|
Size: 30465
Comment:
|
Size: 28158
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 6: | Line 6: |
| If an event is very reproducible and occurs always at the same position (eg. eye blinks and heartbeats), the sensors will always record the same values when it occurs. We can identify the topographies corresponding to this artifact (ie spatial distributions of values at one time point) and remove them from the recordings. This spatial decomposition is the basic idea behind two widely used approaches: the '''SSP '''(Signal-Space Projection) and '''ICA '''(Independent Component Analysis) methods. This introduction tutorial will focus on the SSP approach, as it is a lot simpler and faster but still very efficient for removing blinks and heartbeats from MEG and high-density EEG recordings. You will be able to read later about the ICA decompositions in the advanced tutorials. |
If an event is very reproducible and occurs always at the same position (eg. eye blinks and heartbeats), the sensors will always record the same values when it occurs. We can identify the topographies corresponding to this artifact (ie. the spatial distributions of values at one time point) and remove them from the recordings. This spatial decomposition is the basic idea behind two widely used approaches: the '''SSP '''(Signal-Space Projection) and '''ICA '''(Independent Component Analysis) methods. This introduction tutorial will focus on the SSP approach, as it is a lot simpler and faster but still very efficient for removing blinks and heartbeats from MEG and high-density EEG recordings. The interface for running ICA decompositions is very similar and will be described in an advanced tutorial. |
| Line 15: | Line 15: |
| 1. We start by accumulating as many examples of the artifact we are trying to remove. This is what we've been doing in the previous tutorial with the creation of the "cardiac" and "blink" events. | 1. We start by identifying many examples of the artifact we are trying to remove. This is what we've been doing in the previous tutorial with the creation of the "cardiac" and "blink" events. |
| Line 17: | Line 17: |
| 1. We run a principle components analysis (PCA) on the concatenated artifacts in order to get a decomposition in various spatial components. 1. We should find in the first few principal components some topographies that are very specific of the type of artifact we are targetting. We select those components to remove. |
1. We run a principle components analysis (PCA) on the concatenated artifacts in order to get a decomposition in various spatial components (number of components = number of sensors). 1. If it works well, we can find in the first few principal components some topographies that are very specific of the type of artifact we are targetting. We select those components to remove. |
| Line 20: | Line 20: |
| 1. Whenever some data is recordings are read from this file, the SSP projectors will be applied on the fly to them, to remove the artifact contributions. This approach is fast and memory efficient.<<BR>><<BR>> {{attachment:ssp_intro.gif}} |
1. Whenever some recordings are read from this file, the SSP projectors are applied on the fly to remove the artifact contributions. This approach is fast and memory efficient.<<BR>><<BR>> {{attachment:ssp_intro.gif}} == The order matters == This procedure has to be repeated separately for each artifact type. The order in which you process the artifacts matters, because for removing the second artifact we typically use the recordings cleaned with the first set of SSP projectors. We have to decide which one to process first. It works best if each artifact is defined precisely and as independently as possible from the other artifacts. If the two artifacts happen simulateneously, the SSP projectors calculated for the blink may contain some of the heartbeat topography and vice versa. When trying to remove the second artifact, we might not be able to isolate it clearly anymore. Because the heart beats every second or so, there is a high chance that when the subject blinks there is a heartbeat not too far away in the recordings. Therefore a significant number of the blinks will be contaminated with heartbeats. But we have usually a lot of "clean" heartbeats, we can start by removing those ones. To isolate correctly these two common artifacts, we recommend the following procedure: * Remove the markers "cardiac" that are occurring during a blink (done in the previous tutorial), * Compute the '''cardiac SSP''' (with no eye movements, because we removed the co-occurring events), * Compute the '''blink SSP''' (with no heartbeats, because they've already been taken care of). If you have multiple modalities recorded simultaneously, for example MEG and EEG, you should run twice this entire procedure, once for the EEG only and once for the MEG only. You will always get better results if you '''process the different types of sensors separately'''. Same thing when processing Elekta-Neuromag recordings: you should process separately the magnetometers (MEG MAG) and the gradiometers (MEG GRAD). |
| Line 25: | Line 36: |
| * '''Run #01''': Double-click on the link to show the MEG sensors. * '''Configuration''': Page of '''3 seconds''', view in columns, selection of the "'''CTF LT'''" sensors (the left-temporal sensors will be a good example to show at the same time the two types of artifacts). * In the Record tab, select the menu: '''"Artifacts > SSP: Heartbeats"'''. . o SSP > Compute SSP: Heartbeats . § Display 2D topography for the first spatial component (clearly a heartbeat) . § Select component #1: It is clearly a cardiac component . § Show the influence of the projector on the sensors LT . o SSP > Compute SSP: Eye blinks . § Display the 2D topography for the first spatial component (clearly a blink) . § Show the influence of the projector on the sensors LT == Heartbeats and eye blinks == * Select the two AEF runs in the Process1 box. * Select successively the following processes, then click on [Run]: * '''Compute SSP: Heartbeats''': Event name "cardiac", sensors="MEG", '''do not use existing SSP'''. * '''Compute SSP: Eye blinks''': Event name "blink", sensors="MEG", '''do not use existing SSP'''. {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=ssp_pipeline.gif|ssp_pipeline.gif|height="359",width="527",class="attachment"}} * Double-click on '''Run01 '''to open the MEG. You can change the color of events "standard" and "deviant" to make the figure more readable. * Review the '''EOG '''and '''ECG '''channels and make sure the events detected make sense. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=events.gif|events.gif|height="369",width="557",class="attachment"}} * In the Record tab, menu SSP > '''Select active projectors'''. * Blink: The first component is selected and looks good. * Cardiac: The category is disabled because no component has a value superior to 12%. * Select the first component of the cardiac category and display its topography. * It looks exactly like a cardiac topography, keep it selected and click on [Save]. {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=ssp_result.gif|ssp_result.gif|height="172",width="700",class="attachment"}} * Repeat the same operations for '''Run02''': * Review the events. * Select the first cardiac component. |
