|
Size: 19921
Comment:
|
Size: 22126
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 7: | Line 7: |
| * Default montages for EEG (sensor selection) | * Review signals in time-frequency space * Events processes: Select events names from a list instead of having to type them * Display CTF coils: Show discs instead of squares |
| Line 10: | Line 12: |
| * Mouse ruler: Measure duration and amplitude by dragging the mouse. | * Mouse ruler: Measure amplitude by dragging the mouse. |
| Line 13: | Line 15: |
| * RAW file viewer speed: * Downsample before filtering? (attention to the filter design) |
* RAW file viewer speed (Low priority) :<<BR>> * Consider to change to a format that is faster to read |
| Line 16: | Line 19: |
| * Pre-load next page of recordings * Keep the filter specifications in memory instead of recomputing for every page * Review signals in time-frequency space |
* Keep the filter specifications in memory instead of recomputing for every page<<BR>>(Nice to have) |
| Line 20: | Line 22: |
| * Simulations: https://github.com/lrkrol/SEREEGA == Coregistration == * Template channel file on a subject anatomy: Apply the MNI transform * MEG/EEG registration: Apply the same transformation to multiple runs |
* Simulations: https://github.com/lrkrol/SEREEGA(Low priority) == ECOG/SEEG == * https://www.sciencedirect.com/science/article/pii/S1053811922005559 * Display (high-priority)(Part SEEG grant): * Group display: Overlay multiple channel files in the same figure, coloring contacts by subject/ROI/Cluster/Electrode name * https://neuroimage.usc.edu/forums/t/37617 * iEEG tab must be read-only when multiple files (hide configuration controls) * Bad channels: Contacts greyed out instead of ignored (Marcel H, Germany)<<BR>>(To diff between band and not-recorded) > Rendering of SEEG electrodes: Full surface modelling with surface mesh (see Lead-DBS models + code that generates them?) * Display time in H:M:S instead of S > If there is t0 in H:M:S instead of S > As an option in Display configuration button>x-axis * view_leadfield_sensitivity: Add closing surfaces at cortex limits * Create clusters from anatomical labels (Anne So) : * Identify contacts in a given anatomical region (volume scout, surface mesh, or label in a volume atlas) / allow extracting the signals from all the contacts in an ROI> As a process to select recordings, then Scouts from Volumen Atlas, Create cluster in channel file, then Extract time series. * Group analysis: extract clusters across subjects, display or average signals (see MIA) (Anne So) * Spike detection (Need to check for current toolboxes from scratch)(contact Nicolas R)(Mosher J)(iEEG BIDS): * https://iopscience.iop.org/article/10.1088/1741-2552/ac9259/pdf * Automatic segmentation of CT: * LeGUI: https://github.com/Rolston-Lab/LeGUI/tree/main/LeGUI<<BR>>https://neuroimage.usc.edu/forums/t/automatic-localization-of-seeg-electrodes/36302/7 * GARDEL: http://meg.univ-amu.fr/wiki/GARDEL:presentation * Discussed with Samuel and Christian (ins-amu.fr) * SEEG DEETO Arnulfo 2015: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6 * Used routinely at Niguarda Hospital + other hospitals worldwide, reliable tool. * To be used with SEEG-assistant/3DSlicer: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1545-8 * ECOG Centracchio 2021: https://link.springer.com/content/pdf/10.1007/s11548-021-02325-0.pdf * Classifier on thresholded CT: https://github.com/Jcentracchio/Automated-localization-of-ECoG-electrodes-in-CT-volumes * SEEG Granados 2018 (no code shared): https://link.springer.com/content/pdf/10.1007/s11548-018-1740-8.pdf * ECOG: * Project and display contacts on cortex surface should consider the rigidity of the grids: Contacts cannot rotate, and distance between contacts should remain constant across runs * Method for contacts projection: https://pdfs.semanticscholar.org/f10d/6b899d851f3c4b115404298d7b997cf1d5ab.pdf * ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage) <<BR>>Normalization MNI? solutions with FieldTrip? * Display CT images: Better brightness/contrast adjustment: https://neuroimage.usc.edu/forums/t/automatic-localization-of-seeg-electrodes/36302/8 Range of values is way diff than ones from MRI. Current color maps are not suitable for CT, need to be improved.Together with processing of CT to get electrode positions. * Detection CCEP stim artifacts: Use ImaGIN code ImaGIN_StimDetect https://f-tract.eu/software/imagin/ |
| Line 27: | Line 75: |
| * process_detectbad: * Allow on raw files (for bad channels only) * Add detection on derivative of the signal (see EEGLAB) * Document in tutorial Bad channels * PREP pipeline / EEGLAB (Bigdely-Shamlo 2015) * Improve bad channel/trial detection: * ft_artifact_threshold and ft_rejectartifact * MNE-Python * EEGLAB |
|
| Line 33: | Line 92: |
| * Add note when rejecting trials: https://neuroimage.usc.edu/forums/t/33686 * ICA: * Add Alex's suggestions: https://neuroimage.usc.edu/forums/t/ica-on-very-long-eeg/23556/4 |
* Use it to guide ICA: http://www2.hu-berlin.de/eyetracking-eeg/ * ICA: <<BR>> |
| Line 39: | Line 96: |
| * Add methods: SOBI, Fastica, AMICA/CUDICA/CUDAAMICA (recommended by S Makeig) * Why doesn't the ICA process converge when using 25 components in the EEG tutorial? * Add an option to resample the signals before computing the ICA decomposition |
|
| Line 44: | Line 99: |
