|
Size: 9556
Comment:
|
Size: 7720
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Tutorial 13: Decoding Conditions = ''Author: Seyed-Mahdi Khaligh-Razavi, Francois Tadel, and Dimitrios Pantazis'' |
= Machine learning: Decoding / MVPA = ''Authors: Dimitrios Pantazis''', '''Seyed-Mahdi Khaligh-Razavi, Francois Tadel, '' |
| Line 4: | Line 4: |
| This set of functions allow you to do support vector machine (SVM) and linear discriminant analysis (LDA) classification on your MEG data across time. | This tutorial illustrates how to run MEG decoding using support vector machines (SVM). |
| Line 8: | Line 8: |
| == Description of the MEG data == The data contains the first 15 minutes of MEG recording for one of the subjects in the following study, and can be downloaded from here: https://www.dropbox.com/s/fmu5p3950e2mknc/mem6-0_tsss_mc.fif?dl=0 The study was conducted according to the Declaration of Helsinki and approved by the local ethics committee (Institutional Review Board of the Massachusetts Institute of Technology). Informed consent was obtained from all participants. During MEG recordings participants completed an orthogonal image categorization task. Participants saw 720 stimuli (360 faces, 360 scenes) centrally-presented ( 6 degree visual angle) for 0.5 second at a time and had to decide (without responding) if scenes were indoor / outdoor and faces were male / female. Every 2 to 4 trials (randomly determined), a question mark would appear on the screen for 1 second. At this time, participants were asked to press a button to indicate the category of the last image (male/female, indoor/outdoor), and they were also allowed to blink/swallow. This task was designed to ensure participants were attentive to the images without explicitly encoding them into memory. Participants only responded during the “question mark” trials so that activity during stimulus perception was not contaminated by motor activity. During the task, participants were asked to fixate on a cross centered on the screen and their eye movements were tracked, to ensure any results were not due to differential eye movement patterns. The whole experiment was divided into 16 runs. Each run contained 45 randomly selected images (each of them shown twice per run—not in succession), resulting in a total experiment time of about 50 min. <<BR>>The MEG data that is included here for demonstration purposes contains only the first 15 minutes of the whole session, during which 53 faces and 54 scenes have been observed. |
|
| Line 18: | Line 9: |
| This MEG decoding dataset remains a property of Aude Oliva’s Lab, Computer Science and AI (CSAIL), Massachusetts Institute of Technology, Cambridge, US. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the Lab. If you reference this dataset in your publications, please acknowledge its authors (Seyed-Mahdi Khaligh-Razavi, Francois Tadel, and Dimitrios Pantazis) and cite Brainstorm as indicated on the [[http://neuroimage.usc.edu/brainstorm/CiteBrainstorm|website]]. For questions, please contact us through the forum. |
To reference this dataset in your publications, please cite Cichy et al. (2014). |
| Line 23: | Line 12: |
| These set of functions allow you to do support vector machine (SVM) and linear discriminant analysis (LDA) classification on your MEG data across time. | Two decoding processes are available in Brainstorm: |
| Line 25: | Line 14: |
| '''Input''': the input is channel data from two conditions (e.g. condA and condB) across time. Number of samples per condition should be the same for both condA and condB. Each of them should at least contain two samples. | * Decoding > Classification with SVD * Decoding > Classification with max-correlation classifier |
| Line 27: | Line 17: |
| '''Output''': the output is a decoding curve across time, showing your decoding accuracy (decoding condA vs. condB) at timepoint 't'. | These two processes work in a similar way, but they use a different classifier, so only SVD is demonstrated here. |
| Line 29: | Line 19: |
| We have two condition types: faces, and scenes. We want to decode faces vs. scenes using 306 MEG channels. In the data, the faces are named as condition ‘201’; and the scenes are named as condition ‘203’. | * '''Input''': the input is the channel data from two conditions (e.g. condA and condB) across time. Number of samples per condition do not have to be the same for both condA and condB, but each of them should have enough samples to create k-folds (see parameter below). * '''Output''': the output is a decoding time course, or a temporal generalization matrix (train time x test time). * '''Classifier''': Two methods are offered for the classification of MEG recordings across time: support vector machine (SVM) and max-correlation classifier. |