==== Computation ==== Double-click on the link to show the MEG sensors for '''Run #01'''.<<BR>>In the Record tab, select the menu: '''"Artifacts > SSP: Heartbeats"'''. * '''Event name''': Name of the event to use to calculate the projectors, enter "'''cardiac'''" * '''Sensor types''': Type of sensors for which the projection should be calculated ("'''MEG'''"). Note that you will always get better results if you process the different types of sensors separately. * '''Compute using existing SSP projectors''': You have the option to calculate the projectors from the raw recordings, or from the recordings filtered with the previously computed SSP projectors. Unless you have a good reason for not considering the existing projectors, you should select this option. <<BR>><<BR>> {{attachment:ssp_ecg_process.gif||height="277",width="520"}} After the computation is done, a new figure is displayed, that lets you select the active projectors. * '''On the left''': The projector categories where each row represents the result of an execution of this process (usually one for each sensor type and each artifact). * '''On the right''': The spatial components returned by the PCA decomposition. The percentage indicated between brackets is the singular value for this each component, normalized for this decomposition (percentage = '''S'''i / sum('''S'''i), see technical details at the end of this page). * '''Percentage''': More practically, it indicates the amount of signal that was captured by the component during the decomposition. The highest it is, the more the component is representative of the artifact recordings that were used to calculate it. In the good cases, you would typically see one to three components with values that are significantly higher that the others. * When a component is selected, it means that it is removed from the recordings. A spatial projector is computed and applied to the recordings on the fly when reading from the continuous file. * '''Default selection''': The software selects '''the components with values superior to 12%''' and leaves the others unselected. '''This threshold is completely empirical''' '''and depends on the acquisition device''', you should always review manually the components that you want to remove. <<BR>><<BR>> {{attachment:ssp_select1.gif||height="178",width="366"}} ==== Evaluate the components ==== For the cardiac event, none of the components show a value superior to 12%, therefore the entire "cardiac" projector category was unselected. This doesn't mean that none of those components is actually interesting for us. The percentage indicated for the first value (9%) is much higher than the following ones (5%, 5%, 4%, 3%...), this could indicate that it targets relatively well the cardiac artifact. Let's investigate this in more details. * Select the category ("cardiac") to enable the list on the right. * Click on the first component, then click on the toolbar button '''[Display component topography]'''. This menu shows the spatial distribution of the values recorded by the MEG sensors for this component. Note that you don't have to select the component (ie. check the box) to display it. <<BR>><<BR>> {{attachment:process_select2.gif}} * The second button "'''Display component topography [No magnetic interpolation]'''" produces the same figure but without the reinterpolation of the magnetic fields that is typically applied to the MEG recordings in Brainstorm. This procedure will be detailed later in the introdcution tutorials. * The last button in the toolbar '''[Display component time series]''', opens a figure that represents the evolution of the contribution of this component over time. The higher the value, the more present the selected topography in the recordings. Click on it to show the''' component #1''', and display the '''ECG '''signal at the same time (right-click on the file > ECG > Display time series). <<BR>> We observe that <<BR>><<BR>> {{attachment:process_cardiac_good.gif}} * But <<BR>><<BR>> {{attachment:process_cardiac_bad.gif}} Do it again for components #2 and #3 (the components don't need to be active for that). The other button [No interp] displays the same results without the magnetic field smoothing that is applied by default to MEG topographies. It may help understand some difficult cases. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEcgComp.gif|sspEcgComp.gif|class="attachment"}} ==== Evaluate the correction ==== None of those topographies can be clearly related to a cardiac component. This can have two interpretations: the cardiac artifact is not very strong for this subject and the influence of the heart activity over the MEG sensors is completely buried in the noise or the brain signals, or the characterization of the artifact that we did was not so good and we should refine it by selecting for instance smaller time windows around the cardiac peaks. Here, it's probably due to the subject's morphology. Some people generate strong artifacts in the MEG, some don't. In this case, it is safer to '''unselect '''this "cardiac" category, rather than removing randomly spatial components that we are not sure to identify. == SSP: Eye blinks == Let's try the same thing with the eye blinks. Select the process "'''SSP > Compute SSP: Eye blinks'''". Run it on the event type "blink", that indicate the peaks of the EOG signal. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogFile.gif|sspEogFile.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEog.gif|sspEog.gif|height="214",width="307",class="attachment"}} In this specific case, there is one value that is much higher than the others (21%), and it is selected by default because it is superior to 12%. The first component is most likely a good representation of the artifact we are trying to remove. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogSelect.gif|sspEogSelect.gif|height="214",width="384",class="attachment"}} Select the cardiac category, and display the topographies for the first three components. Component #1 is clearly related with the ocular activity occurring during the blinks, the others are not as clear. Conclusion: the default proposition was good (only component #1 selected). Setting this first projector as active removes from the MEG signals the contributions related with the spatial topography we can see in the first figure, ie. a spatial distribution of sensor values related specifically to the eye blinks. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogComp.gif|sspEogComp.gif|class="attachment"}} While the recordings are displayed in the file viewer, the selection of the active projectors interactively updates the recordings. Go to the first detected "blink" event ('''t=33.600s'''), and select the left frontal sensors ('''Shift+B''' or right-click > Display setup > CTF LF). Then play with the checkboxes to change the active projectors, and see how well this first component of the blink category removes the ocular artifact. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogMeg.gif|sspEogMeg.gif|class="attachment"}} Select the projector category "blink", select the first component, and click on Save, to validate your changes. Starting from now, all the successive operations you apply on this file will integrate this eye blink correction, including the calculation of other projectors. After you closed this window, you can get back to it by selecting the menu "'''SSP > Select active projector'''" in the Record tab, and change again the selection of active projectors. We are not going to calculate the SSP for the other category of events identified on the EOG, the group "'''blink2'''". The events classified in this group could represent eye saccades or other types of blinks, it is not very clear that they correspond to the same physiological event. Most of them do not seem to create any obvious contamination on the MEG recordings, there is probably no need to consider them now. |
| Line 70: | Line 103: |
| = From Continuous = == Signal Space Projection == |
Always check visually the results of the SSP cleaning executed from a script. * Open the MEG sensors for Run #02. * In the Record tab, menu Artifacts > '''Select active projectors'''. <<TAG(Advanced)>> == SSP: Generic == The calculation of the SSP for the heartbeats and the eye blinks are shortcuts to a more generic algorithm "'''Compute SSP: Generic'''". We do not need it here, but the illustration of its properties is interesting to understand the correction of ocular and cardiac artifacts: * '''Event name''': Markers that are used to characterize the artifact * '''Time window''': Time segment to consider before and after each event marker. We want this time window to be longer than the artifact effect itself, in order to have a large number of time samples representative of a normal brain activity. This helps the SVD decomposition to separate the artifact from the ongoing brain activity. * '''Frequency band''': Definition of the band-pass filter that is applied to the recordings before calculating the projector. Usually you would use the same frequency band as we used for the detection, but you may want to try to refine this parameter if the results are not satisfying. * '''Sensor types or names''': List of sensors for which to calculate the projector. You can get better results if you process one sensor type at a time. