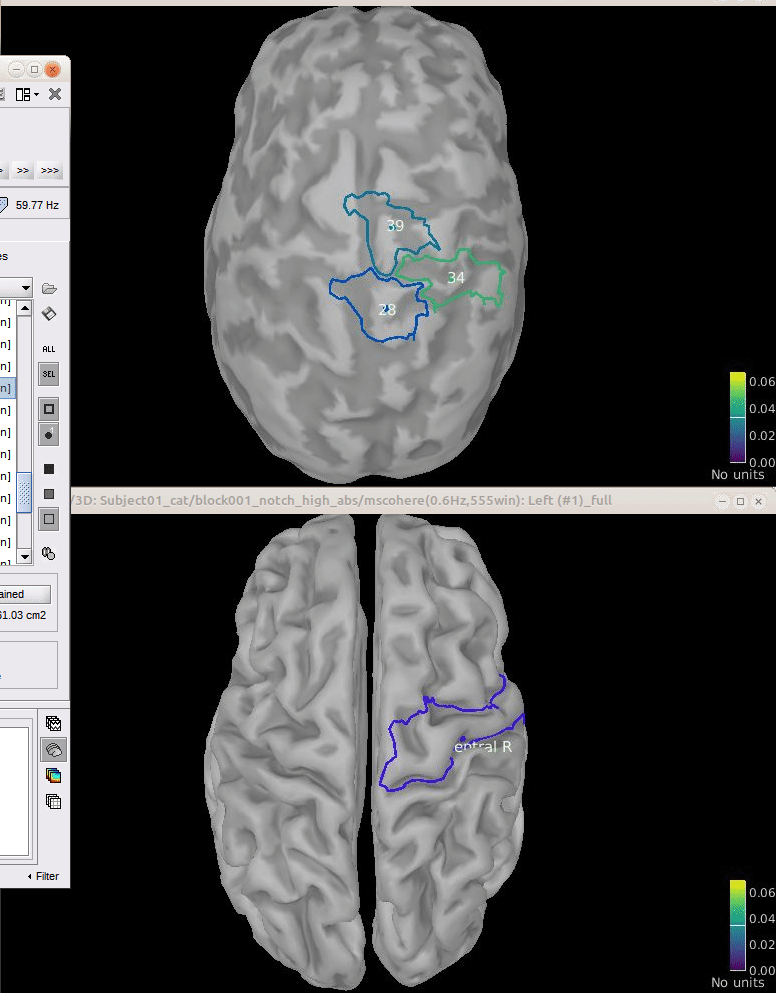
Script
[TO DO] Once we agree on all the steps above.
Additional documentation
Articles
Conway BA, Halliday DM, Farmer SF, Shahani U, Maas P, Weir AI, et al.
Synchronization between motor cortex and spinal motoneuronal pool during the performance of a maintained motor task in man.
The Journal of Physiology. 1995 Dec 15;489(3):917–24.Kilner JM, Baker SN, Salenius S, Hari R, Lemon RN.
Human Cortical Muscle Coherence Is Directly Related to Specific Motor Parameters.
J Neurosci. 2000 Dec 1;20(23):8838–45.Liu J, Sheng Y, Liu H.
Corticomuscular Coherence and Its Applications: A Review.
Front Hum Neurosci. 2019 Mar 20;13:100.
Tutorials
Tutorial: Volume source estimation
Tutorial: Functional connectivity
Forum discussions
[TO DO] Find relevant Forum posts.
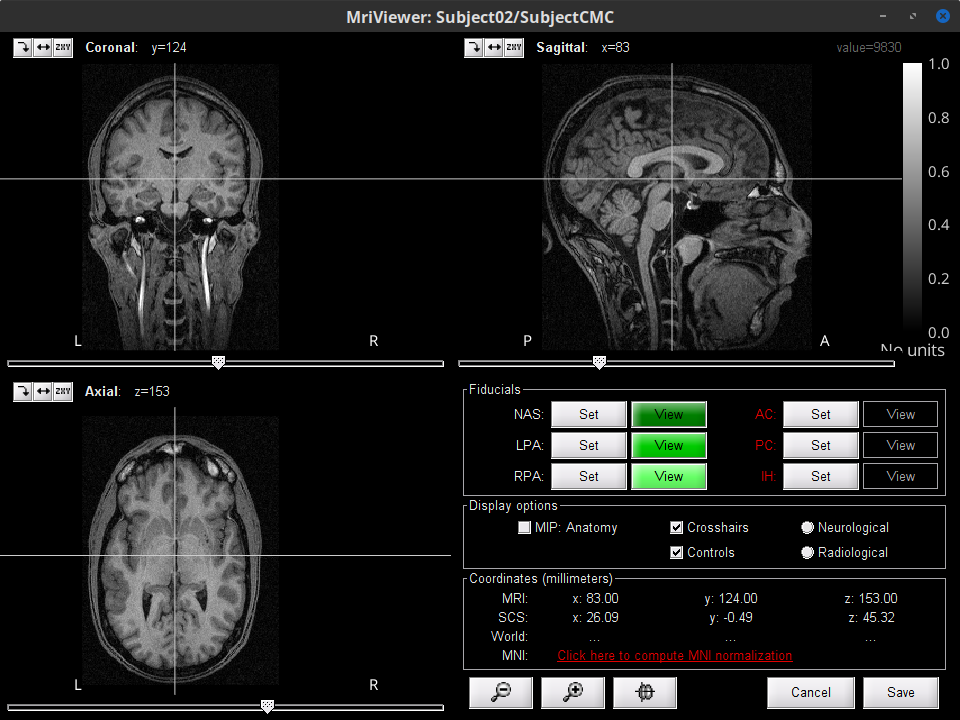
Register the individual anatomy to MNI brain space, for standardization of coordinates: in the MRI viewer click on Click here to compute MNI normalization, use the maff8 method. When the normalization is complete, verify that the locations of the anatomical fiducials are adequate (essentially that they are indeed near the left/right ears and right above the nose) and click on Save.