|
Size: 23550
Comment:
|
Size: 19501
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 7: | Line 7: |
| * Default montages for EEG (sensor selection) | |
| Line 10: | Line 11: |
| * Add filter option to the montage editor: This would allow splitting the signals in various frequency bands. | |
| Line 13: | Line 13: |
| * RAW file viewer: | * RAW file viewer speed: |
| Line 18: | Line 18: |
| * Bad trials: When changing the status of bad to good: remove the bad segments as well, otherwise it is not processed by processes like the PSD. * Review clinical recordings: Reduce the dimensionality of the data with a simple inverse problem, similar to what we do for the magnetic extrapolation ("Regional sources" in BESA, cf S Rampp) |
* Review signals in time-frequency space |
| Line 22: | Line 21: |
| * Import window: Add button to create the corresponding processing pipeline (to generate script or to edit additional options) * BioSemi: Add menu "Convert naming system" to rename channels into 10-10 (A1=>FPz) * Simulations: https://github.com/lrkrol/SEREEGA * Integrate with EYE-EEG (Olaf Dimigen) * Reproduce tutorial: https://www.eyetracking-eeg.org/tutorial.html * Add note when rejecting trials: https://neuroimage.usc.edu/forums/t/33686 |
|
| Line 25: | Line 30: |
| Line 31: | Line 37: |
| * Smooth display from figure_image (ERPimage, raster plot...) | |
| Line 35: | Line 40: |
| * Use boundary() instead of conhull() in all the display functions (ie. 2DDisc) | |
| Line 42: | Line 46: |
| * Thresholding and stat tests the connectivity matrices * Connectivity on unconstrained sources: "Default signal extraction for volume grids should be the time series of the first principal component of the triplet signals after each has been zero-meaned" (SB) * Display of connectivity graphs: * Display as straight lines * Recode 2D graphs * 3D display with anatomical constrains * Display using real position of EEG electrodes * Use new band-pass filters in bst_connectivity ('bst-hfilter' instead of 'bst-fft-fir') * Matrix view of NxN graphs: Add legend of the elements along X and Y axis * Graph view: * Does not display negative values correctly (correlation or difference of coherence) * Re-write using pure Matlab code and smoothed graphics * Fixed scales for intensity sliders * Text bigger * Too much data in appdata * Fixed scales for intensity sliders * Add "=" shortcut for having graphs with similar configurations * Disable zoom in one region (serious bugs) * NxN on sensors: does not place the sensors correctly in space * Coherence: * Average cross-spectra instead of concatenating epochs (to avoid discontinuities)<<BR>>Explore inter-trial approaches (Esther refers to chronux toolbox) |
* Thresholding and stat tests for connectivity matrices * Connect NxN display: * Graph on sensors: does not place the sensors correctly in space * Display as image: Add legend of the elements along X and Y axis * Display as time series: Display warning before trying to open too many signals * Optimize display: use surface() instead of line() for links? (as in figure_3d/PlotFibers) * Time-resolved correlation/coherence: Display as time bands * Weighted Phase Lag Index (WPLI) |
| Line 64: | Line 55: |
| Line 65: | Line 57: |
| * Add p-values | |
| Line 67: | Line 58: |
| * Optimize code | |
| Line 71: | Line 61: |
| * Time-resolved correlation/coherence: Display as time bands * Tutorial coherence [1xN] : Reproduce FieldTrip results? * Connect NxN: Display as time series > Display warning before trying to open too many signals |
|
| Line 76: | Line 63: |
| * Decoding/Classifiers: Implement Dimitrios and scikit-learn algorithms * Allow processes in Python and Java |
* Generate fully reproducible scripts, including all the interactive/graphical parts: * Saving all the interactive operations as process calls * Improving the pipeline editor to handle loops over data files or subjects * Keeping a better track of the provenance of all the data (History, uniform file names) |
| Line 80: | Line 69: |
| * Implement data exchange with MNE-Python: write FIF files from Brainstorm and/or pass python objects in memory instead of FIF files | * https://neuroimage.usc.edu/forums/t/ica-on-very-long-eeg/23556/4 |
| Line 82: | Line 71: |
| * SSS/tSSS cleaning | |
| Line 84: | Line 72: |
| * Point-spread functions (PSFs) and cross-talk functions: https://mne.tools/stable/auto_examples/inverse/plot_psf_ctf_vertices.html#sphx-glr-auto-examples-inverse-plot-psf-ctf-vertices-py * Spatial resolution metrics in source space:<<BR>>https://mne.tools/stable/auto_examples/inverse/plot_resolution_metrics.html#sphx-glr-auto-examples-inverse-plot-resolution-metrics-py |
|
| Line 85: | Line 75: |
| * Chronux toolbox : http://chronux.org/ | |
| Line 95: | Line 86: |
| * ft_prepare_sourcemodel: Compute MNI transformation (linear and non-linear) => Peter | |
| Line 98: | Line 88: |
| * ft_read_atlas('TTatlas+tlrc.BRICK'); | |
| Line 103: | Line 92: |
| * Test speed for writing files: <<BR>>https://undocumentedmatlab.com/articles/improving-fwrite-performance | |
| Line 109: | Line 99: |
| * When computing sources from the pipeline editor: doesn't reselect the options if you click twice on "edit" (works for minnorm, but not for lcmv) | |
| Line 113: | Line 102: |
| * Add Alex's suggestions: https://neuroimage.usc.edu/forums/t/ica-on-very-long-eeg/23556/4 * Automatic classification: ICLabel: https://neuroimage.usc.edu/forums/t/automatic-eeg-ic-ica-classification-for-brainstorm/33785 * Add methods: SOBI, Fastica, AMICA/CUDICA/CUDAAMICA (recommended by S Makeig) |
|
| Line 118: | Line 110: |
| * Use faster methods (MNE-Python?) * Add methods: SOBI, Fastica, AMICA/CUDICA (recommended by S Makeig) |
