|
Size: 12194
Comment:
|
Size: 15292
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Tutorial 8: Artifacts & Frequency filters = | = Tutorial 10: Power spectrum and frequency filters = |
| Line 3: | Line 3: |
We are now going to process our continuous recordings to remove the main sources of noise. Typically, we can expect contaminations coming from the environment (power lines, stimulation equipment, building vibrations) and from the subject (movements, blinks, heartbeats, breathing, teeth clenching, muscle tension, metal in the mouth or the body). In this tutorial, we will focus first on the noise patterns that occur continuously, at specific frequencies. We can correct for these artifacts using '''frequency filters'''. Usually we prefer to run these notch filters, band-pass filters and sinusoid removals '''before any other type of correction''', on the continuous files. They can be applied to the recordings without much supervision, but they may create important artifacts at the beginning and the end of the signals. Processing the entire continuous recordings at once instead of the imported epochs avoids adding these edge effects to all the trials. |
|
| Line 6: | Line 10: |
| = From Auditory = == Detect and remove artifacts == |
== Evaluation of the noise level == Before running any type of cleaning procedure on MEG/EEG recordings, we always recommend to start with a quick evaluation of the noise level. An easy way to do this is to estimate the power spectrum of all the signals over the entire recordings. |
| Line 9: | Line 13: |
| === Spectral evaluation === * One of the typical pre-processing steps consist in getting rid of the contamination due to the power lines (50 Hz or 60Hz). Let's start with the spectral evaluation of this file. * Drag '''ALL''' the "Link to raw file" to the Process1 box, or easier, just drag the node "Subject01", it will select recursively all the files in it. * Run the process "'''Frequency > Power spectrum density (Welch)'''": * Time window: '''[All file]''', Window length: '''4s''', Overlap: '''50%''', Sensor types: '''MEG''' * Note that you need at least 8Gb of RAM to run the PSD on the entire file. If you don't or if you get "Out of memory" errors, you can try running the PSD on a shorter time window. |
* Clear the list of files in the Process1 tab. * Select the three datasets we have linked to our protocol. <<BR>>You can select the three "link to raw file" nodes, the three folders or the entire subject node. <<BR>><<BR>> {{attachment:psd_select_files.gif||height="121",width="400"}} * Click on [Run] to open the pipeline editor window. * Select the process "'''Frequency > Power spectrum density (Welch)'''" * This process evaluates the power of the MEG/EEG signals at different frequencies, using the Welch's method (see [[http://en.wikipedia.org/wiki/Welch's_method|Wikipedia]] or [[http://www.mathworks.com/help/signal/ref/pwelch.html|MathWorks]]). It splits the signals in overlapping windows of a given length, calculates the Fourier Transform (FFT) of each of these short segments, and averages the power of the FFT coefficients for all the overlapping windows. <<BR>><<BR>> {{attachment:psd_options.gif||height="388",width="607"}} * Set the options as follows (click on [Edit] for the additional options): * '''Time window''': [All file] <<BR>>Portion of the input file you want to use for estimating the spectrum. * '''Window length''': 4 seconds <<BR>>Estimator length = length of the overlapping time windows for which we calculate the FFT. The number of time samples in the estimator is the same as the number of frequency bins in the output file. Increasing this parameter increases the output frequency resolution (distance between two frequency bins) but degrades the stability of the estimator, as it also decreases the total number of averaged time windows. * '''Overlap''': 50% <<BR>>How much overlap do we want between two consecutive time windows. * '''Sensor types or names''': MEG <<BR>>Defines the list of channels (names or types) on which you want to apply the process. * '''Frequency definition''': Matlab's FFT default <<BR>>You have the option to directly use the frequency binning returned by the FFT, or run an additional step of averaging these bins in larger frequency bands. Note that you can edit freely these frequency bands. * '''Output''': Save individual PSD value. <<BR>>This option will separately estimate the PSD for each of the three files in input, and create three files in output. If you select the other option (save average), it calculates the same three files but averages them on the fly and saves only one file in the database. * Click on [Run] to start the execution of the process. * '''Troubleshooting''': If you get "Out of memory" errors, try to run this PSD estimation on a shorter time segment. For example, set the time window to [0,100s] instead of the full file. This process starts by loading all the needed recordings in memory, you might not have enough memory available on your system to fit the entire dataset. * It produces three new files, that appear as "depending" on the three datasets. The comments of the files indicate how many overlapping windows could be used to estimate the PSD. "179/4000ms" means 179 windows of 4s each (total 716s). With the 50% overlap, it sums up to a little less than 2x the file length (360s). <<BR>><<BR>> {{attachment:psd_output_files.gif||height="240",width="235"}} |
| Line 16: | Line 29: |
| * Click on '''[Edit]''' and select option "'''Save individual PSD values''' (for each trial)". | == Interpretation of the PSD == ==== File: AEF#01 ==== * Double-click on the new PSD file for run #01 to display it (or right-click > Power spectrum). <<BR>><<BR>> {{attachment:psd_display.gif}} * The power spectrum is displayed in a figure similar to the time series, but the X axis represents the frequencies. Most of the shortcuts described for the recordings are also valid here. Clicking on the white parts of the figure or using the arrow keys of the keyboard move the frequency cursor. The current frequency can also be controlled with a new slider displayed in the Brainstorm window, just below the time panel. * Each black line represent the spectrum for one channel. If you click on a channel, it is selected and highlighted in red. Click again for deselecting. Right-click on a channel to read its name. <<BR>><<BR>> {{attachment:psd_run1.gif||height="194",width="515"}} * The shape of this graph is normal, it does not indicate anything unexpected: * Peaks related with the subject's alpha rhythms: around '''10Hz''' and '''20Hz'''. * Peaks related with the power lines: '''60Hz, 120Hz''' and '''180Hz'''.<<BR>>These datasets were recorded in Canada, where the alternating powerline current is delivered at 60Hz. In Europe you would observe similar peaks at 50Hz, 100Hz and 150Hz. * Add a topography view for the same file, with one of the two methods below: * Right-click on the PSD file > '''2D Sensor cap'''. * Right-click on the spectrum figure > '''2D Sensor cap''' (shortcut: '''Ctrl+T''') * Scroll in frequencies to see the spatial distribution of each frequency bin: <<BR>><<BR>> {{attachment:psd_topo.gif||height="149",width="604"}} * We have already identified two artifacts we will need to remove: the eye movements and the 60Hz+harmonics from the power lines. |