| Line 31: | Line 23: |
| == Creating protocol and loading the data == Go to the file menu and create a new protocol called ''faces_vs_scenes ''with the options shown in the following figure. <<BR>><<BR>> |
In the context of this tutorial, we have two condition types: faces, and objects. We want to decode faces vs. objects using 306 MEG channels. |
| Line 34: | Line 25: |
| {{attachment:1-prot.png}} | == Download and installation == * From the [[http://neuroimage.usc.edu/bst/download.php|Download]] page of this website, download the file: '''subj04NN_sess01-0_tsss.fif''' |
| Line 36: | Line 28: |
| <<BR>> | * Start Brainstorm (Matlab scripts or stand-alone version). * Select the menu File > Create new protocol. Name it "'''TutorialDecoding'''" and select the options: * "'''Yes, use protocol's default anatomy'''", * "'''No, use one channel file per condition'''". <<BR>><<BR>> {{attachment:1_create_new_protocol.jpg||width="400"}} |
| Line 38: | Line 33: |
| After making your protocol, go to the functional data view (sorted by subjects). Then create a new subject ''(let's keep the default name: Subject01)'': to create a new subject right click on the protocol name; then press new subject, choose the default values shown in the following figure; press save. | == Import the recordings == * Go to the "functional data" view (sorted by subjects). * Right-click on the TutorialDecoding folder > New subject > '''Subject01''' <<BR>>Leave the default options you defined for the protocol. * Right click on the subject node (Subject01) > '''Review raw file'''''.'' <<BR>>Select the file format: "'''MEG/EEG: Neuromag FIFF (*.fif)'''"<<BR>>Select the file: sample_decoding/'''mem6-0_tsss_mc.fif''' <<BR>> {{attachment:2_review_raw_file.jpg||width="440"}} <<BR>><<BR>> {{attachment:2_review_raw_file2.jpg||width="440"}} <<BR>><<BR>> * Select "Event channels" to read the triggers from the stimulus channel. <<BR>><<BR>> {{attachment:3_event_channel.jpg||width="320"}} * Double click on the 'Link to raw file' to visualize the raw recordings. Event codes 13-24 indicate responses to face images, and we will combine them to a single group called 'faces'. To do so, select events 13-24 and from the menu select Events->Duplicate groups. Then select Events->Merge groups. * The event codes are duplicated first so we do not lose the original 13-24 event codes. |
| Line 40: | Line 41: |
| <<BR>><<BR>> | {{attachment:4_duplicate_groups_faces.jpg||width="500"}} <<BR>><<BR>> |
| Line 42: | Line 43: |
| {{attachment:2_subj.png}} | {{attachment:5_merge_groups_faces.jpg||width="500"}} |
| Line 44: | Line 45: |
| == Import MEG data == 1. Right click on the subject node (Subject01), and select ‘''review raw file’.'' 1. Change the file format to ‘''MEG/EEG: Neuromag FIF (*.fif)''’; then select the .fif file (''mem6-0_tsss_mc.fif'') you downloaded. 1. Press event channels <<BR>> <<BR>> {{attachment:3_fif.png}} <<BR>><<BR>> 1. In the ‘''mem6-0 tsss mc’'' node right click on ‘''Link to raw file''’; select ‘Import in database’. 1. Set epoch time to [-100, 1000] 1. Time range: [-100,0] 1. From the events select only these two events: 201 (faces), and 203(scenes) <<BR>> <<BR>> {{attachment:4_importfif.png}} <<BR>><<BR>> 1. Then press import. You will get a message saying ‘some epochs are shorter than the others..’ . Press yes. |
* We will not pay attention to MEG/MRI registration because we are not going to compute any source models. The decoding is done on the sensor data. * Right-click on the "Link to raw file" > '''Import in database'''. <<BR>><<BR>> {{attachment:decoding_import.gif||height="317",width="571"}} * Select only two events: 201 (faces) and 203(scenes) * Epoch time: [-100, 1000] ms * Remove DC offset: Time range: [-100, 0] ms * Do not create separate folders for each event type * You will get a message saying "some epochs are shorter than the others". Answer '''yes'''. |
| Line 54: | Line 53: |
| == Decoding conditions == 1. Select ‘Process2’ from the bottom brainstorm window. 1. Drag and drop 40 files from folder ‘201’ to ‘Files A’; and 40 files from folder ‘203’ to ‘Files B’. You can select more than 40 or less. The important thing is that both ‘A’ and ‘B’ should have the same number of files. <<BR>><<BR>> {{attachment:5_dragdrop.png}} <<BR>><<BR>> |
