Tutorial 11: Artifact detection
Authors: Francois Tadel, Elizabeth Bock, John C Mosher, Sylvain Baillet
The previous tutorial illustrated how to remove noise patterns occurring continuously and at specific frequencies. However, most of the events that contaminate the MEG/EEG recordings are not permanent, span over a large frequency range or overlap with the frequencies of the brain signals of interest. Frequency filters are not appropriate to correct for eye movements, breathing movements, heartbeats or other muscle activity.
For getting rid of reproducible artifacts, one popular approach is the Signal-Space Projection (SSP). This method is based on the spatial decomposition of the MEG/EEG recordings for a selection of time samples during which the artifact is present. Therefore we need to identify when each type of artifact is occurring in the recordings. This tutorial shows how to detect automatically some well defined artifacts: the blinks and the heartbeats.
Contents
Observation
Let's start by observing the type of contamination the blinks and heartbeats cause to the MEG recordings.
Run #01: Double-click on the link to show the MEG sensors.
Configuration: Page of 3 seconds, view in columns, selection of the "CTF LT" sensors (the left-temporal sensors will be a good example to show at the same time the two types of artifacts).
EOG: Right-click on the link > EOG > Display time series. Two channels are classified as EOG:
VEOG: Vertical electrooculogram (two electrodes placed below and above one eye)
HEOG: Horizontal electrooculogram (two electrodes placed on the temples of the subject)
- On those traces, there is not much happening for most of the recordings except for a few bumps. This subject is sitting very still and not blinking much. We can expect MEG recordings of a very good quality.
ECG: Right-click on the link > ECG > Display time series.
The electrocardiogram was recorded with a bipolar montage of electrodes across the chest. You can recognize the typical shape of the electric activity of the heart (P, QRS and T waves).Find a blink: Scroll through the recordings using the F3 shortcut until you see a large blink.
- Remember you can change the amplitude scale with many shortcuts (eg. right-click + move).
- To keep the scale fixed between to pages: Uncheck the button [AS] in the figure (auto-scale)
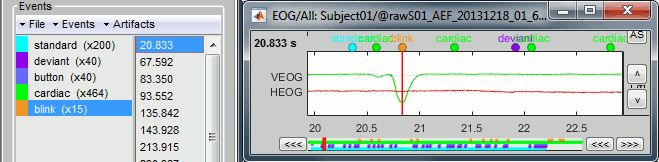
- For instance, you can observe a nice blink at 20.8s (red cursor in the screen capture below).
- On the same page, you should be able to observe the contamination due to a few heartbeats, corresponding to the peaks of the ECG signal (eg. 19.8s, shown as a blue selection below).
The additional data channels (ECG and EOG) contain precious information that we will use for the automatic detection of the blinks and heartbeats. We strongly recommend that you always record these signals in your own experiments, it helps a lot with the data analysis.
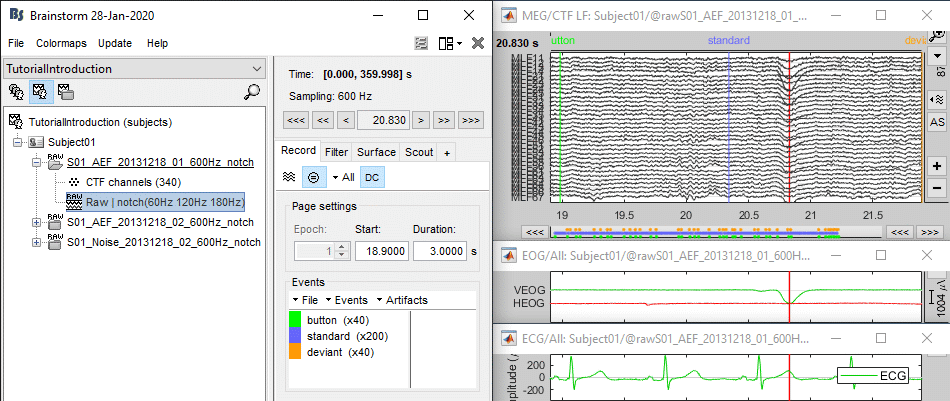
Detection: Heartbeats
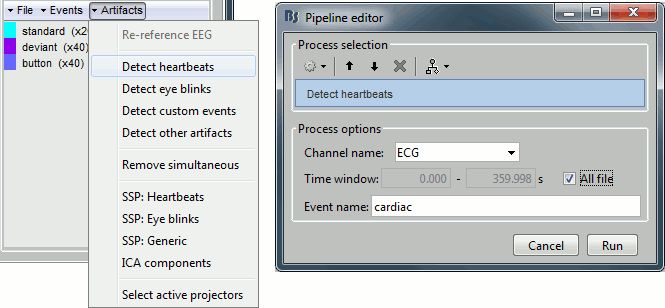
In the Record tab, select the menu: Artifacts > Detect heartbeats.