| Line 18: | Line 43: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd1.gif|psd1.gif|height="394",width="615",class="attachment"}} * Double-click on the new PSD files to display them. * Observations for '''Run''''''01''': |
==== File: AEF#02 ==== * Open the spectrum view for the run AEF#02. <<BR>><<BR>> {{attachment:psd_run2.gif||height="154",width="389"}} * Add a 2D sensor cap view for the same file. Scroll to zoom in/out.<<BR>>To display the sensor names, right-click on the figure > Channels > Display sensors. * This spectrum looks very similar to the run #01: same alpha and power line peaks. * Additionally, we observe higher signal power between '''30Hz and 100Hz''' on many '''occipital sensors'''. This is probably related to some tension in the '''neck muscles''' due to an uncomfortable seating position in the MEG. We will see later whether these channels need to be tagged as bad or not. <<BR>><<BR>> {{attachment:psd_neck.gif||height="198",width="565"}} |
| Line 22: | Line 49: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd_eval01.gif|psd_eval01.gif|height="207",width="394",class="attachment"}} * Peaks related with the power lines: '''60Hz, 120Hz, 180Hz '''(240Hz and 300Hz could be observed as well depending on the window length used for the PSD) |
==== File: Noise recordings ==== * Open the spectrum view for the noise recordings. <<BR>><<BR>> {{attachment:psd_noise.gif||height="139",width="346"}} * This shows the power spectrum of the signals that are recorded when there is no subject in the MEG room. It gives a good and simple representation of the instrumental noise. If you had one bad MEG sensor, you would see it immediately in this graph. Here everything looks good. |
| Line 25: | Line 53: |
| * The drop after '''600Hz''' corresponds to the low-pass filter applied at the acquisition time. * One channel indicates a higher noise than the others in high frequencies: '''MLO52''' (in red). We will probably mark it as bad later, when reviewing the recordings. * Observations for '''Run02''': |
== Notch filter == Let's start with the cleaning for the 60Hz+harmonics. Notch filters are adapted for removing well identified contaminations from systems oscillating at very stable frequencies. |
| Line 30: | Line 56: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd_eval02.gif|psd_eval02.gif|height="208",width="394",class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd_eval02_zoom.gif|psd_eval02_zoom.gif|height="208",width="595",class="attachment"}} * Same peaks related with the power lines: '''60Hz, 120Hz, 180Hz''' * Same drop after '''600Hz'''. * Same noisy channel: '''MLO52'''. * Additionally, we observe higher level of noise in frequencies in the range of 30Hz to 100Hz on '''many occipital sensors'''. This is probably due to some tension in the neck due to an uncomfortable position. We will see later whether those channels need to be tagged as bad. === Power line contamination === * Put '''ALL''' the "Link to raw file" into the Process1 box (or directly the Subject01 folder) * Run the process: '''Pre-process > Notch filter''' |
* Keep all the three datasets selected in the Process1 box. <<BR>>All the filters that are applied to the experiment data also need to be applied to the noise recordings. In the source estimation process, we will need all the files to have similar levels of noise. * Click on [Run] to open the Pipeline editor. * Run the process: '''Pre-process > Notch filter''' <<BR>><<BR>> {{attachment:notch_process.gif||height="281",width="343"}} * Process the entire file at once: '''NO''' |
| Line 42: | Line 62: |
| * The higher harmonics are too high to bother us in this analysis, plus they are not clearly visible in all the recordings. * In output, this process creates new .ds folders in the same folder as the original files, and links the new files to the database. |
* The higher harmonics (240Hz) are not clearly visible, and too high to bother us in this analysis. |
| Line 45: | Line 64: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd3.gif|psd3.gif|class="attachment"}} * Run again the PSD process "'''Frequency > Power spectrum density (Welch)'''" on those new files, with the same parameters, to evaluate the quality of the correction. * Double-click on the new PSD files to open them. |
* This process creates three new datasets, with additional "notch" tags. These output files are saved directly in the Brainstorm database, in a binary format specific to Brainstorm (.bst). <<BR>><<BR>> {{attachment:notch_output.gif||height="235",width="248"}} |
| Line 49: | Line 66: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd5.gif|psd5.gif|height="177",width="416",class="attachment"}} * Zoom in with the mouse wheel to observe what is happening around 60Hz (before / after). |
* If you delete the folders corresponding to the original files (before the filter), your original recordings in the .ds folders are not altered. If you delete the folders corresponding to the filtered files, you will lose your filtered recordings in .bst format. |
| Line 52: | Line 68: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd6.gif|psd6.gif|height="143",width="453",class="attachment"}} * To avoid the confusion later, delete the links to the original files: Select the folders containing the original unfiltered files and press the Delete key (or right-click > File > Delete). |
* To check where the file corresponding to a "Link to raw file" is actually saved on the hard drive, right-click on it > '''File > Show in file explorer'''. |
| Line 55: | Line 70: |
| {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=psd7.gif|psd7.gif|height="184",width="356",class="attachment"}} | == Evaluation of the filter == * Right-click on the Process1 list > '''Clear list'''. * Drag and drop the three filtered files in Process1. <<BR>><<BR>> {{attachment:notch_psd.gif||height="134",width="361"}} |
| Line 57: | Line 74: |