| * Comparison JADE/Infomax: <<BR>> http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0030135 * Dimension reduction with PCA adds artifacts: Not done by default in EEGLAB<<BR>>Contact: Stephen Shall Jones ( shall-jones@infoscience.otago.ac.nz )<<BR>>Student Carl Leichter detailed this in his thesis |
|
| Line 47: | Line 100: |
| * EEGLAB recommends ICA + trial rejection + ICA again: Impossible right now with Brainstorm<<BR>>(http://sccn.ucsd.edu/wiki/Chapter_09:_Decomposing_Data_Using_ICA) | |
| Line 49: | Line 101: |
| Line 50: | Line 103: |
| * Use EYE-EEG: EEGLAB toolbox for eye-tracker guided ICA (Olaf Dimigen): http://www2.hu-berlin.de/eyetracking-eeg/ | * Spectral representation of ICs |
| Line 54: | Line 108: |
| * File format: * Add support to read GDF file format https://github.com/donnchadh/biosig/blob/master/biosig/t200_FileAccess/sload.m * <<BR>> * |
|
| Line 57: | Line 117: |
| * Import window: Add button to create the corresponding processing pipeline (to generate script or to edit additional options) * Record all GUI actions |
* Generate fully reproducible scripts, including all the interactive/graphical parts * Record all GUI actions as script calls * Import window: Add button to create the corresponding processing pipeline (to generate script or to edit additional options). * Adding the list of plugins to the reports |
| Line 64: | Line 126: |
| * Snapshot: Save as image / all figures (similar to Movie/all figure) * Colormaps: * Allow brightness/contrast manipulations on the custom colormaps * Global colormap max: Should get the maximum across all the open files * Copy figures to clipboard (with the screencapture function) |
* Colormaps: Global colormap max: Should get the maximum across all the open files * Snapshot: * Save as image / all figures (similar to Movie/all figure) |
| Line 70: | Line 132: |
| Line 71: | Line 134: |
| * Display CTF coils: Show discs instead of squares * Error message: Add a link to report directly the bug on the forum * Reorganize menus (Dannie's suggestion): {{attachment:dannie_menus.png||width="382",height="237"}} |
|
| Line 77: | Line 137: |
| * Generalize the use of the units (field .DisplayUnits) | * Generalize the use of the units (field .DisplayUnits): Save in source files |
| Line 80: | Line 140: |
| * Thresholding and stat tests for connectivity matrices: * Panel Display: Show only the top N% measures * Documentation |
* Define names and unit labels for each connectivity metric * Null models: (Bratislav M) https://www.nature.com/articles/s41583-022-00601-9 * {{attachment:connect_toolboxes.jpg}} |
| Line 84: | Line 146: |
| * Graph on sensors: does not place the sensors correctly in space | * Graph on sensors: Place M/EEG sensors by location, not by channel order |
| Line 88: | Line 150: |
| * Time-resolved correlation/coherence: Display as time bands | * Time-resolved correlation/coherence: Display as time bands (as done in wavelet, to have same time axis as data) |
| Line 91: | Line 154: |
| * Generate fully reproducible scripts, including all the interactive/graphical parts: * Saving all the interactive operations as process calls * Improving the pipeline editor to handle loops over data files or subjects * Keeping a better track of the provenance of all the data (History, uniform file names) |
|
| Line 97: | Line 156: |
| * https://neuroimage.usc.edu/forums/t/ica-on-very-long-eeg/23556/4 | * BEM single layer (John wants to test it) |
| Line 99: | Line 159: |
| * Reproduce other tutorials / examples * Point-spread functions (PSFs) and cross-talk functions: https://mne.tools/stable/auto_examples/inverse/plot_psf_ctf_vertices.html#sphx-glr-auto-examples-inverse-plot-psf-ctf-vertices-py |
* Reproduce tutorials / examples from FieldTrip and MNE-Python: * FieldTrip ECOG tutorial: http://www.fieldtriptoolbox.org/tutorial/human_ecog * Reproduce tutorials from MNE-Python: https://martinos.org/mne/stable/tutorials.html |
| Line 102: | Line 165: |
| Line 103: | Line 167: |
| Line 104: | Line 169: |
| Line 110: | Line 176: |
| Line 111: | Line 178: |
| Line 112: | Line 180: |
| Line 113: | Line 182: |
| Line 114: | Line 184: |
| Line 115: | Line 186: |
| * Freqanalysis: ITC | |
| Line 117: | Line 187: |
| Line 119: | Line 190: |
| Line 120: | Line 192: |
| * Test speed for writing files: <<BR>>https://undocumentedmatlab.com/articles/improving-fwrite-performance | |
| Line 123: | Line 194: |
| Line 127: | Line 199: |
| Line 128: | Line 201: |
| Line 129: | Line 203: |
| Line 130: | Line 205: |
| * Optimization: bst_timefreq (around l.136), remove evoked in source space: Average should be computed in sensor space instead of source space (requested by Dimitrios) | * Optimization: bst_timefreq (around l.136), remove evoked in source space: Average should be computed in sensor space instead of source space (requested by Dimitrios) |
| Line 132: | Line 207: |
| Line 135: | Line 211: |
| * Review continuous files in time-frequency space (for epilepsy) | |
| Line 137: | Line 212: |