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspGeneric.gif|sspGeneric.gif|height="328",width="330",class="attachment"}} <<TAG(Advanced)>> == Evaluation == === Eye blinks === One efficient way of representing the impact of this artifact correction is to epoch the recordings around the artifacts before and after the correction. Use the same time window as the one used in the SSP options around each marker, and average all the segments of recordings. Those operations are detailed in the next tutorial, we are just presenting the results. You don't need to reproduce them at this time, just remember that it is doable, in case you need it with your own data. This is what the database would look like after those operations: {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=blinkDb.gif|blinkDb.gif|height="263",width="283",class="attachment"}} The following image represents the file '''"blink_uncorrected / Avg: blink"''', which is the average of the 18 identified blinks '''before''' the SSP correction, [-200,+200]ms around the "blink" events. The time series, the 2D topography at the peak (t = 0ms), and the mapping on the cortex of those fields, using the basic inverse model calculated in the previous tutorial. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=blinkBefore.gif|blinkBefore.gif|height="177",width="533",class="attachment"}} The next image represents the file '''"blink_ssp / Avg: blink"''', which is the exact same thing, but '''after '''the SSP correction. The artifact is gone. If you do this and you can still see some clear evoked component in the time series figure, the SSP correction was not efficient: the artifact is not properly identified, or you should select different components using the "Select active projectors" window. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=blinkAfter.gif|blinkAfter.gif|height="177",width="533",class="attachment"}} === Heartbeats === We can do the same thing with the heartbeats: epoch [-40,+40]ms around the "cardiac" events, average the trials, and review the event-related fields at three instants: The peak of the '''P wave''' (t = '''-30ms'''): {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=cardiacP.gif|cardiacP.gif|height="178",width="534",class="attachment"}} The peak of the '''QRS complex''' (t = '''0ms'''): {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=cardiacQRS.gif|cardiacQRS.gif|height="178",width="534",class="attachment"}} The peak of the '''T wave''' (t = '''25ms'''): {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=cardiacT.gif|cardiacT.gif|height="178",width="534",class="attachment"}} Those peaks may look big, but they are in fact much smaller (300 fT for the average of 346 repetitions) than what we observed for the eye blinks (1500 fT for 18 repetitions). You can notice that none of those topographies are similar to the components we obtained after calculating the SSP for the cardiac artifact. We don't know how to correct this artifact, it doesn't look too bad in terms of recordings contamination, so we just leave it uncorrected. <<TAG(Advanced)>> == Troubleshooting == You have calculated the SSP as indicated here but you don't get any good results. No matter what you do, the topographies don't look like the targeted artifact. You can try the following: * Review one by one the events indicating the artifacts, remove the ones that are less clear or that occur close to another artifact. * Select or unselect the option "Compute using existing SSP". * Use the process "Compute SSP: Generic" and modify some parameters: * Reduce the time window around the peak of the artifact. * Use a narrower frequency band: especially the EOG, if the projectors capture some of the alpha oscillations, you can limit the frequency band to [1.5 - 9] Hz. * Change the method: Average / SSP. * If you have multiple acquisition runs, you may try to use all the artifacts from all the runs rather than processing the files one by one. For that, use the Process2 tab instead of Process1. Put the "Link to raw file" of all the runs on both side, Files A (what is used to compute the SSP) and Files B (where the SSP are applied). Always look at what this procedure gives you in output. Most of the time, the artifact cleaning will be an iterative process where to need several time to adjust the options and the order of the different steps in order to get good results. <<TAG(Advanced)>> == Theory == |
| Line 101: | Line 200: |
| For more information on the SSP method, please consult the following publications: * C. D. Tesche, M. A. Uusitalo, R. J. Ilmoniemi, M. Huotilainen, M. Kajola, and O. Salonen, "Signal-space projections of MEG data characterize both distributed and well-localized neuronal sources," Electroencephalogr Clin Neurophysiol, vol. 95, pp. 189-200, 1995. * M. A. Uusitalo and R. J. Ilmoniemi, "Signal-space projection method for separating MEG or EEG into components," Med Biol Eng Comput, vol. 35, pp. 135-40, 1997. == Identify the artifacts == The first step is to identify several repetitions of the artifact (the vectors '''b'''<<HTML(<sub>)>>''1''<<HTML(</sub>)>>...'''b'''<<HTML(<sub>)>>''m''<<HTML(</sub>)>>). We need to set markers in the recordings that indicate when the events that we want to correct for occur. To help with this task, it is recommended to always record with bipolar electrodes the activity of the eyes (electro-oculogram or EOG, vertical and horizontal), the heart (electro-cardiogram or ECG), and possibly other sources of muscular contaminations (electromyogram or EMG). In this example, we are going to use the ECG and vertical EOG traces to mark the cardiac activity and the eye blinks. Two methods can be used, manual or automatic. == Calculate the projectors == === Methodology === |
<<TAG(Advanced)>> == Algorithm == |
| Line 131: | Line 223: |
| === Recommended procedure === We have multiple types of artifact to correct for and it can be tricky to know what artifact to correct first. If you correct for artifact A and then for artifact B, you may get different results than if you correct first B and then A. We propose here a default order for the two artifacts that we have in all the recordings: the blinks and the heartbeats. If two artifacts happen at the same time, the SSP projectors calculated for the blink may contain some of the heartbeat and vice versa. When trying to remove the second artifact, we might not be able to isolate it clearly anymore. To avoid this behavior, we should try to avoid having overlapping artifacts. Because the subject's heart is beating at least once per second, there is a high chance to record a heartbeat next to any eye blink. Therefore a significant number of the blinks will be contaminated with heartbeats. If we decide to remove all the blink events that include some heart activity, we may not have enough blink samples left to build correct SSP projectors. To isolate correctly these two common contaminations we recommend the following procedure: 1. Remove all the cardiac peaks that are occurring next to a blink (we can remove half of the heartbeats, we will still have enough data for calculating the projectors). 1. Calculate the SSP to remove the '''cardiac '''artifact. 1. Calculate the SSP to remove the '''eye blinks''', based on the recordings already corrected for the cardiac artifact. === Compute SSP: Heartbeats === In the Record tab, select the menu "'''SSP > Compute SSP: Heartbeats'''". Set the options: * '''Event name''': Name of the event to use to calculate the projectors, enter "'''cardiac'''" * '''Sensor types''': Type of sensors for which the projection should be calculated ("'''MEG'''"). It doesn't matter if there are other sensor types indicated in the text box: the sensor names or types that do not exist in the recordings are just ignored. Note that you will always get better results if you process the different types of sensors separately. If you have MEG and EEG in the same file, you should run twice this process, once for the EEG only and once for the MEG only. Same thing when processing Elekta-Neuromag recordings, you should process separately the magnetometers (MEG MAG) and the gradiometers (MEG GRAD). * '''Compute using existing SSP projectors''': You have the option to calculate the projectors from the raw recordings, or from the recordings filtered with the previously computed SSP projectors. Unless you have a good reason for not considering the existing SSP projectors, you should always select this option. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogFile.gif|sspEogFile.