| = From Continuous = It is common to have portions of recordings contaminated by events coming from the subject (eye blinks, movements, heartbeats, teeth clenching, implanted stimulators...) or from the environment (stimulation equipment, elevators, cars, trains, building vibrations...). Some occur at specific frequencies and can be removed using '''frequency filters''', as introduced in the first chapter of this tutorial. Some other artifacts have more complex frequency patterns but are well defined, reproducible and can be removed efficiently using '''Signal Space Projections''' ('''SSP'''). This tutorial shows how to apply this technique to correct for the cardiac and ocular artifacts. |
* Run again the PSD process "'''Frequency > Power spectrum density (Welch)'''" on the new files, with the same parameters as before, to evaluate the quality of the correction. * Double-click on the new PSD files to open them. <<BR>><<BR>> {{attachment:notch_evaluation.gif||height="300",width="282"}} * Scroll to zoom in and observe what is happening around 60Hz (before / after). <<BR>><<BR>> {{attachment:notch_zoom.gif||height="151",width="309"}} |
| Line 60: | Line 78: |
| We are going to use the protocol '''TutorialRaw''' created in the previous tutorial [[Tutorials/TutRawViewer|Review continuous recordings and edit markers]]. If you have not followed this tutorial yet, please do it now. | == Some cleaning == To avoid the confusion later, delete the links to the original files: |
| Line 62: | Line 81: |
| == Power line contamination == Notch filters are adapted to remove some well identified contaminations from oscillating systems, such as the power lines 50Hz or 60Hz. Here is an example of how to evaluate and correct fixed frequency artifacts. Important notes: |
* Select the folders containing the original files and press '''Delete''' (or right-click > '''File > Delete''').<<BR>><<BR>> {{attachment:notch_delete.gif||height="194",width="405"}} * Always read the confirmation messages carefully, you will avoid some bad surprises. <<BR>><<BR>> {{attachment:notch_confirmation.gif}} * This is what your database explorer should look like at the end of this tutorial: <<BR>><<BR>> {{attachment:notch_final.gif||height="171",width="296"}} |
| Line 65: | Line 85: |
| 1. Usually you would prefer to apply the frequency filters '''before '''the SSP correction. 1. This approach is limited to CTF and Elekta-Neuromag recordings for now, other file formats will be developed on demand. An alternative approach for other file formats is described in the [[Tutorials/Epilepsy|EEG/Epilepsy tutorial]]. |
<<TAG(Advanced)>> |
| Line 68: | Line 87: |
| === Identify the unwanted frequencies === Drag and drop the "Link to raw file" in the Process1 tab, click on the button [Run] to open the Pipeline editor window and select the process "'''Frequency > Power spectrum density (Welch)'''". This will estimate the power spectrum of the signal using the Welch method. Set the time window to process to '''[0,50]s''', and the window length to '''4 sec''', with an overlap of '''50%''', this will be more than enough to get a good estimate of the spectrum (average of the Fourier transform of 24 windows of 4800 samples each). Leave the other options to the default values. |
== Alternatives to the notch == If the notch filter is not giving satisfying result, you can use two other processes. |
| Line 71: | Line 90: |
| {{attachment:psdRun.gif||height="290",width="463"}} Double click on the "Power (MEG)" file that was created under the "Link to raw file" in the database explorer. This shows the estimate of the power spectrum for the first 50 seconds of the continuous file, for all the sensors, with a logarithmic scale. You can identify four peaks at the following frequencies: 60Hz, 120Hz, 180Hz and 300Hz. The first three are related with the power lines (acquisition in Canada, where the electricity is delivered at 60Hz, plus the harmonics), and are expected to be coming from pure sinusoidal components in the signal. The last one is an artifact of the low-pass filter at 300Hz that was applied on the recordings at the acquisition time. We are going to ignore this one, as it is probably more complex and spanning over several frequency bins. {{attachment:psdBefore.gif||height="172",width="399"}} === Remove: 60Hz and harmonics === Leave the file "Link to raw file" in the Process1 list and now run "'''Pre-process > Notch filter'''".<<BR>>Enter the frequencies to remove ('''60Hz, 120Hz '''and '''180Hz''') and leave the other options to the default values. {{attachment:sinRemoval.gif||height="272",width="333"}} The operation creates a new "Link to raw file" entry in the database, pointing to a new CTF dataset in the same folder as the original continuous file. To check where this file was created: right-click on the file "'''Raw | notch(60Hz 120Hz 180Hz)'''" > '''File '''> '''Show in file explorer'''. This file has the exact same properties as the original file, including the SSP projectors, only the values of the MEG sensors were updated. {{attachment:sinRemovalDb.gif||height="144",width="326"}} === Alternatives === If the notch filter is not giving satisfying result, you can use two other processes: * '''Sinusoid removal''': This process can do a better job at removing precise frequencies by identifying the sinusoidal components and then subtracting them from the signals in time domain (not a frequency filter). |
* '''Sinusoid removal''': This process can do a better job at removing precise frequencies by identifying the sinusoidal components and then subtracting them from the signals in the time domain (not a frequency filter). |
| Line 92: | Line 93: |
| === Evaluate the results === Now run the same frequency analysis: process "'''Frequency > Power spectrum density (Welch)'''", time window = '''[0,50]s''', window length = '''4000ms''', overlap = '''50%'''. Double click on the new "Power (MEG)" file. As expected, the three first peaks due to the power lines are gone. |
<<TAG(Advanced)>> |
| Line 95: | Line 95: |
| {{attachment:psdAfter.gif||height="177",width="411"}} | == Other filters == Other frequency filters could be interesting to run at an early stage of analysis. |
| Line 97: | Line 98: |