- It opens automatically the pipeline editor, with the process "Detect heartbeats" selected.
Channel name: Name of the channel that is used to perform the detection. Select "ECG".
Time window: Time range that the algorithm should scan for amplitude peaks. Leave the default values to process the entire file, or select the option [All file].
Event name: Name of the event group created for saving the detected events. Keep "cardiac".
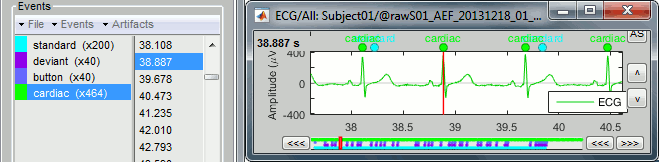
Click on Run. After the process stops, you can see a new event category "cardiac". The 464 occurrences indicate a heart rate around 77bpm.
You can check a few of them, to make sure the "cardiac" markers really indicate the ECG peaks, and that there are not too many peaks that are skipped.
Detection: Blinks
Now do the same thing for the blinks: Menu Artifacts > Detect eye blinks.
Channel name: VEOG
Time window: All file
Event name: Blink
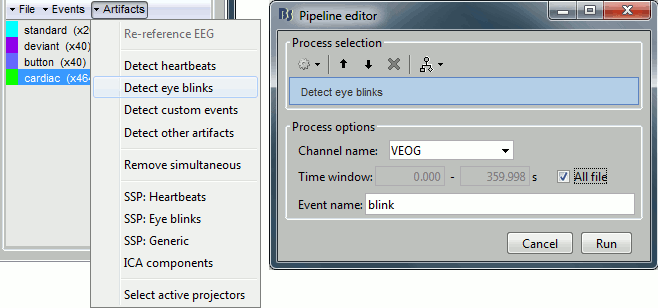
Run, observer the 15 detected blinks.
Remove simultaneous blinks/heartbeats
We will use these event markers as the input to our SSP cleaning method. This technique works well if each artifact is defined as precisely as possible, and as independently as possible from the other artifacts. This means that we should try to avoid having two different artifacts marked at the same time.
Because the heart beats every second or so, there is a high chance that when the subject blinks there is a heartbeat not too far away in the recordings. Because the amplitude of the artifact caused by the heart is negligible compared with amplitudes caused by the eyes, the events "blink" would completely mask the events "cardiac". Cleaning for the heartbeats would also remove a bit of the blinks, causing then the blink cleaning to perform poorly. To avoid this situation, we can remove the markers "cardiac" that are occurring during a blink.
In the Record tab, select the menu "SSP > Remove simultaneous". Set the options:
Remove events named: "cardiac"
When too close to events: "blink"
Minimum delay between events: 250ms
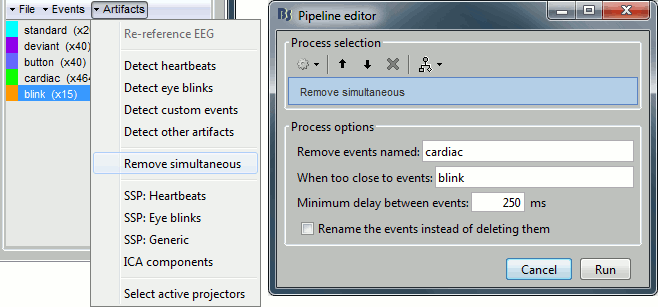
After executing this process, the number of "cardiac" events goes from 464 to 455. The deleted heartbeats were all less than 250ms away from a blink.