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEcg.gif|sspEcg.gif|height="212",width="309",class="attachment"}} After the computation is done, a new figure is displayed, that lets you select the active projectors. On the left you have the projector categories (matrix '''U''') where each row represents the result of an execution of a "Compute SSP " process (usually one for each sensor type and each artifact). On the right, you can activate independently the different components for the selected projector category. Each entry represents the projector created from a column vector '''U'''i (singular vector of the SVD decomposition), and the percentage is the singular value for this vector, normalized for this decomposition (percentage = '''S'''i / sum('''S'''i)). When a projector is selected, it means that it is applied to the recordings on the fly when reading from the continuous file, ie. that the corresponding spatial component is removed from the recordings. The percentage indicates the amount of signal that was captured by this projector during the decomposition. The highest it is, the more the projector is representative of the artifact recordings that were used to calculate it. In the good cases, you would typically see one to three components with values that are significantly higher that the others. By default, the software selects '''the components with values superior to 12%''' and leaves all the others unselected. '''This threshold is completely empirical''' '''and depends on the acquisition device''', you should always review manually the components that you want to remove. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEcgSelect.gif|sspEcgSelect.gif|height="206",width="386",class="attachment"}} For the cardiac event, none of the components show a value superior to 12%, therefore the entire "cardiac" projector category was unselected. This doesn't mean that none of those components is actually interesting for us, let's investigate this. A list of buttons is available to manipulate the projector categories: load, save, rename, delete, and '''display component topography'''. The last one is one of the most useful tool to understand those results. Select the category ("cardiac") to enable the list on the right, click on the first component, then click on the toolbar button to display its topography. Do it again for components #2 and #3 (the components don't need to be active for that). The other button [No interp] displays the same results without the magnetic field smoothing that is applied by default to MEG topographies. It may help understand some difficult cases. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEcgCheck.gif|sspEcgCheck.gif|height="207",width="390",class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEcgComp.gif|sspEcgComp.gif|class="attachment"}} None of those topographies can be clearly related to a cardiac component. This can have two interpretations: the cardiac artifact is not very strong for this subject and the influence of the heart activity over the MEG sensors is completely buried in the noise or the brain signals, or the characterization of the artifact that we did was not so good and we should refine it by selecting for instance smaller time windows around the cardiac peaks. Here, it's probably due to the subject's morphology. Some people generate strong artifacts in the MEG, some don't. In this case, it is safer to '''unselect '''this "cardiac" category, rather than removing randomly spatial components that we are not sure to identify. === Compute SSP: Eye blinks === Let's try the same thing with the eye blinks. Select the process "'''SSP > Compute SSP: Eye blinks'''". Run it on the event type "blink", that indicate the peaks of the EOG signal. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogFile.gif|sspEogFile.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEog.gif|sspEog.gif|height="214",width="307",class="attachment"}} In this specific case, there is one value that is much higher than the others (21%), and it is selected by default because it is superior to 12%. The first component is most likely a good representation of the artifact we are trying to remove. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogSelect.gif|sspEogSelect.gif|height="214",width="384",class="attachment"}} Select the cardiac category, and display the topographies for the first three components. Component #1 is clearly related with the ocular activity occurring during the blinks, the others are not as clear. Conclusion: the default proposition was good (only component #1 selected). Setting this first projector as active removes from the MEG signals the contributions related with the spatial topography we can see in the first figure, ie. a spatial distribution of sensor values related specifically to the eye blinks. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogComp.gif|sspEogComp.gif|class="attachment"}} While the recordings are displayed in the file viewer, the selection of the active projectors interactively updates the recordings. Go to the first detected "blink" event ('''t=33.600s'''), and select the left frontal sensors ('''Shift+B''' or right-click > Display setup > CTF LF). Then play with the checkboxes to change the active projectors, and see how well this first component of the blink category removes the ocular artifact. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspEogMeg.gif|sspEogMeg.gif|class="attachment"}} Select the projector category "blink", select the first component, and click on Save, to validate your changes. Starting from now, all the successive operations you apply on this file will integrate this eye blink correction, including the calculation of other projectors. After you closed this window, you can get back to it by selecting the menu "'''SSP > Select active projector'''" in the Record tab, and change again the selection of active projectors. We are not going to calculate the SSP for the other category of events identified on the EOG, the group "'''blink2'''". The events classified in this group could represent eye saccades or other types of blinks, it is not very clear that they correspond to the same physiological event. Most of them do not seem to create any obvious contamination on the MEG recordings, there is probably no need to consider them now. === Compute SSP : Generic === The calculation of the SSP for the heartbeats and the eye blinks are shortcuts to a more generic algorithm "'''Compute SSP: Generic'''". We do not need it here, but the illustration of its properties is interesting to understand the correction of ocular and cardiac artifacts: * '''Event name''': Markers that are used to characterize the artifact * '''Time window''': Time segment to consider before and after each event marker. We want this time window to be longer than the artifact effect itself, in order to have a large number of time samples representative of a normal brain activity. This helps the SVD decomposition to separate the artifact from the ongoing brain activity. * '''Frequency band''': Definition of the band-pass filter that is applied to the recordings before calculating the projector. Usually you would use the same frequency band as we used for the detection, but you may want to try to refine this parameter if the results are not satisfying. * '''Sensor types or names''': List of sensors for which to calculate the projector. You can get better results if you process one sensor type at a time. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=sspGeneric.gif|sspGeneric.gif|height="328",width="330",class="attachment"}} == Evaluation == === Eye blinks === One efficient way of representing the impact of this artifact correction is to epoch the recordings around the artifacts before and after the correction. Use the same time window as the one used in the SSP options around each marker, and average all the segments of recordings. Those operations are detailed in the next tutorial, we are just presenting the results. You don't need to reproduce them at this time, just remember that it is doable, in case you need it with your own data. This is what the database would look like after those operations: {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=blinkDb.gif|blinkDb.gif|height="263",width="283",class="attachment"}} The following image represents the file '''"blink_uncorrected / Avg: blink"''', which is the average of the 18 identified blinks '''before''' the SSP correction, [-200,+200]ms around the "blink" events. The time series, the 2D topography at the peak (t = 0ms), and the mapping on the cortex of those fields, using the basic inverse model calculated in the previous tutorial. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=blinkBefore.gif|blinkBefore.gif|height="177",width="533",class="attachment"}} The next image represents the file '''"blink_ssp / Avg: blink"''', which is the exact same thing, but '''after '''the SSP correction. The artifact is gone. If you do this and you can still see some clear evoked component in the time series figure, the SSP correction was not efficient: the artifact is not properly identified, or you should select different components using the "Select active projectors" window. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=blinkAfter.gif|blinkAfter.gif|height="177",width="533",class="attachment"}} === Heartbeats === We can do the same thing with the heartbeats: epoch [-40,+40]ms around the "cardiac" events, average the trials, and review the event-related fields at three instants: The peak of the '''P wave''' (t = '''-30ms'''): {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=cardiacP.gif|cardiacP.gif|height="178",width="534",class="attachment"}} The peak of the '''QRS complex''' (t = '''0ms'''): {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=cardiacQRS.gif|cardiacQRS.gif|height="178",width="534",class="attachment"}} The peak of the '''T wave''' (t = '''25ms'''): {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawSsp?action=AttachFile&do=get&target=cardiacT.gif|cardiacT.gif|height="178",width="534",class="attachment"}} Those peaks may look big, but they are in fact much smaller (300 fT for the average of 346 repetitions) than what we observed for the eye blinks (1500 fT for 18 repetitions). You can notice that none of those topographies are similar to the components we obtained after calculating the SSP for the cardiac artifact. We don't know how to correct this artifact, it doesn't look too bad in terms of recordings contamination, so we just leave it uncorrected. == Troubleshooting == You have calculated the SSP as indicated here but you don't get any good results. No matter what you do, the topographies don't look like the targeted artifact. You can try the following: * Review one by one the events indicating the artifacts, remove the ones that are less clear or that occur close to another artifact. * Select or unselect the option "Compute using existing SSP". * Use the process "Compute SSP: Generic" and modify some parameters: * Reduce the time window around the peak of the artifact. * Use a narrower frequency band: especially the EOG, if the projectors capture some of the alpha oscillations, you can limit the frequency band to [1.5 - 9] Hz. * Change the method: Average / SSP. * If you have multiple acquisition runs, you may try to use all the artifacts from all the runs rather than processing the files one by one. For that, use the Process2 tab instead of Process1. Put the "Link to raw file" of all the runs on both side, Files A (what is used to compute the SSP) and Files B (where the SSP are applied). Always look at what this procedure gives you in output. Most of the time, the artifact cleaning will be an iterative process where to need several time to adjust the options and the order of the different steps in order to get good results. |
<<TAG(Advanced)>> == References == For more information on the SSP method, please consult the following publications: * C. D. Tesche, M. A. Uusitalo, R. J. Ilmoniemi, M. Huotilainen, M. Kajola, and O. Salonen, "Signal-space projections of MEG data characterize both distributed and well-localized neuronal sources," Electroencephalogr Clin Neurophysiol, vol. 95, pp. 189-200, 1995. * M. A. Uusitalo and R. J. Ilmoniemi, "Signal-space projection method for separating MEG or EEG into components," Med Biol Eng Comput, vol. 35, pp. 135-40, 1997. <<TAG(Advanced)>> |
| Line 252: | Line 237: |
| * Elekta: jumps in the sensors ? == Additional information == |
* Alpha and cardiac overlapping? |
Tutorial 12: Signal Space Projections (SSP)
Authors: Francois Tadel, Elizabeth Bock, John C Mosher, Sylvain Baillet
As previously said, the frequency filters are not adapted to remove artifacts that are transient or overlapping in frequency domain with the brain signals of interest. Other approaches exist to correct for those artifacts, based on the spatial signature of the artifacts.
If an event is very reproducible and occurs always at the same position (eg. eye blinks and heartbeats), the sensors will always record the same values when it occurs. We can identify the topographies corresponding to this artifact (ie. the spatial distributions of values at one time point) and remove them from the recordings. This spatial decomposition is the basic idea behind two widely used approaches: the SSP (Signal-Space Projection) and ICA (Independent Component Analysis) methods.
This introduction tutorial will focus on the SSP approach, as it is a lot simpler and faster but still very efficient for removing blinks and heartbeats from MEG and high-density EEG recordings. The interface for running ICA decompositions is very similar and will be described in an advanced tutorial.
Contents
Overview
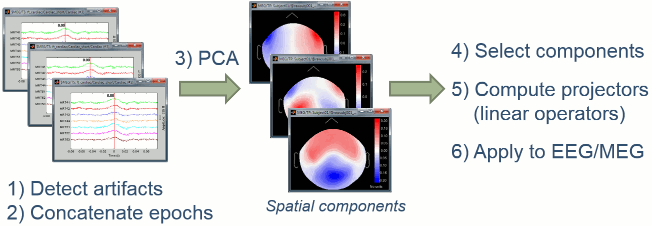
The general SSP objective is to identify the sensor topragraphies that are typical of a specific artifact, then to create spatial projectors to remove the contributions of these topographies from the recordings.
- We start by identifying many examples of the artifact we are trying to remove. This is what we've been doing in the previous tutorial with the creation of the "cardiac" and "blink" events.
- We extract a short time window around each of these event markers and concatenate in time all those small blocks of recordings.
- We run a principle components analysis (PCA) on the concatenated artifacts in order to get a decomposition in various spatial components (number of components = number of sensors).
- If it works well, we can find in the first few principal components some topographies that are very specific of the type of artifact we are targetting. We select those components to remove.
- We compute a linear projector for each spatial component to remove and save them in the database (in the "Link to raw file"). They are not applied immediately to the recordings.
Whenever some recordings are read from this file, the SSP projectors are applied on the fly to remove the artifact contributions. This approach is fast and memory efficient.
The order matters
This procedure has to be repeated separately for each artifact type. The order in which you process the artifacts matters, because for removing the second artifact we typically use the recordings cleaned with the first set of SSP projectors. We have to decide which one to process first.
It works best if each artifact is defined precisely and as independently as possible from the other artifacts. If the two artifacts happen simulateneously, the SSP projectors calculated for the blink may contain some of the heartbeat topography and vice versa. When trying to remove the second artifact, we might not be able to isolate it clearly anymore.
Because the heart beats every second or so, there is a high chance that when the subject blinks there is a heartbeat not too far away in the recordings. Therefore a significant number of the blinks will be contaminated with heartbeats. But we have usually a lot of "clean" heartbeats, we can start by removing those ones. To isolate correctly these two common artifacts, we recommend the following procedure:
- Remove the markers "cardiac" that are occurring during a blink (done in the previous tutorial),
Compute the cardiac SSP (with no eye movements, because we removed the co-occurring events),
Compute the blink SSP (with no heartbeats, because they've already been taken care of).
If you have multiple modalities recorded simultaneously, for example MEG and EEG, you should run twice this entire procedure, once for the EEG only and once for the MEG only. You will always get better results if you process the different types of sensors separately. Same thing when processing Elekta-Neuromag recordings: you should process separately the magnetometers (MEG MAG) and the gradiometers (MEG GRAD).