| You can run a few other pre-processing operations directly on the continuous files, including a band-pass filter. Frequency filters and sinusoid removals are operations that you should apply at very early stages of the analysis, before epoching the recordings. Those operations do not perform well next to the beginning and the end of the signals, they may generate important artifacts. It is therefore much more accurate to filter the entire recordings from the original continuous file at once, rather than filtering small epochs after importing them in the database. | * '''High-pass filter''': Removes the low frequencies from the signals. This could be used for: * Removing the slow fluctuations of the arbitrary baseline value of the MEG sensors (< 0.2Hz). * Removing the artifacts occurring at low frequencies (< 1Hz, for instance breathing). * Doing a baseline correction of the EEG/MEG recordings when there is no clear pre-stimulus baseline in the experiment or in the case of long epochs or resting state recordings (> 5s). * '''Low-pass filter''': Removes the high frequencies from the signals. This could be used for: * Removing some strong noise occurring at high frequencies. * Facilitating the interpretation of the signals: if you are interested only in lower frequencies (for instance < 20Hz), everything in the higher part of the spectrum is just noise in your analysis. * Both filters are accessible using the process '''Pre-process > Band-pass filter'''. |
| Line 99: | Line 107: |
| From now on, we are only going to work on this clean file "Raw | notch(60Hz 120Hz 180Hz)". | '''Important''': Frequency filters and sinusoid removals are operations that you should apply at very early stages of the analysis, before epoching the recordings. These operations do not perform well next to the beginning and the end of the signals, they may generate important artifacts. It is therefore much more efficient to filter the entire recordings from the original continuous file at once, rather than filtering small epochs after importing them in the database. |
| Line 101: | Line 109: |
| = From CTF = == Selecting files to process == First thing to do is to define the files you are going to process. This is done easily by picking files or folders in the database explorer and dropping them in the empty list of the Process1 tab. |
<<TAG(Advanced)>> |
| Line 105: | Line 111: |
| 1. Drag and drop the following nodes in the Process1 list: Right/ERF (recordings), Right (condition), and Subject01 (subject)<<BR>><<BR>> {{attachment:files1.gif}} * The number in the brackets next to each node represents the number of data files that where found in each of them. The node ERF "contains" only itself (1), Subject01/Right contains ERF and Std files (2), and Subject01 contains 2 conditions x 2 recordings (4). * The total number of files, ie. the sum of all those values, appears in the title of the panel "Files to process [7]". 1. The buttons on the left side allow you to select what type of file you want to process: Recordings, sources, time-frequency, other. Now select the second button "Sources". All the counts are updated and now reflect the number of sources files that are found for each node.<<BR>><<BR>> {{attachment:files2.gif}} 1. If you select the third button "Time-frequency", you would see "0" everywhere because there are no time-frequency decompositions in the database yet.<<BR>><<BR>> {{attachment:files3.gif}} 1. Now clear the list from all the files. You may either right-click on the list (popup menu ''Clear list''), or select all the nodes (holding ''Shift ''or ''Crtl ''key) and then press the ''Delete ''key. 1. Select both files Left/ERF and Right/ERF in the tree (holding ''Ctrl ''key), and put the in Process list. We are going to apply some functions on those two files. You cannot distinguish them after they are dropped in the list, because they are both referred as "ERP". If at some point you need to know what is in the list, just leave you mouse over a node for a few seconds, and a tooltip would give you information about it. Just like in the database explorer.<<BR>><<BR>> {{attachment:files4.gif}} |
== On the hard drive == The names of the files generated by the process "Power spectrum density" start with the tag '''timefreq_psd''', they share the same structure as all the files that include a frequency dimension. |
| Line 113: | Line 114: |
| <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/EventMarkers&next=Tutorials/ArtifactsSsp")>> | To explore the contents of a PSD file created in this tutorial, right-click on it and use the popup menus '''File > View file contents''' or '''File > Export to Matlab'''. {{attachment:psd_contents.gif}} * '''TF''': [Nsignals x Ntime x Nfreq] Stores the spectrum information. Nsignals is the number of channels that were selected with the option "MEG" in the PSD process. Nfreq is the number of frequency bins. There is no time dimension (Ntime = 1). * '''Comment''': String used to represent the file in the database explorer. * '''Time''': Window of time over which the file was estimated. * '''TimeBands''': Defined only when you select the option "Group in time bands". <<BR>>Always empty for the PSD files because there is no time dimension. * '''Freqs''': [1 x Nfreq] List of frequencies for which the power spectrum was estimated (in Hz). * '''RefRowNames''': Only used for connectivity results. * '''RowNames''': [Nsignals x 1] Describes the rows of the TF matrix (first dimension). Here it corresponds to the name of the MEG sensors, in the same order as is the .TF field. * '''Measure''': Function currently applied to the FFT coefficients {power, none, magnitude, log, other} * '''Method''': Function that was used to produce this file {psd, hilbert, morlet, corr, cohere, ...} * '''DataFile''': File from which this file was calculated = Parent file in the database explorer. * '''DataType''': Type of file from which this file was calculated (file type of .DataFile). * 'data' = Recordings * 'cluster' = Recordings grouped by clusters of sensors * 'results' = Source activations * 'scouts' = Time series associated with a region of interest * '''SurfaceFile '''/ '''GridLoc '''/ '''GridAtlas '''/ '''Atlas''': Used only when the input file was a source file. * '''nAvg''': Number of input files averaged to produce this file. * '''Options''': Most relevant options that were passed to the function bst_timefreq. * '''History''': Describes all the operations that were performed with Brainstorm on this file. To get a better view of this piece of information, use the menu '''File > View file history'''. <<HTML(<!-- END-PAGE -->)>> <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/PipelineEditor&next=Tutorials/BadChannels")>> |
Tutorial 10: Power spectrum and frequency filters
Authors: Francois Tadel, Elizabeth Bock, John C Mosher, Sylvain Baillet
We are now going to process our continuous recordings to remove the main sources of noise. Typically, we can expect contaminations coming from the environment (power lines, stimulation equipment, building vibrations) and from the subject (movements, blinks, heartbeats, breathing, teeth clenching, muscle tension, metal in the mouth or the body). In this tutorial, we will focus first on the noise patterns that occur continuously, at specific frequencies.