== Source modeling == * Reproduce results in "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x * eLORETA instead of sLORETA? * https://neuroimage.usc.edu/forums/t/compute-eeg-sources-with-sloreta/13425/6 * "eLORETA algorithm is available in the MEG/EEG Toolbox of Hamburg (METH)": https://www.biorxiv.org/content/biorxiv/early/2019/10/17/809285.full.pdf * https://github.com/brainstorm-tools/brainstorm3/issues/114 * Sensitivity maps: https://mne.tools/stable/auto_examples/forward/plot_forward_sensitivity_maps.html * Point-spread and cross-talk functions (code in MNE-Python): * https://www.biorxiv.org/content/biorxiv/early/2019/06/18/672956.full.pdf * https://github.com/olafhauk/EEGMEGResolutionAtlas * Dipoles: * Display dipoles in MRI viewer * Project individual dipoles files on a template * panel_dipoles: Doesn't work with multiple figures * Project sources: Very poor algorithm to project sub-cortical regions and cerebellum * Maximum: * Menu Sources > Maximum value: Doesn't work with volume or mixed head models * Panel Get coordinates: Add button "find maximum" * Sources on surface: Display peak regions over time (time = color) => A.Gramfort * BEM single sphere: Get implementation from MNE * Volume grid: * Optimize: 3D display (better than 9x9 cubes) * Optimize: vol_dilate (with 26 neighbors) * Add eyes models to attract eye activity * Display spectrum scouts (PSD plots when clicking on "Display scouts" on PSD/full cortex) |
* requested feature from the forum: * * https://neuroimage.usc.edu/forums/t/event-export-and-process-find-maximum-value-amplitude/41911/2 * * https://neuroimage.usc.edu/forums/t/custom-process-that-involves-merging-of-channels/40638 * * https://neuroimage.usc.edu/forums/t/swloreta-for-source-localization/41882/4 |
| Line 165: | Line 219: |
| * ICBM152 update: * Move out default anatomy from GitHub repository: * Process with FreeSurfer 7.2 + add FS atlases (Brainnettome, Schaeffer, HCP...) * Add volume atlases (+ reimport ASEG as volatlas) * Add facemask => Use for defacing with any MNI registration * Add T2? * BEM surfaces: Deform fieldtrip BEM surfaces from ICBM152 to subject space with MNI coordinates? |
* Display parcellation values (matrices) in 3D and 2D. * https://github.com/dutchconnectomelab/Simple-Brain-Plot * Scouts * Import SimNIBS4: Use final_tissues_LUT.txt instead of fixed list of tissues: https://neuroimage.usc.edu/forums/t/removing-a-lesioned-area/38414/20 |
| Line 173: | Line 226: |
| * MRI segmentation: <<BR>> * SimNIBS: Replace HEADRECO with CHARM (headreco will be removed in SimNIBS 4) |
|
| Line 180: | Line 231: |
| * Import from SimNIBS (Conform2MNI_nonl.nii.gz, MNI2Conform_nonl.nii.gz) |
|
| Line 181: | Line 234: |
| * Adjust CT contrast better: https://neuroimage.usc.edu/forums/t/automatic-localization-of-seeg-electrodes/36302/10 |
|
| Line 186: | Line 241: |
| Line 187: | Line 243: |
| Line 189: | Line 246: |
| * Removing MNI face mask using MNI coordinates * Atlas switch in 3D MRI figures |
* Removing MNI face mask using MNI coordinates (mask available ICMB152 2023b) |
| Line 192: | Line 250: |
| Line 197: | Line 256: |
* Brain2mesh: Add import of 10-10 positions |
|
| Line 199: | Line 260: |
| Line 202: | Line 264: |
| Line 203: | Line 266: |
| Line 204: | Line 268: |
| Line 205: | Line 270: |
* https://openneuro.org/datasets/ds000256/versions/00002 * https://osf.io/axz5r/ |
|
| Line 208: | Line 277: |
| * Project from one hemisphere to the other using registered spheres/squares (http://neuroimage.usc.edu/forums/t/how-to-create-mirror-roi-in-the-other-hemisphere/5910/8) | |
| Line 212: | Line 281: |
| == ECOG/SEEG == * Contact positions: Import / set / detect * New option: Align on none|inner|cortex to replace ECOG-mid * Project contact positions across subjects or templates (Marcel)<<BR>>Right-click on channel file > Project on default anatomy * Create clusters from anatomical labels: * Identify contacts in a given anatomical region (volume scout, surface mesh, or label in a volume atlas) / allow extracting the signals from all the contacts in an ROI * Automatic segmentation of CT: * GARDEL: http://meg.univ-amu.fr/wiki/GARDEL:presentation * SEEG DEETO Arnulfo 2015: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6 * Used routinely at Niguarda Hospital + other hospitals worldwide, reliable tool. * To be used with SEEG-assistant/3DSlicer: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1545-8 * ECOG Centracchio 2021: https://link.springer.com/content/pdf/10.1007/s11548-021-02325-0.pdf * Classifier on thresholded CT: https://github.com/Jcentracchio/Automated-localization-of-ECoG-electrodes-in-CT-volumes * SEEG Granados 2018 (no code shared): https://link.springer.com/content/pdf/10.1007/s11548-018-1740-8.pdf * ECOG: * Project and display contacts on cortex surface should consider the rigidity of the grids: Contacts cannot rotate, and distance between contacts should remain constant across runs * Method for contacts projection: https://pdfs.semanticscholar.org/f10d/6b899d851f3c4b115404298d7b997cf1d5ab.pdf * ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage) <<BR>>Normalization MNI? solutions with FieldTrip? * Display: * Bad channels: Contacts greyed out instead of ignored (Marcel) * Display time in H:M:S * Display curved SEEG electrodes * Detection CEEP stim artifacts: Use ImaGIN code ImaGIN_StimDetect |