SSP: Heartbeats
Computation
Double-click on the link to show the MEG sensors for Run #01.
In the Record tab, select the menu: "Artifacts > SSP: Heartbeats".
Event name: Name of the event to use to calculate the projectors, enter "cardiac"
Sensor types: Type of sensors for which the projection should be calculated ("MEG"). Note that you will always get better results if you process the different types of sensors separately.
Compute using existing SSP projectors: You have the option to calculate the projectors from the raw recordings, or from the recordings filtered with the previously computed SSP projectors. Unless you have a good reason for not considering the existing projectors, you should select this option.
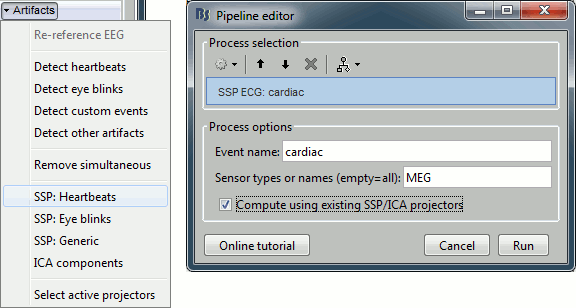
After the computation is done, a new figure is displayed, that lets you select the active projectors.
On the left: The projector categories where each row represents the result of an execution of this process (usually one for each sensor type and each artifact).
On the right: The spatial components returned by the PCA decomposition. The percentage indicated between brackets is the singular value for this each component, normalized for this decomposition (percentage = Si / sum(Si), see technical details at the end of this page).
Percentage: More practically, it indicates the amount of signal that was captured by the component during the decomposition. The highest it is, the more the component is representative of the artifact recordings that were used to calculate it. In the good cases, you would typically see one to three components with values that are significantly higher that the others.
- When a component is selected, it means that it is removed from the recordings. A spatial projector is computed and applied to the recordings on the fly when reading from the continuous file.
Default selection: The software selects the components with values superior to 12% and leaves the others unselected. This threshold is completely empirical and depends on the acquisition device, you should always review manually the components that you want to remove.
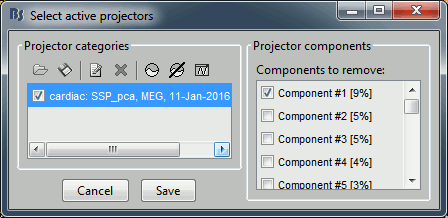
Evaluate the components
For the cardiac event, none of the components show a value superior to 12%, therefore the entire "cardiac" projector category was unselected. This doesn't mean that none of those components is actually interesting for us. The percentage indicated for the first value (9%) is much higher than the following ones (5%, 5%, 4%, 3%...), this could indicate that it targets relatively well the cardiac artifact. Let's investigate this in more details.
- Select the category ("cardiac") to enable the list on the right.
Click on the first component, then click on the toolbar button [Display component topography]. This menu shows the spatial distribution of the values recorded by the MEG sensors for this component. Note that you don't have to select the component (ie. check the box) to display it.
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
The second button "Display component topography [No magnetic interpolation]" produces the same figure but without the reinterpolation of the magnetic fields that is typically applied to the MEG recordings in Brainstorm. This procedure will be detailed later in the introdcution tutorials.
The last button in the toolbar [Display component time series], opens a figure that represents the evolution of the contribution of this component over time. The higher the value, the more present the selected topography in the recordings. Click on it to show the component #1, and display the ECG signal at the same time (right-click on the file > ECG > Display time series).
We observe that
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
- Do it again for components #2 and #3 (the components don't need to be active for that).
The other button [No interp] displays the same results without the magnetic field smoothing that is applied by default to MEG topographies. It may help understand some difficult cases.

Evaluate the correction
None of those topographies can be clearly related to a cardiac component. This can have two interpretations: the cardiac artifact is not very strong for this subject and the influence of the heart activity over the MEG sensors is completely buried in the noise or the brain signals, or the characterization of the artifact that we did was not so good and we should refine it by selecting for instance smaller time windows around the cardiac peaks. Here, it's probably due to the subject's morphology. Some people generate strong artifacts in the MEG, some don't.
In this case, it is safer to unselect this "cardiac" category, rather than removing randomly spatial components that we are not sure to identify.
SSP: Eye blinks
Let's try the same thing with the eye blinks. Select the process "SSP > Compute SSP: Eye blinks". Run it on the event type "blink", that indicate the peaks of the EOG signal.
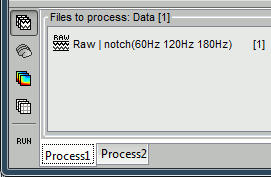
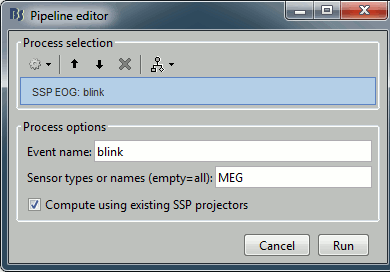
In this specific case, there is one value that is much higher than the others (21%), and it is selected by default because it is superior to 12%. The first component is most likely a good representation of the artifact we are trying to remove.
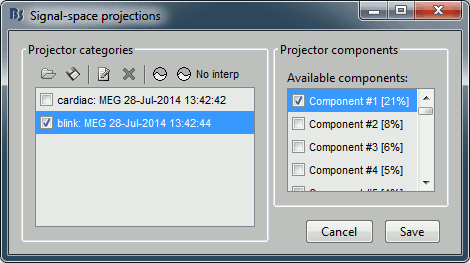
Select the cardiac category, and display the topographies for the first three components. Component #1 is clearly related with the ocular activity occurring during the blinks, the others are not as clear. Conclusion: the default proposition was good (only component #1 selected). Setting this first projector as active removes from the MEG signals the contributions related with the spatial topography we can see in the first figure, ie. a spatial distribution of sensor values related specifically to the eye blinks.
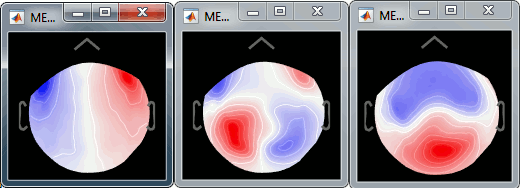
While the recordings are displayed in the file viewer, the selection of the active projectors interactively updates the recordings. Go to the first detected "blink" event (t=33.600s), and select the left frontal sensors (Shift+B or right-click > Display setup > CTF LF). Then play with the checkboxes to change the active projectors, and see how well this first component of the blink category removes the ocular artifact.