|
| Line 121: | Line 111: |
| * S Makeig: Use ICA to select the IC of interest instead of only removing artifacts * Display of spectrum for components (PSD/FFT) |
|
| Line 127: | Line 115: |
| * Save IC time series in database | |
| Line 129: | Line 116: |
| * Other EEGLAB functions: * Step function detection: https://github.com/lucklab/erplab/wiki/Artifact-Detection:-Tutorial |
|
| Line 133: | Line 118: |
| * Spectral flattening (John): * ARIMA(5,0,1): Apply on the signal before any frequency/connectivity/PAC analysis * PSD: * Rewrite to have the same input as coherence (frequency resolution instead of window length) * Allow display of Avg+StdErr |
|
| Line 144: | Line 122: |
| * Matching pursuit: http://m.jneurosci.org/content/36/12/3399.abstract?etoc * Bug: Display logs as negative * Bug: 3D figures: Colormaps with "log" option doesn't work * Bug: Difference of power displayed in log: problems (Soheila) * 2D Layout in spectrum * Make much faster and more memory efficient (C functions coded by Matti ?) * TF scouts: should display average of TF maps * Impossible to keep complex values for unconstrained sources * Pad short epochs with zero values for getting lower frequencies |
|
| Line 154: | Line 123: |
| * Extend clusters tab to display of TF to overlay TF signals (Svet) | |
| Line 156: | Line 124: |
| * Allow baseline normalization of files computed with time bands | |
| Line 160: | Line 127: |
| * Artifact detection: * Artifact rejection like SPM: if bad in 20%, bad everywhere * Test difference between adjacent samples * Events detection: Add option "std" vs "amplitude" * Simulation: * EEGSourceSim: https://www.sciencedirect.com/science/article/pii/S0165027019302341 * Fix units in simulation processes => no *1e-9 in "simulate recordings" * Use "add noise" process from Hui-Ling (in Work/Dev/Divers) * Use field process field "Group" to separate Input/Processing/Output options * Use new Matlab functions: movmean, movsum, movmedian, movmax, movmin, movvar, movstd == Database == * Rename protocol * Faster DB searches (for Emily) * Add buttons to sort files: by name, by comment, by date * MEG protocols: More flexible organization of the database; sub-conditions to allow different runs X different conditions. * Matrix files: Allow to be dependent from other files * Rename multiple files * Default headmodel lost when reloaded: Keep selection on the hard drive (in brainstormstudy.mat) * Auto-save: * protocol.mat can be too big: do not store the results links in it (and recreate when loading)- http://neuroimage.usc.edu/forums/t/abnormally-slow-behavior/2065/10 * Improve auto-save: add tracking file next to protocol.mat, do not save all the time, only when closing app, and reload protocol at stratup if tracking file is still there == Distributed computing == * Options from FieldTrip: * Loose collection of computers: https://github.com/fieldtrip/fieldtrip/tree/master/peer * Alternative, with less limitations: http://research.cs.wisc.edu/htcondor/ * Single multicore machine: https://github.com/fieldtrip/fieldtrip/tree/master/engine * Batch system: https://github.com/fieldtrip/fieldtrip/tree/master/qsub * Documentation: http://fieldtrip.fcdonders.nl/faq#distributed_computing_with_fieldtrip_and_matlab * PSOM: http://psom.simexp-lab.org/ * Various initiatives: http://samirdas.github.io/Data_sharing.html#/ |
|
| Line 198: | Line 129: |
| * Use eLORETA instead of sLORETA? <<BR>>https://neuroimage.usc.edu/forums/t/compute-eeg-sources-with-sloreta/13425/6 | * Reproduce results in "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x * eLORETA instead of sLORETA? * https://neuroimage.usc.edu/forums/t/compute-eeg-sources-with-sloreta/13425/6 * "eLORETA algorithm is available in the MEG/EEG Toolbox of Hamburg (METH)": https://www.biorxiv.org/content/biorxiv/early/2019/10/17/809285.full.pdf * https://github.com/brainstorm-tools/brainstorm3/issues/114 * Sensitivity maps: https://mne.tools/stable/auto_examples/forward/plot_forward_sensitivity_maps.html |
| Line 203: | Line 139: |
| * Display dipoles in MRI viewer | |
| Line 205: | Line 142: |
| * Project sources: Very poor algorithm to project sub-cortical regions and cerebellum (algorithm to fit surfaces should be imrpoved) * Menu head model > Copy to other conditions/subjects (check if applicable first) * Menu Sources > Maximum value: Doesn't work with volume or mixed head models * Mixed head models: * Set loose parameter from the interface |
* Project sources: Very poor algorithm to project sub-cortical regions and cerebellum * Maximum: * Menu Sources > Maximum value: Doesn't work with volume or mixed head models * Panel Get coordinates: Add button "find maximum" * Sources on surface: Display peak regions over time (time = color) => A.Gramfort * BEM single sphere: Get implementation from MNE |
| Line 213: | Line 151: |
| * Menu Sources > Simulate recordings: * Do not close the 3D figures after generating a new file * Add a process equivalent to this menu * Panel Get coordinates: Add button "find maximum" * BEM single sphere: Get implementation from MNE * Unconstrained sources: * Stat and connectivity: what to do? (re-send email John+Sylvain) * Sources on surface: Display peak regions over time (time = color) => A.Gramfort * Process "Extract scouts time series": Add PCA option (replace isnorm with choice PCA/Norm) |
|
| Line 223: | Line 152: |
| * Display source maps on a flat 2D cortex projection (Mollweide projection): https://neuroimage.usc.edu/forums/t/source-model-display-and-output/13940/5 | * Display spectrum scouts (PSD plots when clicking on "Display scouts" on PSD/full cortex) |
| Line 226: | Line 155: |