* Improving the registration between EEG and anatomy templates: * Warping: Improve the basic alignment of the digitized electrodes on the templat, possibly with Cz and other anatomical landmarks * EEG template positions: rework using a standardized Cz position (+ other landmarks) == Forward modeling == * DUNEuro/FEM: * Add lesion mask to SimNIBS: https://simnibs.github.io/simnibs/build/html/documentation/command_line/add_tissues_to_upsampled.html#add-tissues-to-upsampled-doc * GeomtryAdapted: Buggy? * Display differences between leadfields: amplitude of difference (right-click > Compare) * Display sensitivity on FEM surface * OpenMEEG: Detect bad results + exclude from leadfield * BEM single sphere: Get implementation from MNE-Python (John Mosher) * Add eyes models to attract eye activity (Put a dipole in each eye) == Source modeling == * Reproduce results in "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x * eLORETA instead of sLORETA? * https://neuroimage.usc.edu/forums/t/compute-eeg-sources-with-sloreta/13425/6 * https://neuroimage.usc.edu/forums/t/loreta-and-source-localization/30525 * "eLORETA algorithm is available in the MEG/EEG Toolbox of Hamburg (METH)": https://www.biorxiv.org/content/biorxiv/early/2019/10/17/809285.full.pdf * https://github.com/brainstorm-tools/brainstorm3/issues/114 * Point-spread functions (PSFs) and cross-talk functions: https://mne.tools/stable/auto_examples/inverse/plot_psf_ctf_vertices.html#sphx-glr-auto-examples-inverse-plot-psf-ctf-vertices-py https://www.biorxiv.org/content/biorxiv/early/2019/06/18/672956.full.pdf * https://github.com/olafhauk/EEGMEGResolutionAtlas * Dipoles: * Display dipoles in MRI viewer * panel_dipoles: Doesn't work with multiple figures (SOLVED?) * Project sources: Very poor algorithm to project sub-cortical regions and cerebellum * Maximum: * Menu Sources > Maximum value: Doesn't work with volume or mixed head models * Panel Get coordinates: Add button "find maximum" * Sources on surface: Display peak regions over time (time = color) => A.Gramfort * Volume grid: * Optimize: 3D display (better than 3x3 cubes) * Optimize: vol_dilate (with 26 neighbors) |
| Line 237: | Line 331: |
| * Stat on connectivity? | |
| Line 240: | Line 333: |
| Line 245: | Line 339: |
| Line 249: | Line 344: |
| * Add option to process to specify the protocol name * Disable logging of sub-processes (reloading the previous report should only show process_import_bids) |
|
| Line 252: | Line 349: |
| Line 253: | Line 351: |
| Line 254: | Line 353: |
| * Read associated empty room: * https://github.com/brainstorm-tools/brainstorm3/issues/138 |
* Use BIDS-Matlab? * Test datasets: * See list of test datasets in process_import_bids.m * ds004085 / ds004473: Check response epoch + BUG with coordinate interpretation |
| Line 257: | Line 361: |
| * Add events tsv, channel tsv, EEG, iEEG * BIDS-Matlab? |
* EEG, iEEG: Add events.tsv, channel.tsv, electrodes.tsv * Anatomy: Add t1w.json (including fiducials) * Use BIDS-Matlab? * EDF+ reader: Add resampling of channels with different sampling rates |
| Line 260: | Line 368: |
| Line 263: | Line 372: |
| Line 264: | Line 374: |
| * EEG File formats: * XDF: https://github.com/sccn/xdf |
* EEG File formats:<<BR>> |
| Line 267: | Line 376: |
| Line 268: | Line 378: |
| Line 269: | Line 380: |
| Line 270: | Line 382: |
| Line 271: | Line 384: |
| Line 278: | Line 392: |
| Line 279: | Line 394: |
| * MINC MRI: Add support for "voxel to world" transformation (vox2ras) similarly to .nii | |
| Line 281: | Line 397: |
| * Java-free Matlab: All references of functions below must be removed * '''JavaFrame''': screencapture.m (used for screen captures of videos) * '''Actxcontrol''': Used for video-EEG * uihtml + JavaScript callbacks? * ActiveX in .NET app? * Pure Java framce + VLC java plugin? * Other video player? * '''Javacomponent''': * mri_editMask * figure_mri * process_bandpass * List .jar files used from Matlab distribution (e.g. dom) => Check all the import calls |
|
| Line 283: | Line 416: |
| * Remove ICBM152 default anatomy from repo | |
| Line 292: | Line 425: |
| * easyh5 | |
| Line 298: | Line 430: |
* MNE-Python 1.0: Test and update install documentation |
|
| Line 299: | Line 433: |
| * Update the organization of derivatives folder (same for ECOG tutorial) * Add review of literature for the resting state MEG |
* Update the organization of derivatives folder (full FS folders) |
| Line 302: | Line 435: |
| Line 304: | Line 438: |
| Line 305: | Line 440: |
| * FieldTrip ECOG tutorial: http://www.fieldtriptoolbox.org/tutorial/human_ecog * Reproduce tutorials from MNE-Python: https://martinos.org/mne/stable/tutorials.html |
|
| Line 308: | Line 442: |
| Line 310: | Line 445: |
| Line 311: | Line 447: |
* https://archive.physionet.org/ [data and tools] |
|
| Line 316: | Line 455: |
| Line 319: | Line 459: |
| Line 320: | Line 461: |
| * Doesn't close properly on some Linux systems * Focus requests change workspace when processing constantly (Linux systems) |
* Doesn't close properly on some Linux systems (SOLVED?) * Focus requests change workspace when processing constantly (Linux systems) (SOLVED?) |
| Line 326: | Line 467: |
| Line 327: | Line 469: |
| Line 328: | Line 471: |
| Line 329: | Line 473: |
| Line 330: | Line 475: |
| Line 336: | Line 482: |
| Line 337: | Line 484: |
| * Interpolations: Use scatteredInterpolant, griddedInterpolant, triangulation.nearestNeighbor (2014b) |
What's next
A roadmap to the future developments of Brainstorm.
Contents
Recordings
- Review signals in time-frequency space
- Events processes: Select events names from a list instead of having to type them
- Display CTF coils: Show discs instead of squares
- Sleep scoring wish list (Emily C):
- Configurable horizontal lines (for helping detecting visually some thresholds)
- Mouse ruler: Measure amplitude by dragging the mouse.