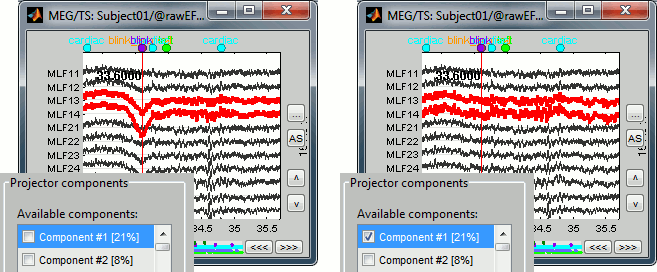
Select the projector category "blink", select the first component, and click on Save, to validate your changes. Starting from now, all the successive operations you apply on this file will integrate this eye blink correction, including the calculation of other projectors. After you closed this window, you can get back to it by selecting the menu "SSP > Select active projector" in the Record tab, and change again the selection of active projectors.
We are not going to calculate the SSP for the other category of events identified on the EOG, the group "blink2". The events classified in this group could represent eye saccades or other types of blinks, it is not very clear that they correspond to the same physiological event. Most of them do not seem to create any obvious contamination on the MEG recordings, there is probably no need to consider them now.
Run #02: Running from a script
Let's perform the same detection operations on Run #02, using this time the Process1 box.
- Close everything with the [X] button at the top-right corner of the Brainstorm window.
Select the run AEF #02 in the Process1 box, then select the following processes:
Events > Detect heartbeats: Select channel ECG, check "All file", event name "cardiac".
Events > Detect eye blinks: Select channel VEOG, check "All file", event name "blink".
Events > Remove simultaneous: Remove "cardiac", too close to "blink", delay 250ms.
Always check visually the results of the SSP cleaning executed from a script.
- Open the MEG sensors for Run #02.
In the Record tab, menu Artifacts > Select active projectors.
SSP: Generic
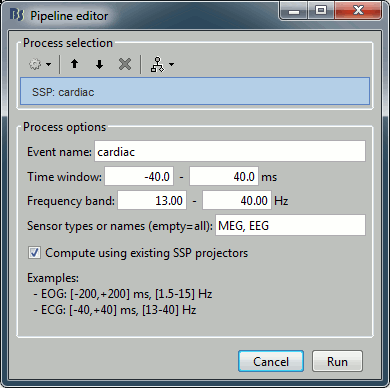
The calculation of the SSP for the heartbeats and the eye blinks are shortcuts to a more generic algorithm "Compute SSP: Generic". We do not need it here, but the illustration of its properties is interesting to understand the correction of ocular and cardiac artifacts:
Event name: Markers that are used to characterize the artifact
Time window: Time segment to consider before and after each event marker. We want this time window to be longer than the artifact effect itself, in order to have a large number of time samples representative of a normal brain activity. This helps the SVD decomposition to separate the artifact from the ongoing brain activity.
Frequency band: Definition of the band-pass filter that is applied to the recordings before calculating the projector. Usually you would use the same frequency band as we used for the detection, but you may want to try to refine this parameter if the results are not satisfying.
Sensor types or names: List of sensors for which to calculate the projector. You can get better results if you process one sensor type at a time.
Evaluation
Eye blinks
One efficient way of representing the impact of this artifact correction is to epoch the recordings around the artifacts before and after the correction. Use the same time window as the one used in the SSP options around each marker, and average all the segments of recordings. Those operations are detailed in the next tutorial, we are just presenting the results. You don't need to reproduce them at this time, just remember that it is doable, in case you need it with your own data. This is what the database would look like after those operations:
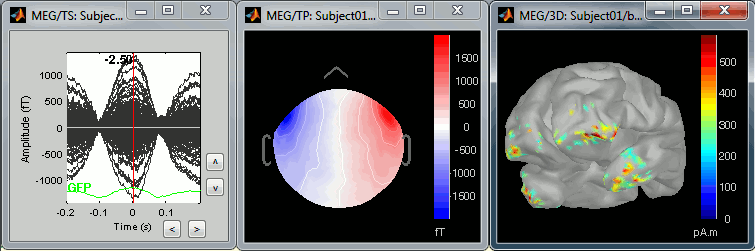
The following image represents the file "blink_uncorrected / Avg: blink", which is the average of the 18 identified blinks before the SSP correction, [-200,+200]ms around the "blink" events. The time series, the 2D topography at the peak (t = 0ms), and the mapping on the cortex of those fields, using the basic inverse model calculated in the previous tutorial.
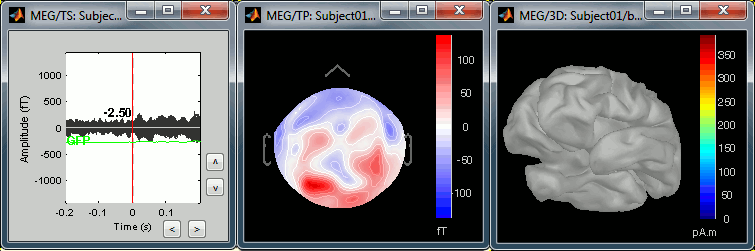
The next image represents the file "blink_ssp / Avg: blink", which is the exact same thing, but after the SSP correction. The artifact is gone. If you do this and you can still see some clear evoked component in the time series figure, the SSP correction was not efficient: the artifact is not properly identified, or you should select different components using the "Select active projectors" window.
Heartbeats
We can do the same thing with the heartbeats: epoch [-40,+40]ms around the "cardiac" events, average the trials, and review the event-related fields at three instants:
The peak of the P wave (t = -30ms):

The peak of the QRS complex (t = 0ms):
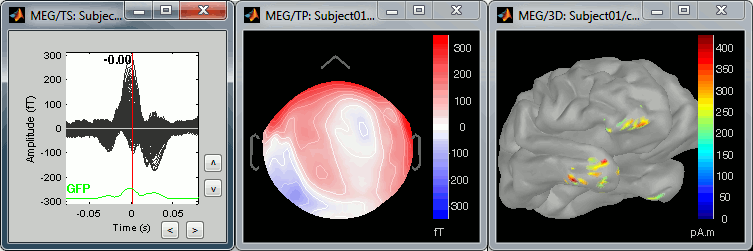
The peak of the T wave (t = 25ms):
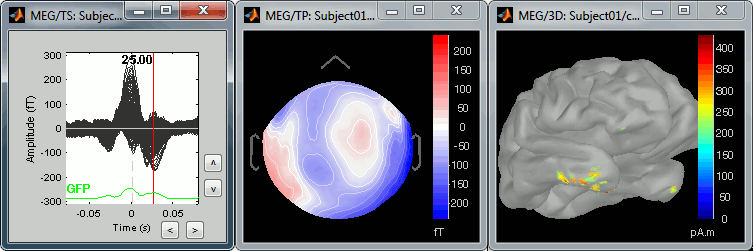
Those peaks may look big, but they are in fact much smaller (300 fT for the average of 346 repetitions) than what we observed for the eye blinks (1500 fT for 18 repetitions). You can notice that none of those topographies are similar to the components we obtained after calculating the SSP for the cardiac artifact. We don't know how to correct this artifact, it doesn't look too bad in terms of recordings contamination, so we just leave it uncorrected.