| * Multi-Scale Brain Parcellator (Lausanne2008): * [[https://github.com/sebastientourbier/multiscalebrainparcellatorhttps://hub.docker.com/r/sebastientourbier/multiscalebrainparcellator|https://github.com/sebastientourbier/multiscalebrainparcellator]] * [[https://github.com/sebastientourbier/multiscalebrainparcellatorhttps://hub.docker.com/r/sebastientourbier/multiscalebrainparcellator|https://hub.docker.com/r/sebastientourbier/multiscalebrainparcellator]] * https://multiscalebrainparcellator.readthedocs.io/en/latest/ * MNI transformation: Use SPM non-linear MNI transformation y_... * Registration: * Getting electrode positions from 3D scanners: https://sccn.ucsd.edu/wiki/Get_chanlocs * GARDEL: http://meg.univ-amu.fr/wiki/GARDEL:presentation * Use the same registration for multiple recording sessions that have already re-registered previously (eg. with MaxFilter) * When linking multiple EEG recordings including 3D positions, do the registration only once and copy it to all the runs * Compute non-linear MNI registration instead of linear * Select and remove bad digitized head points before automatic coregistration * Load the MNE -transf.fif: http://neuroimage.usc.edu/forums/showthread.php?2830 |
* BEM surfaces: Deform fieldtrip BEM surfaces from ICBM152 to subject space with MNI coordinates? * Simple-brain-plot: https://github.com/dutchconnectomelab/Simple-Brain-Plot * MRI segmentation: <<BR>> * SimNIBS: Replace HEADRECO with CHARM (headreco will be removed in SimNIBS 4) * MNI normalization: More options: * DARTEL / SHOOT * BrainSuite (wait for Anand) |
| Line 245: | Line 169: |
| * Render surface envelope in the MRI as a thin line instead of the full interpolation matrix * Edit fiducials: Replace 6 text boxes with 1 for easy copy-paste (see fiducials.m) * Optimize computation interpolation MRI-surface (tess_tri_interp) => spm_mesh_to_grid |
* Render surface envelope in the MRI as a thin line instead of the full interpolation matrix<<BR>>Or use inpolyhedron to get a surface mask and then erode it to get the volume envelope * Surface>Volume interpolation: Use '''spm_mesh_to_grid''' instead of tess_tri_interp * Defacing: * https://afni.nimh.nih.gov/pub/dist/doc/htmldoc/tutorials/refacer/refacer_run.html * Removing MNI face mask using MNI coordinates * Atlas switch in 3D MRI figures * Bug import anatomy: Requested nVert > high-resolution cortex surface: Creates an empty cortex_0V * ICBM152 update: * Process with FreeSurfer 7.2 + add FS atlases (Brainnettome, Schaeffer, HCP...) * Add volume atlases (+ reimport ASEG as volatlas) * Add facemask => Use for defacing with any MNI registration * Add T2? |
| Line 249: | Line 182: |
| * Add new labels to all BrainSuite anatomy templates | |
| Line 251: | Line 183: |
| * Pediatric head atlases: https://www.pedeheadmod.net/pediatric-head-atlases-v1-2/ | * Use for volume coregistration (rigid / non-rigid) * USCBrain: Add default electrodes positions * Remove BrainSuite1 when not needed anymore * Templates for different ages: * MNI: https://www.bic.mni.mcgill.ca/ServicesAtlases/NIHPD-obj1 * Pediatric head atlases: https://www.pedeheadmod.net/pediatric-head-atlases-v1-2/ * https://iopscience.iop.org/article/10.1088/2057-1976/ab4c76 * https://www.biorxiv.org/content/biorxiv/early/2020/02/09/2020.02.07.939447.full.pdf * John Richards: https://www.nitrc.org/frs/?group_id=1361 * Neurodev database: https://jerlab.sc.edu/projects/neurodevelopmental-mri-database/ |
| Line 254: | Line 197: |
| * Display scouts in a tree: hemisphere, region, subregion * Sort scouts by region in process options * Downsample to atlas: allow on timefreq/connect files |
|
| Line 259: | Line 199: |
| * Major bug when importing surfaces for an MRI that was re-oriented manually * Surface>Volume interpolation: Use spm_mesh_to_grid * Bug: Hide scouts in the preview of the grid for volume head models |
|
| Line 267: | Line 204: |
| * Add history: Save modifications and transformations applied to the channel files (Marcel) | |
| Line 269: | Line 205: |
| * Add menu to import implantation channel file in imported recordings | * Create clusters from anatomical labels: * Identify contacts in a given anatomical region (volume scout, surface mesh, or label in a volume atlas) / allow extracting the signals from all the contacts in an ROI |
| Line 272: | Line 209: |
| * Arnulfo: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6 * MAP07 / SPM: https://www.epi.ch/_files/Artikel_Epileptologie/Huppertz_2_13.pdf |
* SEEG DEETO Arnulfo 2015: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6 * Used routinely at Niguarda Hospital + other hospitals worldwide, reliable tool. * To be used with SEEG-assistant/3DSlicer: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1545-8 * ECOG Centracchio 2021: https://link.springer.com/content/pdf/10.1007/s11548-021-02325-0.pdf * Classifier on thresholded CT: https://github.com/Jcentracchio/Automated-localization-of-ECoG-electrodes-in-CT-volumes * SEEG Granados 2018 (no code shared): https://link.springer.com/content/pdf/10.1007/s11548-018-1740-8.pdf |
| Line 277: | Line 218: |
| * ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage) | * ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage) <<BR>>Normalization MNI? solutions with FieldTrip? |
| Line 285: | Line 226: |
| * Stat on connectivity? * Stat on unconstrained sources? * Stat/time series: Hide lines going down to zero (Dimitrios: https://neuroimage.usc.edu/forums/t/common-source-activation-across-subjects-and-conditions/1152/21) * Cluster stat: Add frequency selection option |
|
| Line 286: | Line 231: |
| * Which functions to use? * Write panel similar to Process1 and Process2 to allow the |