Running from a script
Let's perform the same detection operations on Run #02, using this time the Process1 box.
- Close everything with button [X] on the top-right corner of the Brainstorm window.
- Select the run AEF #02 in the Process1 box, select successively:
Events > Detect heartbeats: Select channel ECG, check "All file", event name "cardiac".
Events > Detect eye blinks: Select channel VEOG, check "All file", event name "blink".
Events > Remove simultaneous: Remove "cardiac", too close to "blink", delay 250ms.
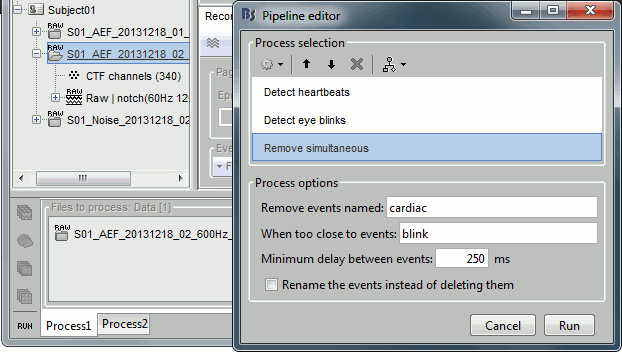
- Open the Run#02 recordings (MEG+EOG+ECG) and verify that the detection gave correct results.
Custom detection
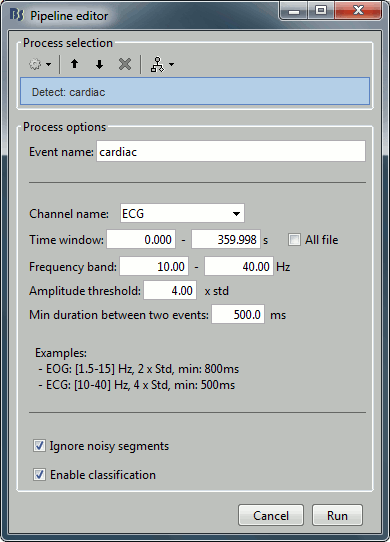
Those two processes "Detect heartbeats" and "Detect eye blinks" are in reality shortcuts for a generic process "Detect custom events". This process can be used for detecting any kind of event based on the signal power in a specific frequency band. We are not going to use it here, but it is interesting to introduce it to understand how the blinks and heartbeats detection work. The logic is the following:
- The channel to analyze is read from the continuous file, for a given time window.
Frequency band: The signal is filtered in a frequency band where the artifact is easy to detect. For EOG: 1.5-15Hz ; for ECG: 10-40Hz.
Threshold: An event of interest is detected if the absolute value of the filtered signal value goes over a given number of times the standard deviation. For EOG: 2xStd, for ECG: 4xStd
Minimum duration between two events: If the filtered signal crosses the threshold several times in relation with the same artifact (like it would be the case for muscular activity recordings on an EMG channel), we don't want to trigger several events but just one at the beginning of the activity. This parameter would indicate the algorithm to take only the maximum value over the given time window; it also prevents from detecting other events immediately after a successful detection. For the ECG, this value is set to 500ms, because it is very unlikely that the heart rate of the subject goes over 120 beats per minute.
Ignore the noisy segments: If this option is selected, the detection is not performed on the segments that are much noisier than the rest of the recordings.
Enable classification: If this option is selected, the events are classified by shape in different categories, based on correlation measure. In the end, only the categories that have more than 5 occurrences are kept, all the other successful detections are ignored.
In case of failure
If you cannot get your artifacts to be detected automatically, you can browse through the recordings and mark all the artifacts manually, as explained in the tutorial Event markers.
Additional documentation
Related tutorial: Detect subject movements