== Select files == Select the Process2 tab at the bottom of the Brainstorm window. |
| Line 58: | Line 56: |
| === a) Cross-validation === | * Drag and drop '''40 files''' from group '''201''' to the left (Files A). * Drag and drop '''40 files''' from group '''203''' to the right (Files B). * You can select more than 40 or less. The important thing is that both ‘A’ and ‘B’ should have the same number of files. <<BR>><<BR>> {{attachment:decoding_selectfiles.gif||height="369",width="398"}} == Cross-validation == |
| Line 61: | Line 63: |
| 1. Select run -> decoding conditions -> classification with cross validation <<BR>><<BR>> {{attachment:6_decodingcrossval.png}} <<BR>><<BR>> Here, you will have three choices for cross-validation. If you have Matlab Statistics and Machine Learning Toolbox, you can use ‘Matlab SVM’ or ‘Matlab LDA’. You can also install the ‘LibSVM’ toolbox (https://www.csie.ntu.edu.tw/~cjlin/libsvm/ ); and addpath it to your Matlab session. LibSVM may be faster. The LibSVM cross-validation won’t be stratified. However, if you select Matlab SVM/LDA, it will do a k-fold stratified cross-validation for you, meaning that each fold will contain the same proportions of the two types of class labels. | * Select process "'''Decoding > Classification with cross-validation'''": <<BR>><<BR>> {{attachment:cv_process.gif||height="388",width="346"}} * '''Low-pass cutoff frequency''': If set, it will apply a low-pass filter to all the input recordings. * '''Matlab SVM/LDA''': Require Matlab's Statistics and Machine Learning Toolbox. <<BR>>These methods do a k-fold stratified cross-validation for you, meaning that each fold will contain the same proportions of the two types of class labels (option "Number of folds"). * '''LibSVM''': Requires the LibSVM toolbox ([[http://www.csie.ntu.edu.tw/~cjlin/libsvm/#download|download]] and add to your path).<<BR>>The LibSVM cross-validation may be faster but will not be stratified. |
| Line 63: | Line 68: |
| You can also set the number of folds for cross-validation; and the cut-off frequency for low-pass filtering – this is to smooth your data. | * The process will take some time. The results are then saved in a file in the new decoding folder. <<BR>><<BR>> {{attachment:cv_file.gif||height="167",width="268"}} * If you double click on it you will see a decoding curve across time. <<BR>><<BR>> {{attachment:cv_plot.gif||height="172",width="355"}} |
| Line 65: | Line 71: |
| . 2- To continue, set the values as shown below: <<BR>><<BR>> {{attachment:7_crossvaloptions.png}} <<BR>><<BR>> 3- The process will take some time.The results are then saved in a file (‘Matlab SVM Decoding_201_203’) under the new ‘decoding’ node. If you double click on it you will see a decoding curve across time (shown below). <<BR>><<BR>> {{attachment:8_crossvalres1.png}} <<BR>><<BR>> {{attachment:9_crossvalres2.png}} <<BR>><<BR>> The internal brainstorm plot is not perfect for this purpose. To make a proper plot you can plot the results yourself. Right click on the result file (Matlab SVM Decoding_201_203), select 'export to Matlab' from the menu. Give it a name ‘decodingcurve’. Then using Matlab plot function you can plot the decoding accuracies across time. ''<<BR>>'' ''Matlab code: <<BR>>'' * ''figure; '' * ''plot(decodingcurve.Value);'' <<BR>><<BR>> {{attachment:10_crossvalres3.png}} <<BR>><<BR>> === b) Permutation === |
== Permutation == |
| Line 81: | Line 74: |
| 1. Select run -> decoding conditions -> classification with permutation 1. Set the values as the following: |
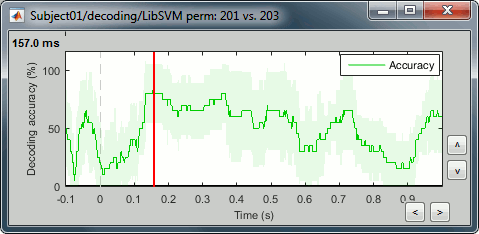
* Select process "'''Decoding > Classification with cross-validation'''". Set options as below: <<BR>><<BR>> {{attachment:perm_process.gif||height="366",width="311"}} * '''Trial bin size''': If greater than 1, the training data will be randomly grouped into bins of the size you determine here. The samples within each bin are then averaged (we refer to this as sub-averaging); the classifier is then trained using the averaged samples. For example, if you have 40 faces and 40 scenes, and you set the trial bin size to 5; then for each condition you will have 8 bins each containing 5 samples. Seven bins from each condition will be used for training, and the two left-out bins (one face bin, one scene bin) will be used for testing the classifier performance. * The results are saved in a file in the new decoding folder. <<BR>><<BR>> {{attachment:perm_file.gif||height="142",width="284"}} * Right-click > Display as time series (or double click). <<BR>><<BR>> {{attachment:perm_plot.gif||height="169",width="346"}} |