* Write panel similar to Process1 and Process2 |
| Line 290: | Line 234: |
| * Quality control before statistics, on condition averages across subjects:<<BR>>mean(baseline)/std(baseline): shows bad subject quickly. * Use SurfStat: Impements interesting things, like an analytical cluster-based p-value correction (Random-field theory which is used in SPM) - Peter * Export to R or SPSS for advanced stat |
* Multivariate stim-response analysis: https://github.com/mickcrosse/mTRF-Toolbox |
| Line 295: | Line 237: |
| * '''XDF import''': Use FieldTRip or the EEGLAB plugin, contact Martin Bleichner (Oldenburg)<<BR>>https://github.com/sccn/xdf/blob/master/xdf_sample.xdf | * BIDS import: * Read real fiducials (OMEGA) / transformation matrices: * https://groups.google.com/g/bids-discussion/c/BeyUeuNGl7I * https://github.com/bids-standard/bids-specification/issues/752#issuecomment-795880992 * Read associated empty room * BIDS export: * Add events tsv, channel tsv, EEG, iEEG * BIDS-Matlab? * Support for OpenJData / JNIfTI: https://github.com/brainstorm-tools/brainstorm3/issues/284 |
| Line 299: | Line 249: |
| * FieldTrip: Import/Export time-frequency: * Export: http://neuroimage.usc.edu/forums/t/export-time-frequency-to-fieldtrip/1968 * Import: http://neuroimage.usc.edu/forums/t/import-time-frequency-data-from-fieldtrip/2644 |
* SPM .mat/.dat: Fix the import of the EEG/SEEG coordinates * EEG File formats: * XDF: https://github.com/sccn/xdf * XLTEK: https://github.com/danielmhanover/OpenXLT * Persyst .lay: https://github.com/ieeg-portal/Persyst-Reader * Nervus .eeg: https://github.com/ieeg-portal/Nervus-Reader * Biopac .acq: https://github.com/ieeg-portal/Biopac-Reader * BCI2000 Input (via EEGLAB plugin) |
| Line 303: | Line 258: |
| * Use reader from MNE-Python: mne.io.read_raw_kit (doesn't require Yokogawa slow library) | * Use reader from MNE-Python: mne.io.read_raw_kit (skip Yokogawa slow library) |
| Line 309: | Line 264: |
| * EEG File formats: * EEG CeeGraph * EGI: Finish support for epoched files (formats 3,5,7) * XLTEK: https://github.com/danielmhanover/OpenXLT * Persyst .lay: https://github.com/ieeg-portal/Persyst-Reader * Nervus .eeg: https://github.com/ieeg-portal/Nervus-Reader * Biopac .acq: https://github.com/ieeg-portal/Biopac-Reader * gTec EEG recordings: Read directly from the HDF5 files instead of the Matlab exports. * BCI2000 Input (via EEGLAB plugin) |
|
| Line 319: | Line 265: |
| * Review raw on all the file formats (ASCII EEG and Cartool missing) * SPM .mat/.dat: Fix the import of the EEG/SEEG coordinates * Get acquisition date from files: Missing for 4D |
|
| Line 324: | Line 267: |
| * All tutorial datasets in BIDS (including introduction tutorials) * Count GitHub clones in the the download stats * Deface the MRIs of all the tutorials |
|
| Line 325: | Line 271: |
| * Update the organization of derivatives folder (same for ECOG tutorial) | |
| Line 329: | Line 276: |
| * Rat PAC + high gamma (Soheila) | |
| Line 332: | Line 278: |
| * FieldTrip cortico-muscular coherence tutorial: http://www.fieldtriptoolbox.org/tutorial/coherence | |
| Line 336: | Line 281: |
| * BIDS-EEG example datasets * Stand-alone ICA tutorial * Move all the files to download to the cloud for faster download everywhere in the world * Provide secure way of sending password over HTTPS for: * Account creation * Forum exchanges * org.brainstorm.dialog.CloneControl * Workflows FieldTrip: http://www.fieldtriptoolbox.org/faq/what_types_of_datasets_and_their_respective_analyses_are_used_on_fieldtrip * Count GitHub clones in the the download stats * Deface the MRIs of all the tutorials * Clean up the wiki: * Remove all the wiki pages that are not used * Check all the links in all the pages * Check that all the TODO blocks have been properly handled * Remove useless images from all tutorials * Update page count on the main tutorials page |
* Reproduce results from "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x * Stand-alone ICA tutorial |
| Line 364: | Line 294: |
| * MacOS 10.14.5 (Mojave): * Toggle buttons do not show their status * Panel Record: Text is too large for text boxes |
|
| Line 368: | Line 301: |
| * in_bst_data_multi: If trials have different sizes, output is random (the one of the first file) * Canolty maps computation: Fix progress bar |
== Distributed computing == * Options from FieldTrip: * Loose collection of computers: https://github.com/fieldtrip/fieldtrip/tree/master/peer * Single multicore machine: https://github.com/fieldtrip/fieldtrip/tree/master/engine * Batch system: https://github.com/fieldtrip/fieldtrip/tree/master/qsub * Documentation: https://www.fieldtriptoolbox.org/faq/what_are_the_different_approaches_i_can_take_for_distributed_computing/ * PSOM: http://psom.simexp-lab.org/ * Google: https://www.youtube.com/watch?v=LLMXV3o2FT0 * https://edu.google.com/why-google/case-studies/unc-chapel-hill-gcp/ |
| Line 372: | Line 313: |
| * bst_bsxfun: After 2016b, we can use directly the scalar operators (./ .* ...) instead of bsxfun. Update bst_bsxfun to skip the use of bsxfun when possible. | * Replace all calls to inpolyhd.m with inpolyhedron.m (10x faster) |
| Line 374: | Line 315: |
| * Hide Java panels instead of deleting them | |
| Line 377: | Line 317: |
| * bst_warp and channel_project: Use tess_parametrize_new instead of tess_parametrize * Shared kernels: "get bad channels" operation in a different way (reading all the files is too slow) * Optimize bst_get: * Now study and subject have necessarily the same folder name * Replace big switch with separate functions * Fix all the 'todo' blocks in the code |
What's next
A roadmap to the future developments of Brainstorm.
