Hello Brainstorm experts
I am attempting source reconstruction with EEG data. I am in the stage to compute the head model right now. I would like to confirm my concerns during the past steps.
My steps from the very beginning:
1: Import BISD dataset
2: Convert DWI to DTI
3: Compute mesh with SimNIBS
4: Align EEG electrode
5: Compute head model
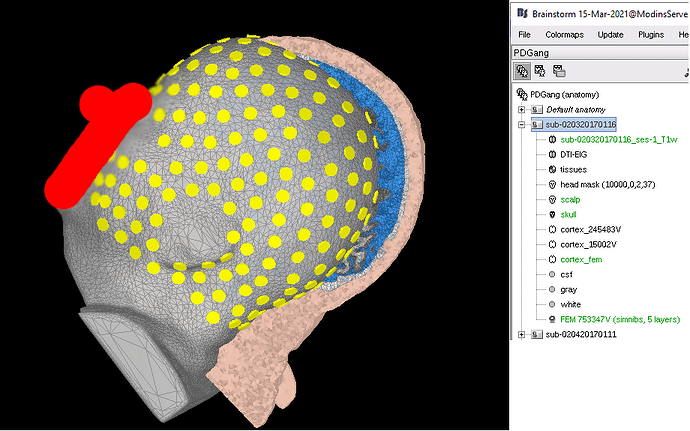
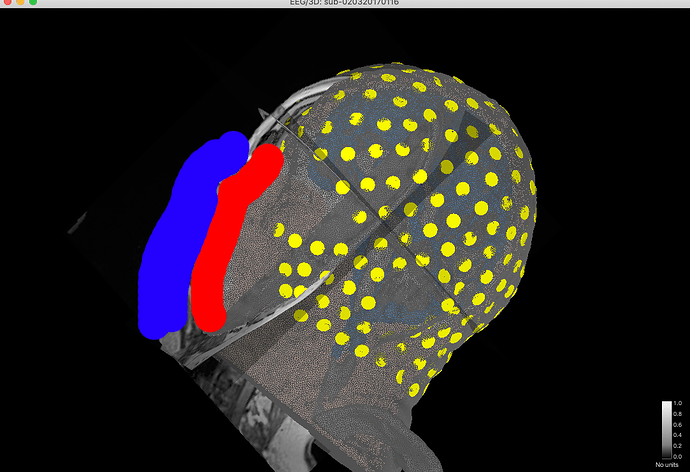
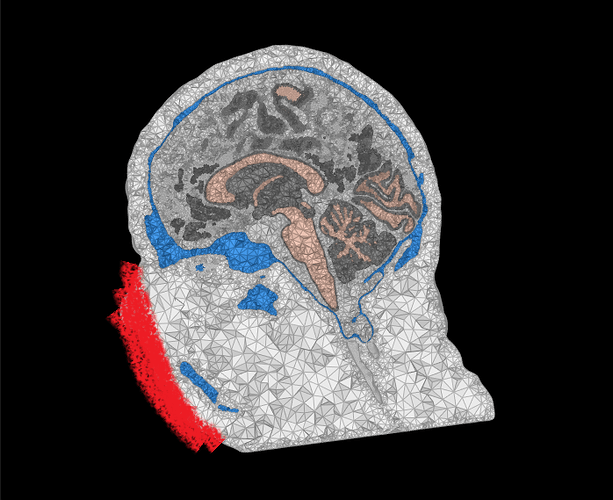
The first issue I observed is that the different space between T1 and FEM mesh. It was noticed when I was trying to align my EEG electrode in Step 4 as shown in the screenshot below. If I understand correctly once I import from the BIDS dataset, there is automated preprocessing for the T1 image and generate head mask and other tissue. Since SimNIBS will do a better job with CAT12 and SPM segmentation, I directly generate the mesh with that. The result of SimNIBS is a 5 layers mesh in the space that is not consistent with the previously generated head mask, so what I did is to extract all the 5 layers first and activate the head mask from the 5 layers for my EEG alignment. I wonder is a correct way of doing it or it is wrong?
The second concern is about the group analysis, I wanted to extract the time series estimated from a specific brain region after the inverse solution. I check the tutorial online, Volume atlases. It seems the atlas can be transformed to the subject space, does that mean I can do this for each subject and then analyze the time series for my group analysis instead of creating a group grid? If I create a group grid, does my FEM mesh(different space issue in Question 1) will be automated coregistered with the template correctly?
The last issue I am considering is the channel number of the head model. Since the head model can be generated independent from EEG recordings, I couldn't tell whether the EEG recording aftering clearning will still keep the same channels. For example, I make the head model based on 256 channels, but cleaned EEG recording may have just 230 channels, would it still work with the head model when estimating the source?
Thank you very much for your answer, if anything is not clear don't hesitate to request. Thanks.