| Line 84: | Line 79: |
| <<BR>><<BR>> {{attachment:11_permoptions.png}} <<BR>><<BR>> If the ‘Trial bin size’ is greater than 1, the training data will be randomly grouped into bins of the size you determine here. The samples within each bin are then averaged (we refer to this as sub-averaging); the classifier is then trained using the averaged samples. For example, if you have 40 faces and 40 scenes, and you set the trial bin size to 5; then for each condition you will have 8 bins each containing 5 samples. Seven bins from each condition will be used for training, and the two left-out bins (one face bin, one scene bin) will be used for testing the classifier performance. . 3- The results are saved under the ‘decoding ’ node . If you selected ‘Matlab SVM’, the file name will be ‘Matlab SVM Decoding-permutation_201_203’ . Double click on it. You will get the plot blow: <<BR>><<BR>> {{attachment:12_permres1.png}} <<BR>><<BR>> This might not be very intuitive. You can export the decoding results into Matlab and plot it yourself. If you export the decoding results into Matlab, the imported structure will have two important fields: * a) Value: this is the mean decoding accuracy across all permutations * b) Std: this is the standard deviation across all permutations. If you plot the mean value (decodingcurve.Value), below is what you will get. You also have access to the standard deviation (decodingcurve.Std), in case you want to plot it. <<BR>><<BR>> {{attachment:13_permres2.png}} <<BR>><<BR>> |
== Acknowledgment == This work was supported by the ''McGovern Institute Neurotechnology Program'' to PIs: Aude Oliva and Dimitrios Pantazis. ''http://mcgovern.mit.edu/technology/neurotechnology-program'' |
| Line 102: | Line 83: |
| 1. Khaligh-Razavi, S-M; Bainbridge, W; Pantazis, D; and Oliva, A. (2015) Introducing Brain Memorability: an MEG study (in preparation). 1. |
1. Khaligh-Razavi SM, Bainbridge W, Pantazis D, Oliva A (2016)<<BR>>[[http://biorxiv.org/content/early/2016/04/22/049700.abstract|From what we perceive to what we remember: Characterizing representational dynamics of visual memorability]]. ''bioRxiv'', 049700. 1. Cichy RM, Pantazis D, Oliva A (2014)<<BR>>[[http://www.nature.com/neuro/journal/v17/n3/full/nn.3635.html|Resolving human object recognition in space and time]], Nature Neuroscience, 17:455–462. == Additional documentation == * Forum: Decoding in source space: http://neuroimage.usc.edu/forums/showthread.php?2719 <<EmbedContent(http://neuroimage.usc.edu/bst/get_feedback.php?Tutorials/Decoding)>> |
Machine learning: Decoding / MVPA
Authors: Dimitrios Pantazis, Seyed-Mahdi Khaligh-Razavi, Francois Tadel,
This tutorial illustrates how to run MEG decoding using support vector machines (SVM).
Contents
License
To reference this dataset in your publications, please cite Cichy et al. (2014).
Description of the decoding functions
Two decoding processes are available in Brainstorm:
Decoding > Classification with SVD
Decoding > Classification with max-correlation classifier
These two processes work in a similar way, but they use a different classifier, so only SVD is demonstrated here.
Input: the input is the channel data from two conditions (e.g. condA and condB) across time. Number of samples per condition do not have to be the same for both condA and condB, but each of them should have enough samples to create k-folds (see parameter below).
Output: the output is a decoding time course, or a temporal generalization matrix (train time x test time).
Classifier: Two methods are offered for the classification of MEG recordings across time: support vector machine (SVM) and max-correlation classifier.
In the context of this tutorial, we have two condition types: faces, and objects. We want to decode faces vs. objects using 306 MEG channels.
Download and installation
From the Download page of this website, download the file: subj04NN_sess01-0_tsss.fif
- Start Brainstorm (Matlab scripts or stand-alone version).
Select the menu File > Create new protocol. Name it "TutorialDecoding" and select the options:
"Yes, use protocol's default anatomy",
"No, use one channel file per condition".