Contents
Recordings
- Default montages for EEG (sensor selection)
- Sleep scoring wish list (Emily C):
- Configurable horizontal lines (for helping detecting visually some thresholds)
- Mouse ruler: Measure duration and amplitude by dragging the mouse.
- Automatic spindle detector
https://neuroimage.usc.edu/forums/t/page-overlap-while-reviewing-raw-file-a-way-to-set-to-0/11229/13
- RAW file viewer speed:
- Downsample before filtering? (attention to the filter design)
- Add parameter to make the visual downsampling more or less aggressive
- Pre-load next page of recordings
- Keep the filter specifications in memory instead of recomputing for every page
- Review signals in time-frequency space
- MEG/EEG registration: Apply the same transformation to multiple runs
- Create heat maps: Maybe with matlab function heatmap?
- Import window: Add button to create the corresponding processing pipeline (to generate script or to edit additional options)
BioSemi: Add menu "Convert naming system" to rename channels into 10-10 (A1=>FPz)
Simulations: https://github.com/lrkrol/SEREEGA
- Integrate with EYE-EEG (Olaf Dimigen)
Reproduce tutorial: https://www.eyetracking-eeg.org/tutorial.html
Add note when rejecting trials: https://neuroimage.usc.edu/forums/t/33686
Interface
Add a warning when computing a forward model with > 100000 sources (check selection)
- Snapshot: Save as image / all figures (similar to Movie/all figure)
Generalize the use of the units (field .DisplayUnits): Rewrite processes to save the units correctly
- Colormaps:
- Allow brightness/contrast manipulations on the custom colormaps
- Global colormap max: Should get the maximum across all the open files
- Copy figures to clipboard (with the screencapture function)
Contact sheets & movies: use average of time windows instead of single instants, for each picture.
- Contact sheets: Allow explicit list of times in input (+ display as in MNE-Python with TS)
- Display CTF coils: Show discs instead of squares
- Progress bar: Add a "Cancel" button
- Error message: Add a link to report directly the bug on the forum
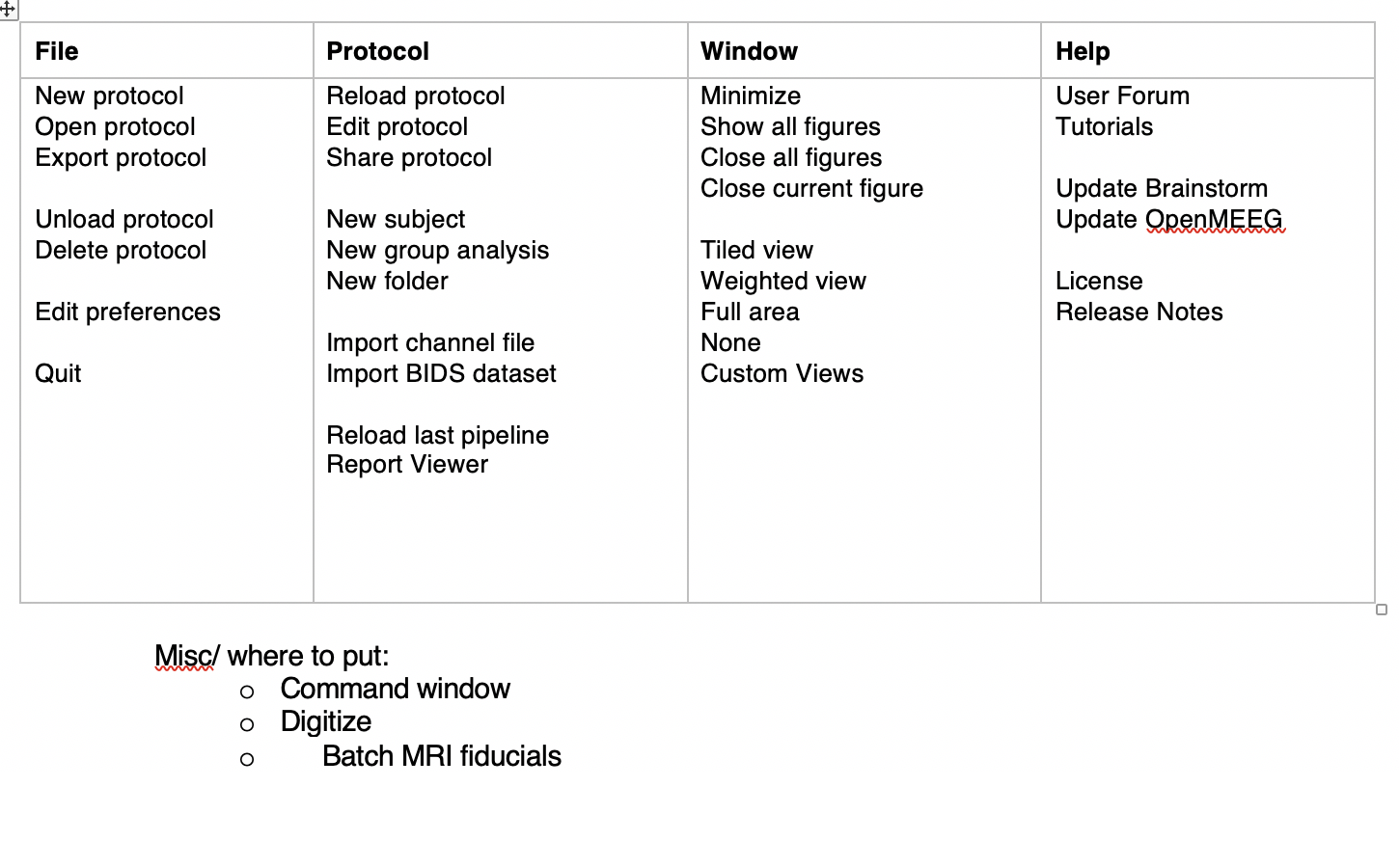
Reorganize menus (Dannie's suggestion):
Connectivity
- Thresholding and stat tests for connectivity matrices
- Connect NxN display:
- Graph on sensors: does not place the sensors correctly in space
- Display as image: Add legend of the elements along X and Y axis
- Display as time series: Display warning before trying to open too many signals
- Optimize display: use surface() instead of line() for links? (as in figure_3d/PlotFibers)
- Time-resolved correlation/coherence: Display as time bands
- Weighted Phase Lag Index (WPLI)
- Granger: Check for minimum time window (Esther: min around 500-1000 data points)
- PLV:
- Remove evoked
- Add time integration
- Unconstrained sources