- Automatic spindle detector
https://neuroimage.usc.edu/forums/t/page-overlap-while-reviewing-raw-file-a-way-to-set-to-0/11229/13
RAW file viewer speed (Low priority) :
- Consider to change to a format that is faster to read
- Add parameter to make the visual downsampling more or less aggressive
Keep the filter specifications in memory instead of recomputing for every page
(Nice to have)
BioSemi: Add menu "Convert naming system" to rename channels into 10-10 (A1=>FPz)
Simulations: https://github.com/lrkrol/SEREEGA(Low priority)
ECOG/SEEG
https://www.sciencedirect.com/science/article/pii/S1053811922005559
- Display (high-priority)(Part SEEG grant):
- Group display: Overlay multiple channel files in the same figure, coloring contacts by subject/ROI/Cluster/Electrode name
- iEEG tab must be read-only when multiple files (hide configuration controls)
Bad channels: Contacts greyed out instead of ignored (Marcel H, Germany)
(To diff between band and not-recorded) > Rendering of SEEG electrodes: Full surface modelling with surface mesh (see Lead-DBS models + code that generates them?)Display time in H:M:S instead of S > If there is t0 in H:M:S instead of S > As an option in Display configuration button>x-axis
- view_leadfield_sensitivity: Add closing surfaces at cortex limits
- Create clusters from anatomical labels (Anne So) :
Identify contacts in a given anatomical region (volume scout, surface mesh, or label in a volume atlas) / allow extracting the signals from all the contacts in an ROI> As a process to select recordings, then Scouts from Volumen Atlas, Create cluster in channel file, then Extract time series.
- Group analysis: extract clusters across subjects, display or average signals (see MIA) (Anne So)
- Group display: Overlay multiple channel files in the same figure, coloring contacts by subject/ROI/Cluster/Electrode name
- Spike detection (Need to check for current toolboxes from scratch)(contact Nicolas R)(Mosher J)(iEEG BIDS):
- Automatic segmentation of CT:
- Discussed with Samuel and Christian (ins-amu.fr)
SEEG DEETO Arnulfo 2015: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-015-0511-6
- Used routinely at Niguarda Hospital + other hospitals worldwide, reliable tool.
To be used with SEEG-assistant/3DSlicer: https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1545-8
ECOG Centracchio 2021: https://link.springer.com/content/pdf/10.1007/s11548-021-02325-0.pdf
Classifier on thresholded CT: https://github.com/Jcentracchio/Automated-localization-of-ECoG-electrodes-in-CT-volumes
SEEG Granados 2018 (no code shared): https://link.springer.com/content/pdf/10.1007/s11548-018-1740-8.pdf
- ECOG:
- Project and display contacts on cortex surface should consider the rigidity of the grids: Contacts cannot rotate, and distance between contacts should remain constant across runs
Method for contacts projection: https://pdfs.semanticscholar.org/f10d/6b899d851f3c4b115404298d7b997cf1d5ab.pdf
ECOG: Brain shift: When creating contact positions on a post-implantation image, the brain shift should be taken into account for creating images of the ECOG contacts on the pre-op brain => iELVis (http://ielvis.pbworks.com/w/page/116347253/FrontPage)
Normalization MNI? solutions with FieldTrip?
Display CT images: Better brightness/contrast adjustment: https://neuroimage.usc.edu/forums/t/automatic-localization-of-seeg-electrodes/36302/8 Range of values is way diff than ones from MRI. Current color maps are not suitable for CT, need to be improved.Together with processing of CT to get electrode positions.
Detection CCEP stim artifacts: Use ImaGIN code ImaGIN_StimDetect https://f-tract.eu/software/imagin/
Pre-processing
- process_detectbad:
- Allow on raw files (for bad channels only)
- Add detection on derivative of the signal (see EEGLAB)
- Document in tutorial Bad channels
- PREP pipeline / EEGLAB (Bigdely-Shamlo 2015)
- Improve bad channel/trial detection:
- ft_artifact_threshold and ft_rejectartifact
- MNE-Python
- EEGLAB
- Integrate with EYE-EEG (Olaf Dimigen)
Reproduce tutorial: https://www.eyetracking-eeg.org/tutorial.html
- Create EYE-EEG plugin + processes (Raphael Lambert)
- Process: Detect sacades (extended events) + fixations
- Improved ICA
- Eye-movement related potentials
Use it to guide ICA: http://www2.hu-berlin.de/eyetracking-eeg/
ICA:
Automatic classification: ICLabel: https://neuroimage.usc.edu/forums/t/automatic-eeg-ic-ica-classification-for-brainstorm/33785
- Exploration: Add window with spectral decomposition (useful for muscle artifacts)
- Export IC time series (and then compute their spectrum): solves the problem above
- Import ICA matrices available in EEGLAB .set files
ICA+machine learning: https://www.ncbi.nlm.nih.gov/pubmed/28497769
Automated artifact rejection: https://arxiv.org/abs/1612.08194
- Spectral representation of ICs
- SSP:
Display warning if changing the ChannelFlag while there is a Projector applied
- File format:
- Add support to read GDF file format
https://github.com/donnchadh/biosig/blob/master/biosig/t200_FileAccess/sload.m
Reproducibility toolbox
- Generate fully reproducible scripts, including all the interactive/graphical parts
- Record all GUI actions as script calls
- Import window: Add button to create the corresponding processing pipeline (to generate script or to edit additional options).
- Adding the list of plugins to the reports
- Better provenance: History fields, uniform file names...
Interface
Add a warning when computing a forward model with > 100000 sources (check selection)
- Colormaps: Global colormap max: Should get the maximum across all the open files
- Snapshot:
- Save as image / all figures (similar to Movie/all figure)
Contact sheets & movies: use average of time windows instead of single instants, for each picture.