Troubleshooting
You have calculated the SSP as indicated here but you don't get any good results. No matter what you do, the topographies don't look like the targeted artifact. You can try the following:
- Review one by one the events indicating the artifacts, remove the ones that are less clear or that occur close to another artifact.
- Select or unselect the option "Compute using existing SSP".
- Use the process "Compute SSP: Generic" and modify some parameters:
- Reduce the time window around the peak of the artifact.
- Use a narrower frequency band: especially the EOG, if the projectors capture some of the alpha oscillations, you can limit the frequency band to [1.5 - 9] Hz.
- Change the method: Average / SSP.
- If you have multiple acquisition runs, you may try to use all the artifacts from all the runs rather than processing the files one by one. For that, use the Process2 tab instead of Process1. Put the "Link to raw file" of all the runs on both side, Files A (what is used to compute the SSP) and Files B (where the SSP are applied).
Always look at what this procedure gives you in output. Most of the time, the artifact cleaning will be an iterative process where to need several time to adjust the options and the order of the different steps in order to get good results.
Theory
The Signal-Space Projection (SSP) is one approach to rejection of external disturbances. Here is a short description of the method by Matti Hämäläinen, from the MNE 2.7 reference manual, section 4.16.
Unlike many other noise-cancellation approaches, SSP does not require additional reference sensors to record the disturbance fields. Instead, SSP relies on the fact that the magnetic field distributions generated by the sources in the brain have spatial distributions sufficiently different from those generated by external noise sources. Furthermore, it is implicitly assumed that the linear space spanned by the significant external noise patterns has a low dimension.
Without loss of generality we can always decompose any n-channel measurement b(t) into its signal and noise components as:
b(t) = bs(t) + bn(t)
Further, if we know that bn(t) is well characterized by a few field patterns b1...bm, we can express the disturbance as
bn(t) = Ucn(t) + e(t) ,
where the columns of U constitute an orthonormal basis for b1...bm, cn(t) is an m-component column vector, and the error term e(t) is small and does not exhibit any consistent spatial distributions over time, i.e., Ce = E{eeT} = I. Subsequently, we will call the column space of U the noise subspace. The basic idea of SSP is that we can actually find a small basis set b1...bm such that the conditions described above are satisfied. We can now construct the orthogonal complement operator
P⊥ = I - UUT
and apply it to b(t) yielding
b(t) ≈ P⊥bs(t) ,
since P⊥bn(t) = P⊥Ucn(t) ≈ 0. The projection operator P⊥ is called the signal-space projection operator and generally provides considerable rejection of noise, suppressing external disturbances by a factor of 10 or more. The effectiveness of SSP depends on two factors:
The basis set b1...bm should be able to characterize the disturbance field patterns completely and
The angles between the noise subspace space spanned by b1...bm and the signal vectors bs(t) should be as close to π/2 as possible.
If the first requirement is not satisfied, some noise will leak through because P⊥bn(t) ≠ 0. If the any of the brain signal vectors bs(t) is close to the noise subspace not only the noise but also the signal will be attenuated by the application of P⊥ and, consequently, there might by little gain in signal-to-noise ratio.
Since the signal-space projection modifies the signal vectors originating in the brain, it is necessary to apply the projection to the forward solution in the course of inverse computations.
Algorithm
We have now 346 examples of heartbeats, 18 examples of blinks, and 12 examples of saccades. Those are sufficient numbers of repetitions to calculate reliable signal-space projections. The logic of the computation is the following:
- Take a small time window around each marker to capture the full effect of the artifact, plus some clean brain signals before and after. The default time window is [-200,+200]ms for eye blinks, and [-40,+40]ms for the heartbeats.
- Filter the MEG or EEG signals in a frequency band of interest, in which the artifact is the most visible.
Concatenate all those time blocks into a big matrix A = [b1, ..., bm]
Compute the singular value decomposition of this matrix A: [U,S,V] = svd(A, 'econ')
The singular vectors Ui with the highest singular values Si are an orthonormal basis of the artifact subspace that we want to subtract from the recordings. The software selects by default the vectors with eigenvalues above a certain threshold: Si > 12% * sum(Si)
Then it is possible to redefine interactively the selected components.Calculate the projection operator: P⊥i = I - UiUiT
Apply this projection on the MEG or EEG recordings F: F = P⊥iF
The process has to be repeated separately several times:
- for each sensor type (EEG, MEG magnetometers, MEG gradiometers) and
- for each artifact, starting with the one that leads to the strongest contamination in amplitude on the sensors (in the case: the eye blinks)
- If you have difficulties removing one artifact or the other, you may try to process them in a different order. You may also try removing some of the artifact markers in the case of co-occurring artifacts. If a lot of the blinks happen at the same time as heartbeats, you may end up calculating projectors that mix both effects, but that do not remove efficiently one or the other. In this case, remove all the markers that happen in the segments contaminated by multiple artifacts.
Steps #1 to #5 are done automatically by the processes "SSP > Compute SSP" in the Record tab: the results, the vectors Ui, are saved in the database link ("Link to raw file") and in the channel file.
Steps #6 and #7 are calculated on the fly when reading a block of recordings from the continuous file: when using the raw viewer, running a process a process on the continuous file, or importing epochs in the database.
Step #8 is the manual control of the process. Take some time to understand what you are trying to remove and how to do it. Never trust blindly any fully automated artifact cleaning algorithm, always check manually what is removed from the recordings, and do not give up if the first results are not satisfying.
References
For more information on the SSP method, please consult the following publications:
- C. D. Tesche, M. A. Uusitalo, R. J. Ilmoniemi, M. Huotilainen, M. Kajola, and O. Salonen, "Signal-space projections of MEG data characterize both distributed and well-localized neuronal sources," Electroencephalogr Clin Neurophysiol, vol. 95, pp. 189-200, 1995.
- M. A. Uusitalo and R. J. Ilmoniemi, "Signal-space projection method for separating MEG or EEG into components," Med Biol Eng Comput, vol. 35, pp. 135-40, 1997.
Additional documentation
McGill guidelines
- SSP cookbook
- Alpha and cardiac overlapping?