We can correct for these artifacts using frequency filters. Usually we prefer to run these notch filters, band-pass filters and sinusoid removals before any other type of correction, on the continuous files. They can be applied to the recordings without much supervision, but they may create important artifacts at the beginning and the end of the signals. Processing the entire continuous recordings at once instead of the imported epochs avoids adding these edge effects to all the trials.
Contents
Evaluation of the noise level
Before running any type of cleaning procedure on MEG/EEG recordings, we always recommend to start with a quick evaluation of the noise level. An easy way to do this is to estimate the power spectrum of all the signals over the entire recordings.
- Clear the list of files in the Process1 tab.
Select the three datasets we have linked to our protocol.
You can select the three "link to raw file" nodes, the three folders or the entire subject node.
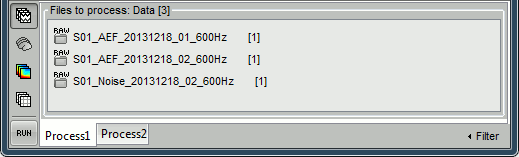
- Click on [Run] to open the pipeline editor window.
Select the process "Frequency > Power spectrum density (Welch)"
This process evaluates the power of the MEG/EEG signals at different frequencies, using the Welch's method (see Wikipedia or MathWorks). It splits the signals in overlapping windows of a given length, calculates the Fourier Transform (FFT) of each of these short segments, and averages the power of the FFT coefficients for all the overlapping windows.
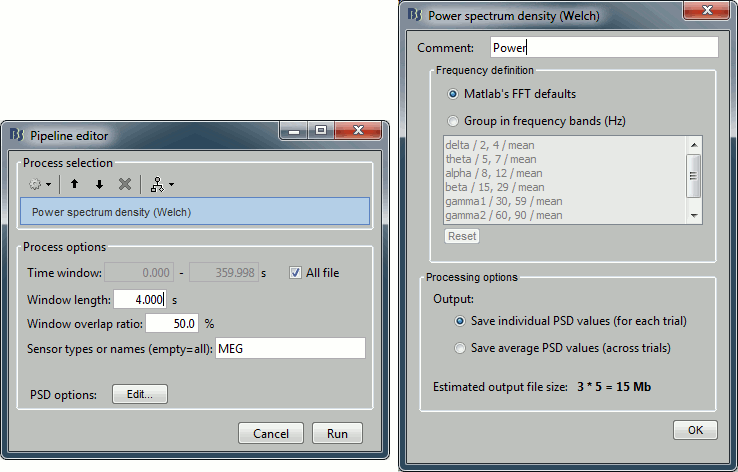
- Set the options as follows (click on [Edit] for the additional options):
Time window: [All file]
Portion of the input file you want to use for estimating the spectrum.Window length: 4 seconds
Estimator length = length of the overlapping time windows for which we calculate the FFT. The number of time samples in the estimator is the same as the number of frequency bins in the output file. Increasing this parameter increases the output frequency resolution (distance between two frequency bins) but degrades the stability of the estimator, as it also decreases the total number of averaged time windows.Overlap: 50%
How much overlap do we want between two consecutive time windows.Sensor types or names: MEG
Defines the list of channels (names or types) on which you want to apply the process.Frequency definition: Matlab's FFT default
You have the option to directly use the frequency binning returned by the FFT, or run an additional step of averaging these bins in larger frequency bands. Note that you can edit freely these frequency bands.Output: Save individual PSD value.
This option will separately estimate the PSD for each of the three files in input, and create three files in output. If you select the other option (save average), it calculates the same three files but averages them on the fly and saves only one file in the database.
- Click on [Run] to start the execution of the process.
Troubleshooting: If you get "Out of memory" errors, try to run this PSD estimation on a shorter time segment. For example, set the time window to [0,100s] instead of the full file. This process starts by loading all the needed recordings in memory, you might not have enough memory available on your system to fit the entire dataset.
It produces three new files, that appear as "depending" on the three datasets. The comments of the files indicate how many overlapping windows could be used to estimate the PSD. "179/4000ms" means 179 windows of 4s each (total 716s). With the 50% overlap, it sums up to a little less than 2x the file length (360s).
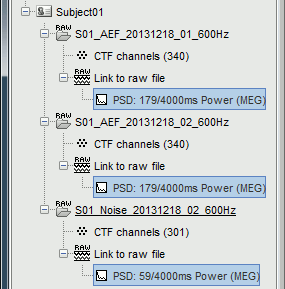
Interpretation of the PSD
File: AEF#01
Double-click on the new PSD file for run #01 to display it (or right-click > Power spectrum).