- Contact sheets: Allow explicit list of times in input (+ display as in MNE-Python with TS)
Database
- Save iHeadModel somewhere in the datbase structure
Generalize the use of the units (field .DisplayUnits): Save in source files
Connectivity
- Define names and unit labels for each connectivity metric
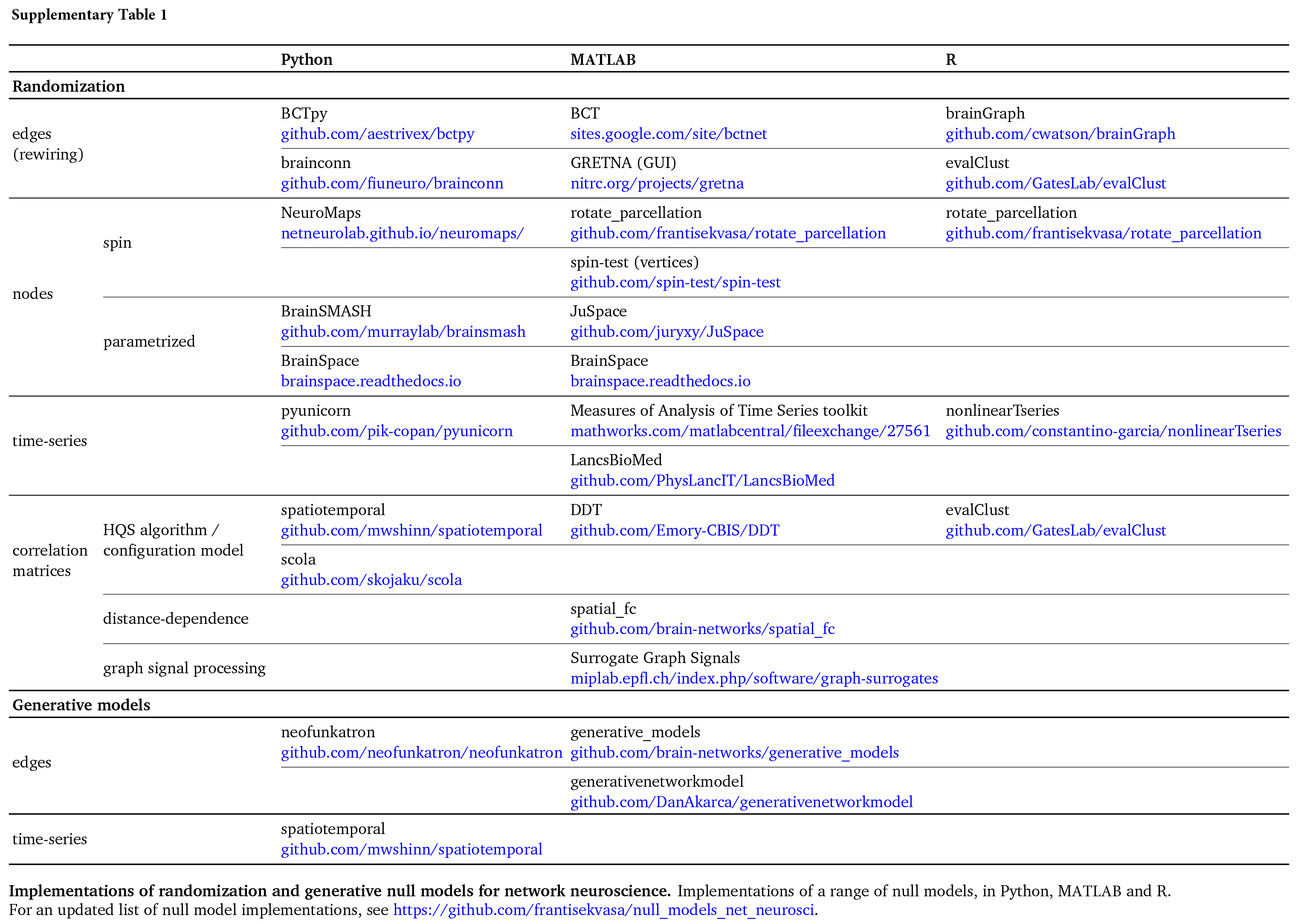
Null models: (Bratislav M) https://www.nature.com/articles/s41583-022-00601-9
- Connect NxN display:
- Graph on sensors: Place M/EEG sensors by location, not by channel order
- Display as image: Add legend of the elements along X and Y axis
- Display as time series: Display warning before trying to open too many signals
- Optimize display: use surface() instead of line() for links? (as in figure_3d/PlotFibers)
- Time-resolved correlation/coherence: Display as time bands (as done in wavelet, to have same time axis as data)
Processes
- Add MNE-Python functions:
- scikit-learn classifiers
- BEM single layer (John wants to test it)
https://neuroimage.usc.edu/forums/t/best-way-to-export-to-mne-python/12704/3
Reproduce tutorials / examples from FieldTrip and MNE-Python:
FieldTrip ECOG tutorial: http://www.fieldtriptoolbox.org/tutorial/human_ecog
Reproduce tutorials from MNE-Python: https://martinos.org/mne/stable/tutorials.html
Spatial resolution metrics in source space:
https://mne.tools/stable/auto_examples/inverse/plot_resolution_metrics.html#sphx-glr-auto-examples-inverse-plot-resolution-metrics-py- Change the graphic renderer from Matlab
Chronux toolbox : http://chronux.org/
Add FieldTrip functions:
- ft_sourceanalysis:
- Check noise covariance
- Check all the options of all the methods
- Single trial reconstructions + noise covariance?
Filters?? http://www.fieldtriptoolbox.org/example/common_filters_in_beamforming
Beamformers: Save ftSource.avg.mom
http://www.fieldtriptoolbox.org/workshop/meg-uk-2015/fieldtrip-beamformer-demohttp://www.fieldtriptoolbox.org/tutorial/beamformingextended
- Baseline? Two inputs?