- Add warning when running of short windows (because of filters)
Processes
- Generate fully reproducible scripts, including all the interactive/graphical parts:
- Saving all the interactive operations as process calls
- Improving the pipeline editor to handle loops over data files or subjects
- Keeping a better track of the provenance of all the data (History, uniform file names)
- Add MNE-Python functions:
- scikit-learn classifiers
https://neuroimage.usc.edu/forums/t/ica-on-very-long-eeg/23556/4
https://neuroimage.usc.edu/forums/t/best-way-to-export-to-mne-python/12704/3
- Reproduce other tutorials / examples
Point-spread functions (PSFs) and cross-talk functions: https://mne.tools/stable/auto_examples/inverse/plot_psf_ctf_vertices.html#sphx-glr-auto-examples-inverse-plot-psf-ctf-vertices-py
Spatial resolution metrics in source space:
https://mne.tools/stable/auto_examples/inverse/plot_resolution_metrics.html#sphx-glr-auto-examples-inverse-plot-resolution-metrics-py- Change the graphic renderer from Matlab
Chronux toolbox : http://chronux.org/
Add FieldTrip functions:
- ft_sourceanalysis:
- Check noise covariance
- Check all the options of all the methods
- Single trial reconstructions + noise covariance?
Filters?? http://www.fieldtriptoolbox.org/example/common_filters_in_beamforming
Beamformers: Save ftSource.avg.mom
http://www.fieldtriptoolbox.org/workshop/meg-uk-2015/fieldtrip-beamformer-demohttp://www.fieldtriptoolbox.org/tutorial/beamformingextended
- Baseline? Two inputs?
- ft_prepare_heamodel: Add support from BEM surfaces from the Brainstorm database
- Freqanalysis: ITC
ft_volumereslice: http://www.fieldtriptoolbox.org/faq/how_change_mri_orientation_size_fov
- ft_freqanalysis
- ft_combineplanar
- ft_sourceanalysis:
- Optimization:
Test speed for writing files:
https://undocumentedmatlab.com/articles/improving-fwrite-performance- Use CUDA for speeding up some operations (filtering, wavelets, etc)
- Use Matlab Coder to optimize: Wavelets, bandpass filter, sinusoid removal
- Pipeline editor:
- Bug: After "convert to continuous", the time of the following processes should change
- Add loops over subjects/conditions/trial groups
- Events: Allow selection from a drop-down list (similar to option "channelname" in panel_process_selection)
ITC: Inter-trial coherence (see MNE reports for group tutorial)
http://www.sciencedirect.com/science/article/pii/S1053811916304232- ICA:
Add Alex's suggestions: https://neuroimage.usc.edu/forums/t/ica-on-very-long-eeg/23556/4
Automatic classification: ICLabel: https://neuroimage.usc.edu/forums/t/automatic-eeg-ic-ica-classification-for-brainstorm/33785
- Add methods: SOBI, Fastica, AMICA/CUDICA/CUDAAMICA (recommended by S Makeig)
- Why doesn't the ICA process converge when using 25 components in the EEG tutorial?