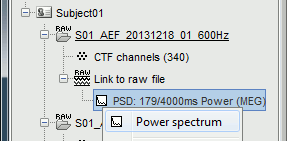
- The power spectrum is displayed in a figure similar to the time series, but the X axis represents the frequencies. Most of the shortcuts described for the recordings are also valid here. Clicking on the white parts of the figure or using the arrow keys of the keyboard move the frequency cursor. The current frequency can also be controlled with a new slider displayed in the Brainstorm window, just below the time panel.
Each black line represent the spectrum for one channel. If you click on a channel, it is selected and highlighted in red. Click again for deselecting. Right-click on a channel to read its name.
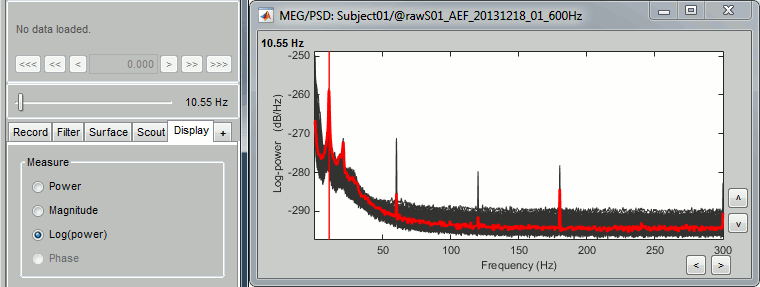
- The shape of this graph is normal, it does not indicate anything unexpected:
Peaks related with the subject's alpha rhythms: around 10Hz and 20Hz.
Peaks related with the power lines: 60Hz, 120Hz and 180Hz.
These datasets were recorded in Canada, where the alternating powerline current is delivered at 60Hz. In Europe you would observe similar peaks at 50Hz, 100Hz and 150Hz.
- Add a topography view for the same file, with one of the two methods below:
Right-click on the PSD file > 2D Sensor cap.
Right-click on the spectrum figure > 2D Sensor cap (shortcut: Ctrl+T)
Scroll in frequencies to see the spatial distribution of each frequency bin:
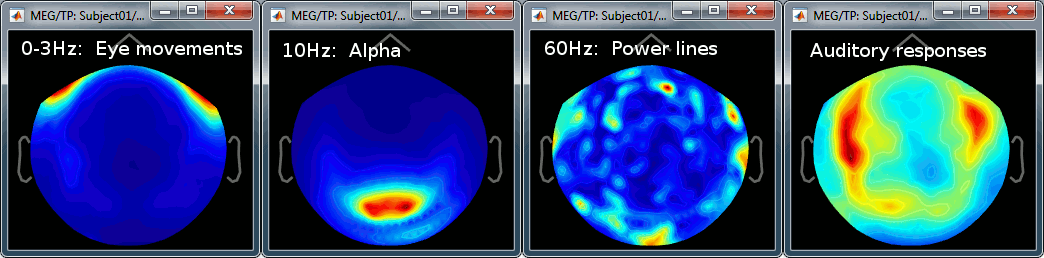
- We have already identified two artifacts we will need to remove: the eye movements and the 60Hz+harmonics from the power lines.
File: AEF#02
Open the spectrum view for the run AEF#02.
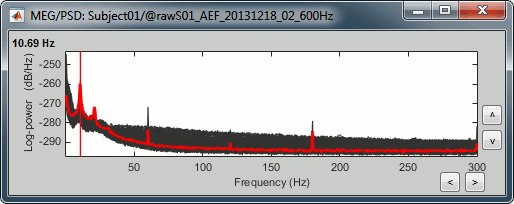
Add a 2D sensor cap view for the same file. Scroll to zoom in/out.
To display the sensor names, right-click on the figure > Channels > Display sensors.- This spectrum looks very similar to the run #01: same alpha and power line peaks.
Additionally, we observe higher signal power between 30Hz and 100Hz on many occipital sensors. This is probably related to some tension in the neck muscles due to an uncomfortable seating position in the MEG. We will see later whether these channels need to be tagged as bad or not.
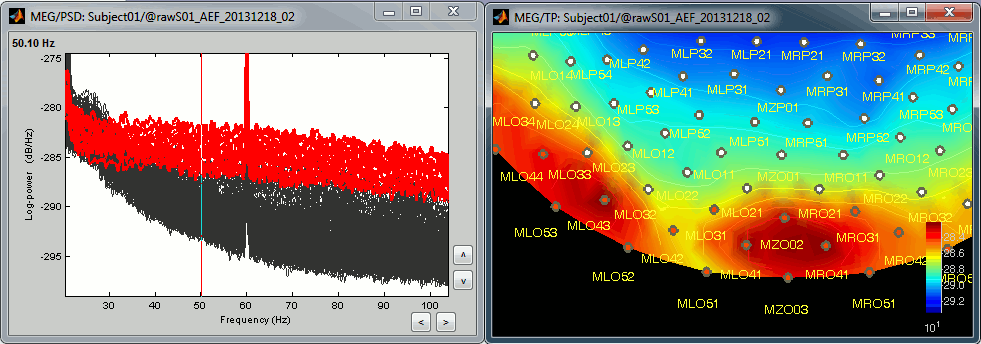
File: Noise recordings
Open the spectrum view for the noise recordings.
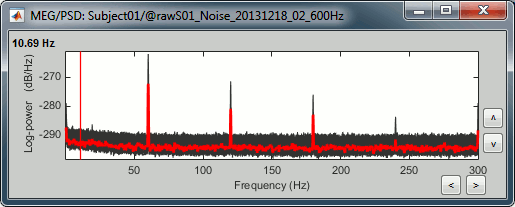
- This shows the power spectrum of the signals that are recorded when there is no subject in the MEG room. It gives a good and simple representation of the instrumental noise. If you had one bad MEG sensor, you would see it immediately in this graph. Here everything looks good.