- ft_prepare_heamodel: Add support from BEM surfaces from the Brainstorm database
ft_volumereslice: http://www.fieldtriptoolbox.org/faq/how_change_mri_orientation_size_fov
- ft_freqanalysis
- ft_combineplanar
- ft_sourceanalysis:
- Optimization:
- Use CUDA for speeding up some operations (filtering, wavelets, etc)
- Use Matlab Coder to optimize: Wavelets, bandpass filter, sinusoid removal
- Pipeline editor:
- Bug: After "convert to continuous", the time of the following processes should change
- Add loops over subjects/conditions/trial groups
- Events: Allow selection from a drop-down list (similar to option "channelname" in panel_process_selection)
ITC: Inter-trial coherence (see MNE reports for group tutorial)
http://www.sciencedirect.com/science/article/pii/S1053811916304232Remove line noise: http://www.nitrc.org/projects/cleanline
- Time-frequency:
- Optimization: bst_timefreq (around l.136), remove evoked in source space: Average should be computed in sensor space instead of source space (requested by Dimitrios)
Short-time Fourier transform: http://www.mikexcohen.com/lectures.html
- Hilbert with time bands very slow on very long files (eg. 3600s at 1000Hz) because the time vector is still full (10^7 values): save compressed time vector instead.
- When normalizing with baseline: Propagate with the edge effects marked in TFmask
- Allow running TF on montages
- Bug when computing TF on constrained and unconstrained scouts at the same time (in mixed head models for instance): uses only the constrained information and doesn't sum the 3 orientations for the unconstrained regions.
- requested feature from the forum:
* https://neuroimage.usc.edu/forums/t/event-export-and-process-find-maximum-value-amplitude/41911/2
* https://neuroimage.usc.edu/forums/t/custom-process-that-involves-merging-of-channels/40638
* https://neuroimage.usc.edu/forums/t/swloreta-for-source-localization/41882/4
Anatomy
- Display parcellation values (matrices) in 3D and 2D.
Import SimNIBS4: Use final_tissues_LUT.txt instead of fixed list of tissues: https://neuroimage.usc.edu/forums/t/removing-a-lesioned-area/38414/20
Simple-brain-plot: https://github.com/dutchconnectomelab/Simple-Brain-Plot
- MNI normalization: More options:
- DARTEL / SHOOT
BrainSuite (wait for Anand)
- Import from SimNIBS (Conform2MNI_nonl.nii.gz, MNI2Conform_nonl.nii.gz)
- MRI Viewer:
Adjust CT contrast better: https://neuroimage.usc.edu/forums/t/automatic-localization-of-seeg-electrodes/36302/10
- Pan in zoomed view (shift + click + move?)
- Zoom in/out with mouse (shift + scroll?)
- Ruler tool to measure distances
- Display scouts as additional volumes
Render surface envelope in the MRI as a thin line instead of the full interpolation matrix
Or use inpolyhedron to get a surface mask and then erode it to get the volume envelopeSurface>Volume interpolation: Use spm_mesh_to_grid instead of tess_tri_interp
- Defacing:
https://afni.nimh.nih.gov/pub/dist/doc/htmldoc/tutorials/refacer/refacer_run.html
- Removing MNI face mask using MNI coordinates (mask available ICMB152 2023b)
Bug import anatomy: Requested nVert > high-resolution cortex surface: Creates an empty cortex_0V
BrainSuite:
- Use same colors for left and right for anatomical atlases
- Use for volume coregistration (rigid / non-rigid)
- USCBrain: Add default electrodes positions
Remove BrainSuite1 when not needed anymore
- Brain2mesh: Add import of 10-10 positions
- Templates for different ages:
MNI: https://www.bic.mni.mcgill.ca/ServicesAtlases/NIHPD-obj1
Pediatric head atlases: https://www.pedeheadmod.net/pediatric-head-atlases-v1-2/
https://www.biorxiv.org/content/biorxiv/early/2020/02/09/2020.02.07.939447.full.pdf
John Richards: https://www.nitrc.org/frs/?group_id=1361
Neurodev database: https://jerlab.sc.edu/projects/neurodevelopmental-mri-database/
- Scouts:
- Display edges in the middle of the faces instead of the vertices
- Parcellating volume grids: scikit-learn.cluster.Ward
Geodesic distance calculations:
https://www.mathworks.com/matlabcentral/fileexchange/6110-toolbox-fast-marching- Improving the registration between EEG and anatomy templates:
- Warping: Improve the basic alignment of the digitized electrodes on the templat, possibly with Cz and other anatomical landmarks
- EEG template positions: rework using a standardized Cz position (+ other landmarks)
Forward modeling
- DUNEuro/FEM:
Add lesion mask to SimNIBS: https://simnibs.github.io/simnibs/build/html/documentation/command_line/add_tissues_to_upsampled.html#add-tissues-to-upsampled-doc
GeomtryAdapted: Buggy?
Display differences between leadfields: amplitude of difference (right-click > Compare)
- Display sensitivity on FEM surface
- OpenMEEG: Detect bad results + exclude from leadfield
- BEM single sphere: Get implementation from MNE-Python (John Mosher)
- Add eyes models to attract eye activity (Put a dipole in each eye)
Source modeling
Reproduce results in "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x
- eLORETA instead of sLORETA?
https://neuroimage.usc.edu/forums/t/compute-eeg-sources-with-sloreta/13425/6
https://neuroimage.usc.edu/forums/t/loreta-and-source-localization/30525
"eLORETA algorithm is available in the MEG/EEG Toolbox of Hamburg (METH)": https://www.biorxiv.org/content/biorxiv/early/2019/10/17/809285.full.pdf
Point-spread functions (PSFs) and cross-talk functions: https://mne.tools/stable/auto_examples/inverse/plot_psf_ctf_vertices.html#sphx-glr-auto-examples-inverse-plot-psf-ctf-vertices-py https://www.biorxiv.org/content/biorxiv/early/2019/06/18/672956.full.pdf
- Dipoles:
- Display dipoles in MRI viewer
- panel_dipoles: Doesn't work with multiple figures (SOLVED?)