- Add an option to resample the signals before computing the ICA decomposition
- Exploration: Add window with spectral decomposition (useful for muscle artifacts)
- Export IC time series (and then compute their spectrum): solves the problem above
Comparison JADE/Infomax:
http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0030135Dimension reduction with PCA adds artifacts: Not done by default in EEGLAB
Contact: Stephen Shall Jones ( shall-jones@infoscience.otago.ac.nz )
Student Carl Leichter detailed this in his thesis- Import ICA matrices available in EEGLAB .set files
EEGLAB recommends ICA + trial rejection + ICA again: Impossible right now with Brainstorm
(http://sccn.ucsd.edu/wiki/Chapter_09:_Decomposing_Data_Using_ICA)ICA+machine learning: https://www.ncbi.nlm.nih.gov/pubmed/28497769
Automated artifact rejection: https://arxiv.org/abs/1612.08194
Use EYE-EEG: EEGLAB toolbox for eye-tracker guided ICA (Olaf Dimigen): http://www2.hu-berlin.de/eyetracking-eeg/
- SSP:
Display warning if changing the ChannelFlag while there is a Projector applied
Remove line noise: http://www.nitrc.org/projects/cleanline
- Time-frequency:
- Optimization: bst_timefreq (around l.136), remove evoked in source space: Average should be computed in sensor space instead of source space (requested by Dimitrios)
Short-time Fourier transform: http://www.mikexcohen.com/lectures.html
- Hilbert with time bands very slow on very long files (eg. 3600s at 1000Hz) because the time vector is still full (10^7 values): save compressed time vector instead.
- When normalizing with baseline: Propagate with the edge effects marked in TFmask
- Allow running TF on montages
- Review continuous files in time-frequency space (for epilepsy)
- Bug when computing TF on constrained and unconstrained scouts at the same time (in mixed head models for instance): uses only the constrained information and doesn't sum the 3 orientations for the unconstrained regions.
Source modeling
Reproduce results in "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x
- eLORETA instead of sLORETA?
https://neuroimage.usc.edu/forums/t/compute-eeg-sources-with-sloreta/13425/6
"eLORETA algorithm is available in the MEG/EEG Toolbox of Hamburg (METH)": https://www.biorxiv.org/content/biorxiv/early/2019/10/17/809285.full.pdf
Sensitivity maps: https://mne.tools/stable/auto_examples/forward/plot_forward_sensitivity_maps.html
- Point-spread and cross-talk functions (code in MNE-Python):
- Dipoles:
- Display dipoles in MRI viewer
- Project individual dipoles files on a template
- panel_dipoles: Doesn't work with multiple figures
- Project sources: Very poor algorithm to project sub-cortical regions and cerebellum
- Maximum:
Menu Sources > Maximum value: Doesn't work with volume or mixed head models
- Panel Get coordinates: Add button "find maximum"
Sources on surface: Display peak regions over time (time = color) => A.Gramfort
- BEM single sphere: Get implementation from MNE
- Volume grid:
- Optimize: 3D display (better than 9x9 cubes)
- Optimize: vol_dilate (with 26 neighbors)
- Add eyes models to attract eye activity
- Display spectrum scouts (PSD plots when clicking on "Display scouts" on PSD/full cortex)
Anatomy
- BEM surfaces: Deform fieldtrip BEM surfaces from ICBM152 to subject space with MNI coordinates?
Simple-brain-plot: https://github.com/dutchconnectomelab/Simple-Brain-Plot
MRI segmentation:
- SimNIBS: Replace HEADRECO with CHARM (headreco will be removed in SimNIBS 4)
- MNI normalization: More options:
- DARTEL / SHOOT
BrainSuite (wait for Anand)
- MRI Viewer:
- Pan in zoomed view (shift + click + move?)
- Zoom in/out with mouse (shift + scroll?)
- Ruler tool to measure distances
- Display scouts as additional volumes
Render surface envelope in the MRI as a thin line instead of the full interpolation matrix
Or use inpolyhedron to get a surface mask and then erode it to get the volume envelopeSurface>Volume interpolation: Use spm_mesh_to_grid instead of tess_tri_interp
- Defacing:
https://afni.nimh.nih.gov/pub/dist/doc/htmldoc/tutorials/refacer/refacer_run.html
- Removing MNI face mask using MNI coordinates
- Atlas switch in 3D MRI figures
Bug import anatomy: Requested nVert > high-resolution cortex surface: Creates an empty cortex_0V
- ICBM152 update:
Process with FreeSurfer 7.2 + add FS atlases (Brainnettome, Schaeffer, HCP...)
- Add volume atlases (+ reimport ASEG as volatlas)
Add facemask => Use for defacing with any MNI registration
- Add T2?
BrainSuite:
- Use same colors for left and right for anatomical atlases
- Use for volume coregistration (rigid / non-rigid)
- USCBrain: Add default electrodes positions
Remove BrainSuite1 when not needed anymore
- Templates for different ages:
MNI: https://www.bic.mni.mcgill.ca/ServicesAtlases/NIHPD-obj1
Pediatric head atlases: https://www.pedeheadmod.net/pediatric-head-atlases-v1-2/
https://www.biorxiv.org/content/biorxiv/early/2020/02/09/2020.02.07.939447.full.pdf
John Richards: https://www.nitrc.org/frs/?group_id=1361
Neurodev database: https://jerlab.sc.edu/projects/neurodevelopmental-mri-database/
- Scouts:
- Display edges in the middle of the faces instead of the vertices
Project from one hemisphere to the other using registered spheres/squares (http://neuroimage.usc.edu/forums/t/how-to-create-mirror-roi-in-the-other-hemisphere/5910/8)
- Parcellating volume grids: scikit-learn.cluster.Ward
Geodesic distance calculations:
https://www.mathworks.com/matlabcentral/fileexchange/6110-toolbox-fast-marching
ECOG/SEEG
- Contact positions: Import / set / detect
- New option: Align on none|inner|cortex to replace ECOG-mid
- Project contact positions across subjects or templates (Marcel)
- Create clusters from anatomical labels:
- Identify contacts in a given anatomical region (volume scout, surface mesh, or label in a volume atlas) / allow extracting the signals from all the contacts in an ROI
- Automatic segmentation of CT:
SEEG DEETO Arnulfo 2015: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6
- Used routinely at Niguarda Hospital + other hospitals worldwide, reliable tool.
