|
Size: 5631
Comment:
|
Size: 5609
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Bad channels and bad segments = ''Authors: Francois Tadel, Elizabeth Bock'' |
= Tutorial 11: Bad channels [Under construction] = ''Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet'' It is common during the acquisition to have a few sensors that are recording values that will not be usable in the data analysis. In MEG, a sensor can be damaged or unstable. In EEG, the quality of the connection between the electrode and the subject scalp is sometimes too low to record anything interesting. It is important to identify the sensors with poor signal quality at an early stage of the pre-processing, because the efficiency of the artifact removal will depend on it. If you try to remove blink and cardiac artifacts with some bad sensors it may not work very well, and worse, it will propagate the bad signals to all the channels. This tutorial will explain the various ways we have to handle the bad channels. Note that the recordings from this auditory experiment do not contain any bad sensors, therefore the entire tutorial is optional. If you are not interested, you can skip it and will still be able to follow the next tutorials. |
| Line 6: | Line 12: |
| = From auditory = | |
| Line 41: | Line 48: |
| * Note that if you click on a row in this window, it will select the corresponding channel in the time series and topography figures. | * Note that if you click on a row in this window, it will select the corresponding channel in the time series and topography figures. => TO CHECK |
| Line 52: | Line 59: |
| {{{ ADD BAD CHANNEL MONTAGE }}} = From auditory = == Bad segments == * At this point, you should review the entire files, by pages of a few seconds scrolling with the F3 key, to identify all the bad channels and the noisy segments of recordings. Do this with the the EOG channel open at the same time to identify saccades or blinks that were not completely corrected with the SSP projectors. As this is a complicated task that requires some expertise, we have prepared a list of bad segments for these datasets. * Open '''Run01'''. In the Record tab, select '''File > Add events from file''': * File name: sample_auditory/data/S01_AEF_20131218_01_notch/'''events_bad_01.mat ''' * File type: Brainstorm (events*.mat) * It adds '''12 bad segments''' to the file. |
== Montage: Bad channels == On the hard drive |
| Line 63: | Line 62: |
| * Open '''Run02'''. In the Record tab, select '''File > Add events from file''': * File name: sample_auditory/data/S01_AEF_20131218_02_notch/'''events_bad_02.mat ''' * File type: Brainstorm (events*.mat) * It adds '''9 bad segments''' and '''16 saccades''' to the file. <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/ArtifactsSsp&next=Tutorials/Epoching")>> |
<<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/ArtifactsFilter&next=Tutorials/ArtifactsDetect")>> |
Tutorial 11: Bad channels [Under construction]
Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet
It is common during the acquisition to have a few sensors that are recording values that will not be usable in the data analysis. In MEG, a sensor can be damaged or unstable. In EEG, the quality of the connection between the electrode and the subject scalp is sometimes too low to record anything interesting.
It is important to identify the sensors with poor signal quality at an early stage of the pre-processing, because the efficiency of the artifact removal will depend on it. If you try to remove blink and cardiac artifacts with some bad sensors it may not work very well, and worse, it will propagate the bad signals to all the channels.
This tutorial will explain the various ways we have to handle the bad channels. Note that the recordings from this auditory experiment do not contain any bad sensors, therefore the entire tutorial is optional. If you are not interested, you can skip it and will still be able to follow the next tutorials.
Bad channels
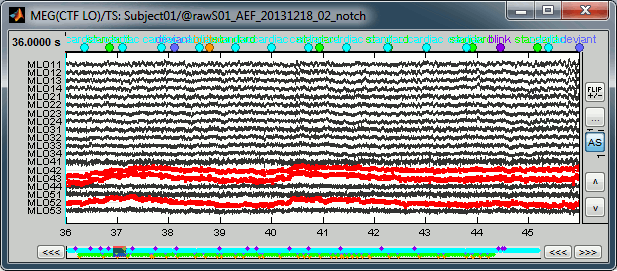
- During the visual exploration, some channels appeared generally noisier than the others. Example:
Right-click on Run01 > Good/bad channels > Mark some channels as bad
> MRT51, MLO52
Right-click on Run02 > Good/bad channels > Mark some channels as bad
> MRT51, MLO52, MLO42, MLO43
From CTF
Bad channels
If you find out that a sensor has unexpected values, that are not coherent with the other surrounding sensors, you can choose to ignore it in the displays and in the source estimation process. In the channel file, each channel has a flag which indicates if it is good or bad. You may also import recordings files that already have some bad channels defined.
Select a few channels with one of the method described above: click on the time series, click on the sensors dots, right-click and move to select a group of sensors. Then right-click in one of the figures and check out the Channels sub-menu:
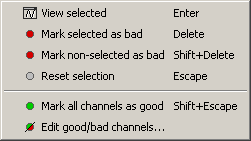
View selected: Show the time series of the selected sensors
Mark selected as bad: Remove sensors from the display and all the further computations
Mark non-selected as bad: Keep only the selected channels
Reset selection: Unselect all the selected sensors
Mark all channels as good: Brings back all the channels to display
Edit good/bad channels: Opens an interface that looks like the channel editor, but with one extra column to edit the status (good or bad) of each channel.
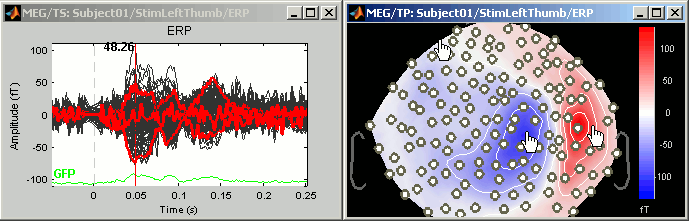
Mark channels as bad: Right-click > Channels > Mark selected as bad, or press Delete key. The sensors should disappear in all figures, and the topography view (2D sensors cap) is updated so that the interpolation on the 2D surface now ignores the bad channels.
Get the channels back: two options
Right-click on figure > Channels > Mark all channels as good
Right-click on figure > Channels > Edit good/bad channels... : this menu open a window very similar to the Channel Editor window introduced in previous tutorials, but without the annoying location and orientation values, and with green and red dots. Click on the dots to mark a channel as good or bad.

Note that if you click on a row in this window, it will select the corresponding channel in the time series and topography figures. => TO CHECK
- Close this window to save the changes.
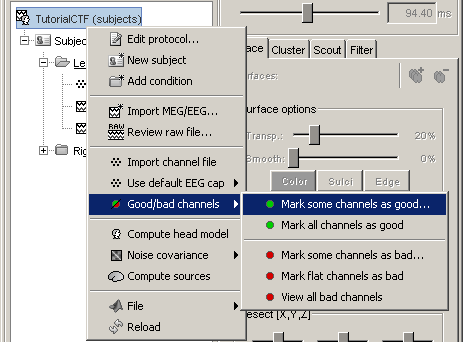
Batching this from the database explorer:
- You will find a "Channels" menu for any node in the tree that contains recordings.
- If you do this on the level of a node, the operation will be applied recursively to all the recordings contained in the node. This way, you can quickly set that the channel 63 is always bad, or that electrode 43 is not working for subject #12, without having to visualize all the recordings one after the other.
The View all bad channels command displays the list of all the bad channels in all the files in the Messages tab, in main Brainstorm window.
Important notes:
The good/bad channel flags are stored in the recordings files, not in the channel files. So if you marked some channels as bad in the ERF data file, there are still considered as good in the Std data file.
Montage: Bad channels
On the hard drive