- Project sources: Very poor algorithm to project sub-cortical regions and cerebellum
- Maximum:
Menu Sources > Maximum value: Doesn't work with volume or mixed head models
- Panel Get coordinates: Add button "find maximum"
Sources on surface: Display peak regions over time (time = color) => A.Gramfort
- Volume grid:
- Optimize: 3D display (better than 3x3 cubes)
- Optimize: vol_dilate (with 26 neighbors)
Statistics
- Stat on unconstrained sources?
Stat/time series: Hide lines going down to zero (Dimitrios: https://neuroimage.usc.edu/forums/t/common-source-activation-across-subjects-and-conditions/1152/21)
- Cluster stat: Add frequency selection option
- ANOVA:
- Write panel similar to Process1 and Process2
- Output = 1 file per effect, all grouped in a node "ANOVA"
- Display several ANOVA maps (from several files) on one single figure, using a "graphic accumulator", towards which one can send any type of graphic object
Multivariate stim-response analysis: https://github.com/mickcrosse/mTRF-Toolbox
Input / output
- BIDS import:
- Add option to process to specify the protocol name
- Disable logging of sub-processes (reloading the previous report should only show process_import_bids)
- Full support for iEEG and EEG
- Read real fiducials (OMEGA) / transformation matrices:
- Use BIDS-Matlab?
- Test datasets:
- See list of test datasets in process_import_bids.m
- ds004085 / ds004473: Check response epoch + BUG with coordinate interpretation
- BIDS export:
- EEG, iEEG: Add events.tsv, channel.tsv, electrodes.tsv
- Anatomy: Add t1w.json (including fiducials)
- Use BIDS-Matlab?
- EDF+ reader: Add resampling of channels with different sampling rates
Support for OpenJData / JNIfTI: https://github.com/brainstorm-tools/brainstorm3/issues/284
- DICOM converter:
- Add dcm2nii (MRICron)
- Add MRIConvert
- SPM .mat/.dat: Fix the import of the EEG/SEEG coordinates
EEG File formats:
Persyst .lay: https://github.com/ieeg-portal/Persyst-Reader
Nervus .eeg: https://github.com/ieeg-portal/Nervus-Reader
Biopac .acq: https://github.com/ieeg-portal/Biopac-Reader
- BCI2000 Input (via EEGLAB plugin)
- 4D file format:
- Use reader from MNE-Python: mne.io.read_raw_kit (skip Yokogawa slow library)
- Reference gradiometers: Keep the orientation of the first or second coil?
- Reference gradiometers: Add the sensor definition from coil_def.dat
- Validate with phantom recordings that noise compensation is properly taken into account
- The noise compensation is considered to be always applied on the recordings, not sure this assumption is always correct
- 4D phantom tutorial (JM Badier?)
- BST-BIN: Add compression to .bst
- MINC MRI: Add support for "voxel to world" transformation (vox2ras) similarly to .nii
Distribution
- Java-free Matlab: All references of functions below must be removed
JavaFrame: screencapture.m (used for screen captures of videos)
Actxcontrol: Used for video-EEG
uihtml + JavaScript callbacks?
- ActiveX in .NET app?
- Pure Java framce + VLC java plugin?
- Other video player?
Javacomponent:
- mri_editMask
- figure_mri
- process_bandpass
List .jar files used from Matlab distribution (e.g. dom) => Check all the import calls
Cleanup GitHub repository:
- Move external I/O libraries as plugins:
- mne-matlab
- CEDS64ML
- edfimport
- eeprobe
- son
- ricoh
- yokogawa
Documentation
- All tutorial datasets in BIDS (including introduction tutorials)
- Deface the MRIs of all the tutorials
Count GitHub clones in the the download stats
- MNE-Python 1.0: Test and update install documentation
- Tutorial OMEGA/BIDS:
- Update the organization of derivatives folder (full FS folders)
- Download example datasets directly from the OMEGA repository
New tutorials:
Other public datasets: https://github.com/INCF/BIDS-examples/tree/bep008_meg/
- EEG/research
Cam-CAN database: https://camcan-archive.mrc-cbu.cam.ac.uk/dataaccess/<<BR>>(download new datasets, including maxfiltered files and manual fiducial placements)
- MEG steady-state / high-gamma visual / frequency tagging
Reproduce results from "Simultaneous human intracerebral stimulation and HD-EEG, ground-truth for source localization methods": https://www.nature.com/articles/s41597-020-0467-x
- Stand-alone ICA tutorial
https://archive.physionet.org/ [data and tools]
Current bugs
- Image viewer:
- Difficult to get to 100%
- Buggy on some systems
- 2DLayout:
- (TF) Units are weird with % values
- (TF) Difficult to navigate in frequencies: Scaling+changing frequency resets the scaling
- Progress bar:
- Doesn't close properly on some Linux systems (SOLVED?)
- Focus requests change workspace when processing constantly (Linux systems) (SOLVED?)
Distributed computing
Options from FieldTrip:
Loose collection of computers: https://github.com/fieldtrip/fieldtrip/tree/master/peer
Single multicore machine: https://github.com/fieldtrip/fieldtrip/tree/master/engine
Batch system: https://github.com/fieldtrip/fieldtrip/tree/master/qsub
Documentation: https://www.fieldtriptoolbox.org/faq/what_are_the_different_approaches_i_can_take_for_distributed_computing/
Geeky programming details
- Replace all calls to inpolyhd.m with inpolyhedron.m (10x faster)
Interface scaling: Rewrite class IconLoader to scale only once the icons at startup instead of at each request of an icon (might improve the speed of the rendering of the tree)
- Processes with "radio" and "radio_line" options: Replace with "radio_label" and "radio_linelabel"