To be used with SEEG-assistant/3DSlicer: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1545-8
ECOG Centracchio 2021: https://link.springer.com/content/pdf/10.1007/s11548-021-02325-0.pdf
Classifier on thresholded CT: https://github.com/Jcentracchio/Automated-localization-of-ECoG-electrodes-in-CT-volumes
SEEG Granados 2018 (no code shared): https://link.springer.com/content/pdf/10.1007/s11548-018-1740-8.pdf
- ECOG:
- Project and display contacts on cortex surface should consider the rigidity of the grids: Contacts cannot rotate, and distance between contacts should remain constant across runs
Method for contacts projection: https://pdfs.semanticscholar.org/f10d/6b899d851f3c4b115404298d7b997cf1d5ab.pdf
ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage)
Normalization MNI? solutions with FieldTrip?
- Display:
- Bad channels: Contacts greyed out instead of ignored (Marcel)
- Display time in H:M:S
- Display curved SEEG electrodes
Detection CEEP stim artifacts: Use ImaGIN code ImaGIN_StimDetect
Statistics
- Stat on connectivity?
- Stat on unconstrained sources?
Stat/time series: Hide lines going down to zero (Dimitrios: https://neuroimage.usc.edu/forums/t/common-source-activation-across-subjects-and-conditions/1152/21)
- Cluster stat: Add frequency selection option
- ANOVA:
- Write panel similar to Process1 and Process2
- Output = 1 file per effect, all grouped in a node "ANOVA"
- Display several ANOVA maps (from several files) on one single figure, using a "graphic accumulator", towards which one can send any type of graphic object
Multivariate stim-response analysis: https://github.com/mickcrosse/mTRF-Toolbox
Input / output
- BIDS import:
- Read real fiducials (OMEGA) / transformation matrices:
- Read associated empty room
- BIDS export:
- Add events tsv, channel tsv, EEG, iEEG
- BIDS-Matlab?
Support for OpenJData / JNIfTI: https://github.com/brainstorm-tools/brainstorm3/issues/284
- DICOM converter:
- Add dcm2nii (MRICron)
- Add MRIConvert
- SPM .mat/.dat: Fix the import of the EEG/SEEG coordinates
- EEG File formats:
Persyst .lay: https://github.com/ieeg-portal/Persyst-Reader
Nervus .eeg: https://github.com/ieeg-portal/Nervus-Reader
Biopac .acq: https://github.com/ieeg-portal/Biopac-Reader
- BCI2000 Input (via EEGLAB plugin)
- 4D file format:
- Use reader from MNE-Python: mne.io.read_raw_kit (skip Yokogawa slow library)
- Reference gradiometers: Keep the orientation of the first or second coil?
- Reference gradiometers: Add the sensor definition from coil_def.dat
- Validate with phantom recordings that noise compensation is properly taken into account
- The noise compensation is considered to be always applied on the recordings, not sure this assumption is always correct
- 4D phantom tutorial (JM Badier?)
- BST-BIN: Add compression to .bst
Distribution & documentation
- All tutorial datasets in BIDS (including introduction tutorials)
Count GitHub clones in the the download stats
- Deface the MRIs of all the tutorials
- Tutorial OMEGA/BIDS:
- Update the organization of derivatives folder (same for ECOG tutorial)
- Add review of literature for the resting state MEG
- Download example datasets directly from the OMEGA repository
New tutorials:
Other public datasets: https://github.com/INCF/BIDS-examples/tree/bep008_meg/
- EEG/research
FieldTrip ECOG tutorial: http://www.fieldtriptoolbox.org/tutorial/human_ecog
Reproduce tutorials from MNE-Python: https://martinos.org/mne/stable/tutorials.html
Cam-CAN database: https://camcan-archive.mrc-cbu.cam.ac.uk/dataaccess/<<BR>>(download new datasets, including maxfiltered files and manual fiducial placements)
- MEG steady-state / high-gamma visual / frequency tagging
Reproduce results from "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x
- Stand-alone ICA tutorial
Current bugs
- Image viewer:
- Difficult to get to 100%
- Buggy on some systems
- 2DLayout:
- (TF) Units are weird with % values
- (TF) Difficult to navigate in frequencies: Scaling+changing frequency resets the scaling
- Progress bar:
- Doesn't close properly on some Linux systems
- Focus requests change workspace when processing constantly (Linux systems)
- MacOS 10.14.5 (Mojave):
- Toggle buttons do not show their status
- Panel Record: Text is too large for text boxes
- MacOS bugs:
- Buttons {Yes,No,Cancel} listed backwards
- Record tab: Text of epoch number is too big
- Colormap menus: Do not work well on compiled MacOSX 10.9.5 and 10.10
Distributed computing
Options from FieldTrip:
Loose collection of computers: https://github.com/fieldtrip/fieldtrip/tree/master/peer
Single multicore machine: https://github.com/fieldtrip/fieldtrip/tree/master/engine
Batch system: https://github.com/fieldtrip/fieldtrip/tree/master/qsub
Documentation: https://www.fieldtriptoolbox.org/faq/what_are_the_different_approaches_i_can_take_for_distributed_computing/
Geeky programming details
- Replace all calls to inpolyhd.m with inpolyhedron.m (10x faster)
Interface scaling: Rewrite class IconLoader to scale only once the icons at startup instead of at each request of an icon (might improve the speed of the rendering of the tree)
- Processes with "radio" and "radio_line" options: Replace with "radio_label" and "radio_linelabel"
- Interpolations: Use scatteredInterpolant, griddedInterpolant, triangulation.nearestNeighbor (2014b)