Notch filter
Let's start with the cleaning for the 60Hz+harmonics. Notch filters are adapted for removing well identified contaminations from systems oscillating at very stable frequencies.
Keep all the three datasets selected in the Process1 box.
All the filters that are applied to the experiment data also need to be applied to the noise recordings. In the source estimation process, we will need all the files to have similar levels of noise.- Click on [Run] to open the Pipeline editor.
Run the process: Pre-process > Notch filter
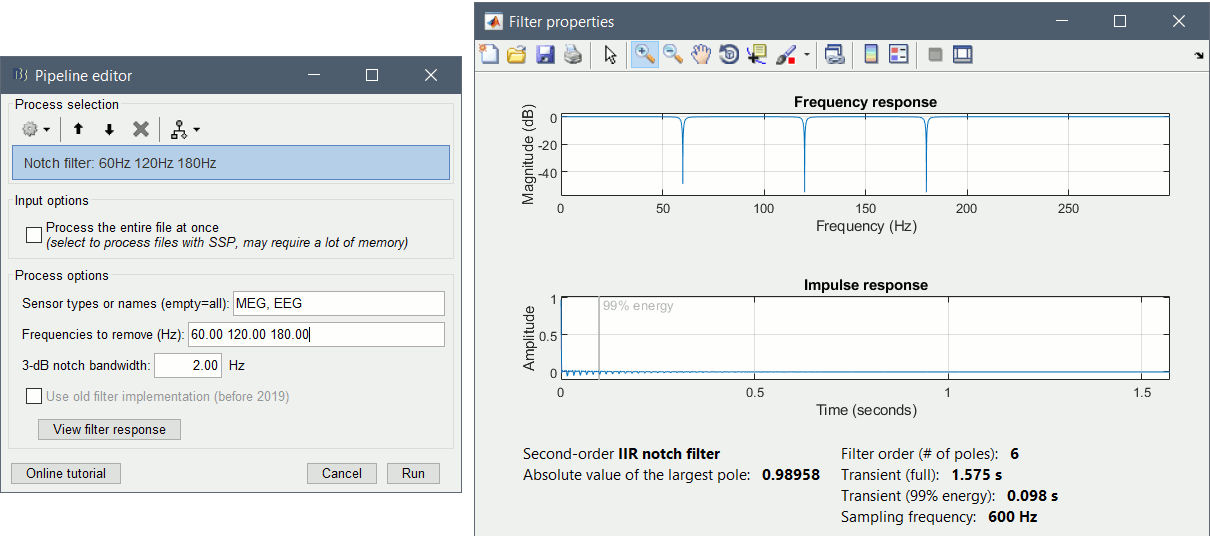
Process the entire file at once: NO
Select the frequencies: 60, 120, 180 Hz
Sensor types or names: MEG
- The higher harmonics (240Hz) are not clearly visible, and too high to bother us in this analysis.
This process creates three new datasets, with additional "notch" tags. These output files are saved directly in the Brainstorm database, in a binary format specific to Brainstorm (.bst).
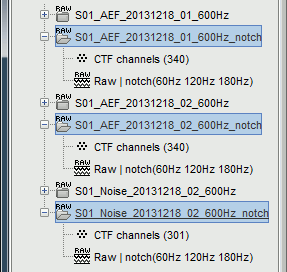
- If you delete the folders corresponding to the original files (before the filter), your original recordings in the .ds folders are not altered. If you delete the folders corresponding to the filtered files, you will lose your filtered recordings in .bst format.
To check where the file corresponding to a "Link to raw file" is actually saved on the hard drive, right-click on it > File > Show in file explorer.
Evaluation of the filter
Right-click on the Process1 list > Clear list.
Drag and drop the three filtered files in Process1.
Run again the PSD process "Frequency > Power spectrum density (Welch)" on the new files, with the same parameters as before, to evaluate the quality of the correction.
Double-click on the new PSD files to open them.
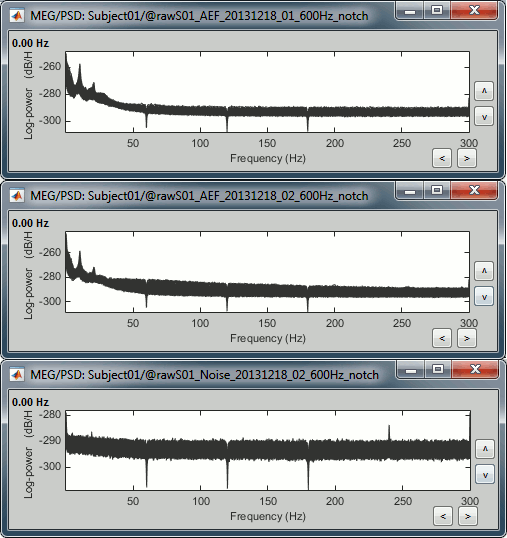
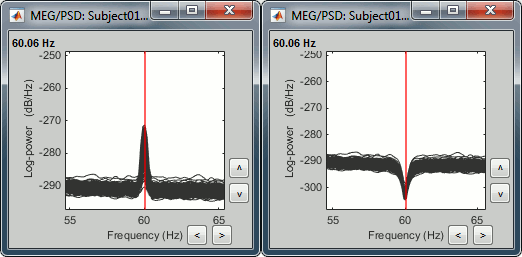
Scroll to zoom in and observe what is happening around 60Hz (before / after).
Some cleaning
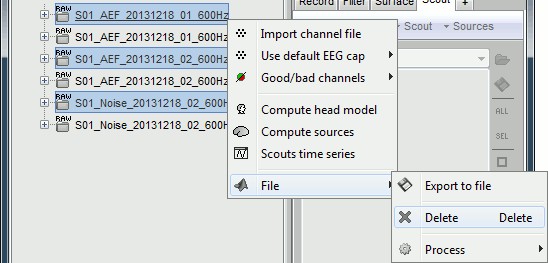
To avoid the confusion later, delete the links to the original files:
Select the folders containing the original files and press Delete (or right-click > File > Delete).
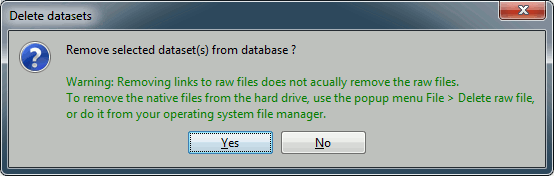
Always read the confirmation messages carefully, you will avoid some bad surprises.
This is what your database explorer should look like at the end of this tutorial:
Alternatives to the notch
If the notch filter is not giving satisfying result, you can use two other processes.
Sinusoid removal: This process can do a better job at removing precise frequencies by identifying the sinusoidal components and then subtracting them from the signals in the time domain (not a frequency filter).
Band-stop filter: This process can be useful for removing larger segments of the spectrum, in case the power line peaks are spread over numerous frequency bins or for suppressing other types of artifacts.
Other filters
Other frequency filters could be interesting to run at an early stage of analysis.
High-pass filter: Removes the low frequencies from the signals. This could be used for:
Removing the slow fluctuations of the arbitrary baseline value of the MEG sensors (< 0.2Hz).
Removing the artifacts occurring at low frequencies (< 1Hz, for instance breathing).
Doing a baseline correction of the EEG/MEG recordings when there is no clear pre-stimulus baseline in the experiment or in the case of long epochs or resting state recordings (> 5s).
Low-pass filter: Removes the high frequencies from the signals. This could be used for:
- Removing some strong noise occurring at high frequencies.
Facilitating the interpretation of the signals: if you are interested only in lower frequencies (for instance < 20Hz), everything in the higher part of the spectrum is just noise in your analysis.
Both filters are accessible using the process Pre-process > Band-pass filter.
Important: Frequency filters and sinusoid removals are operations that you should apply at very early stages of the analysis, before epoching the recordings. These operations do not perform well next to the beginning and the end of the signals, they may generate important artifacts. It is therefore much more efficient to filter the entire recordings from the original continuous file at once, rather than filtering small epochs after importing them in the database.
On the hard drive
The names of the files generated by the process "Power spectrum density" start with the tag timefreq_psd, they share the same structure as all the files that include a frequency dimension.
To explore the contents of a PSD file created in this tutorial, right-click on it and use the popup menus File > View file contents or File > Export to Matlab.
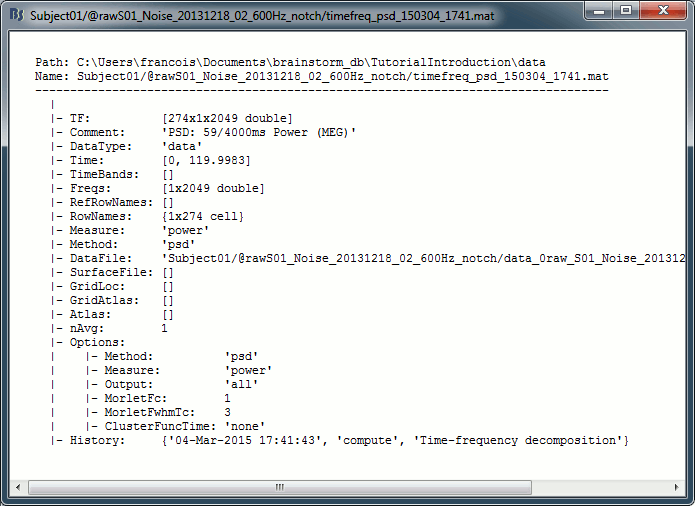
TF: [Nsignals x Ntime x Nfreq] Stores the spectrum information. Nsignals is the number of channels that were selected with the option "MEG" in the PSD process. Nfreq is the number of frequency bins. There is no time dimension (Ntime = 1).
Comment: String used to represent the file in the database explorer.
Time: Window of time over which the file was estimated.
TimeBands: Defined only when you select the option "Group in time bands".
Always empty for the PSD files because there is no time dimension.Freqs: [1 x Nfreq] List of frequencies for which the power spectrum was estimated (in Hz).
RefRowNames: Only used for connectivity results.
RowNames: [Nsignals x 1] Describes the rows of the TF matrix (first dimension). Here it corresponds to the name of the MEG sensors, in the same order as is the .TF field.
Measure: Function currently applied to the FFT coefficients {power, none, magnitude, log, other}
Method: Function that was used to produce this file {psd, hilbert, morlet, corr, cohere, ...}
DataFile: File from which this file was calculated = Parent file in the database explorer.
DataType: Type of file from which this file was calculated (file type of .DataFile).
- 'data' = Recordings
- 'cluster' = Recordings grouped by clusters of sensors
- 'results' = Source activations
- 'scouts' = Time series associated with a region of interest
SurfaceFile / GridLoc / GridAtlas / Atlas: Used only when the input file was a source file.
nAvg: Number of input files averaged to produce this file.
Options: Most relevant options that were passed to the function bst_timefreq.
History: Describes all the operations that were performed with Brainstorm on this file. To get a better view of this piece of information, use the menu File > View file history.