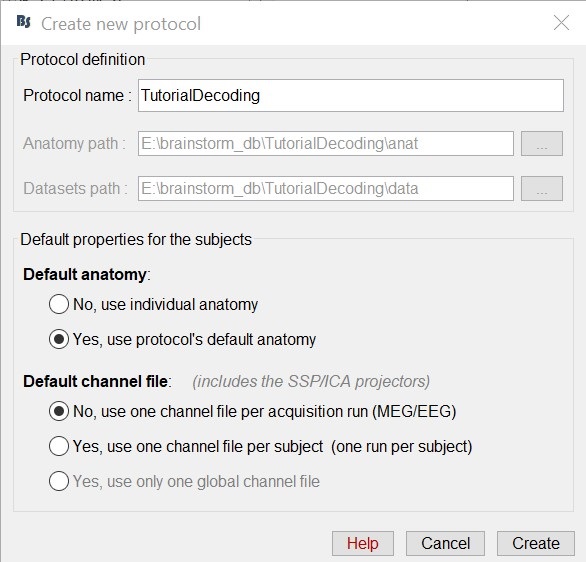
Import the recordings
- Go to the "functional data" view (sorted by subjects).
Right-click on the TutorialDecoding folder > New subject > Subject01
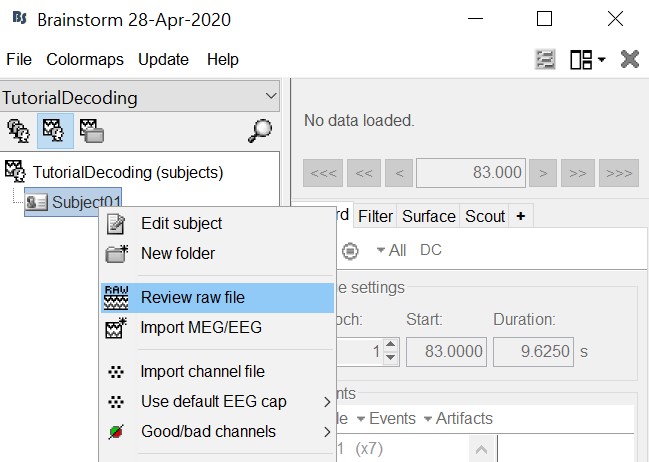
Leave the default options you defined for the protocol.Right click on the subject node (Subject01) > Review raw file.
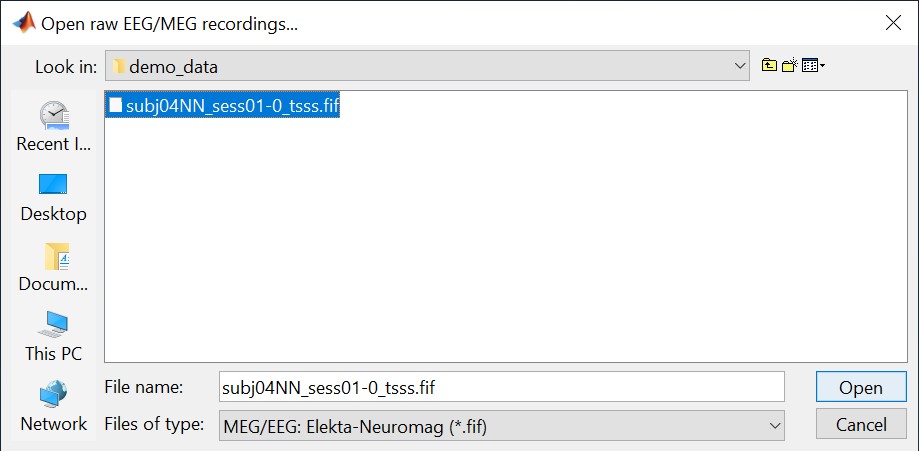
Select the file format: "MEG/EEG: Neuromag FIFF (*.fif)"
Select the file: sample_decoding/mem6-0_tsss_mc.fif
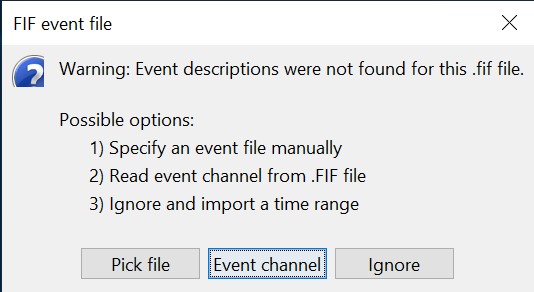
Select "Event channels" to read the triggers from the stimulus channel.
Double click on the 'Link to raw file' to visualize the raw recordings. Event codes 13-24 indicate responses to face images, and we will combine them to a single group called 'faces'. To do so, select events 13-24 and from the menu select Events->Duplicate groups. Then select Events->Merge groups.
- The event codes are duplicated first so we do not lose the original 13-24 event codes.
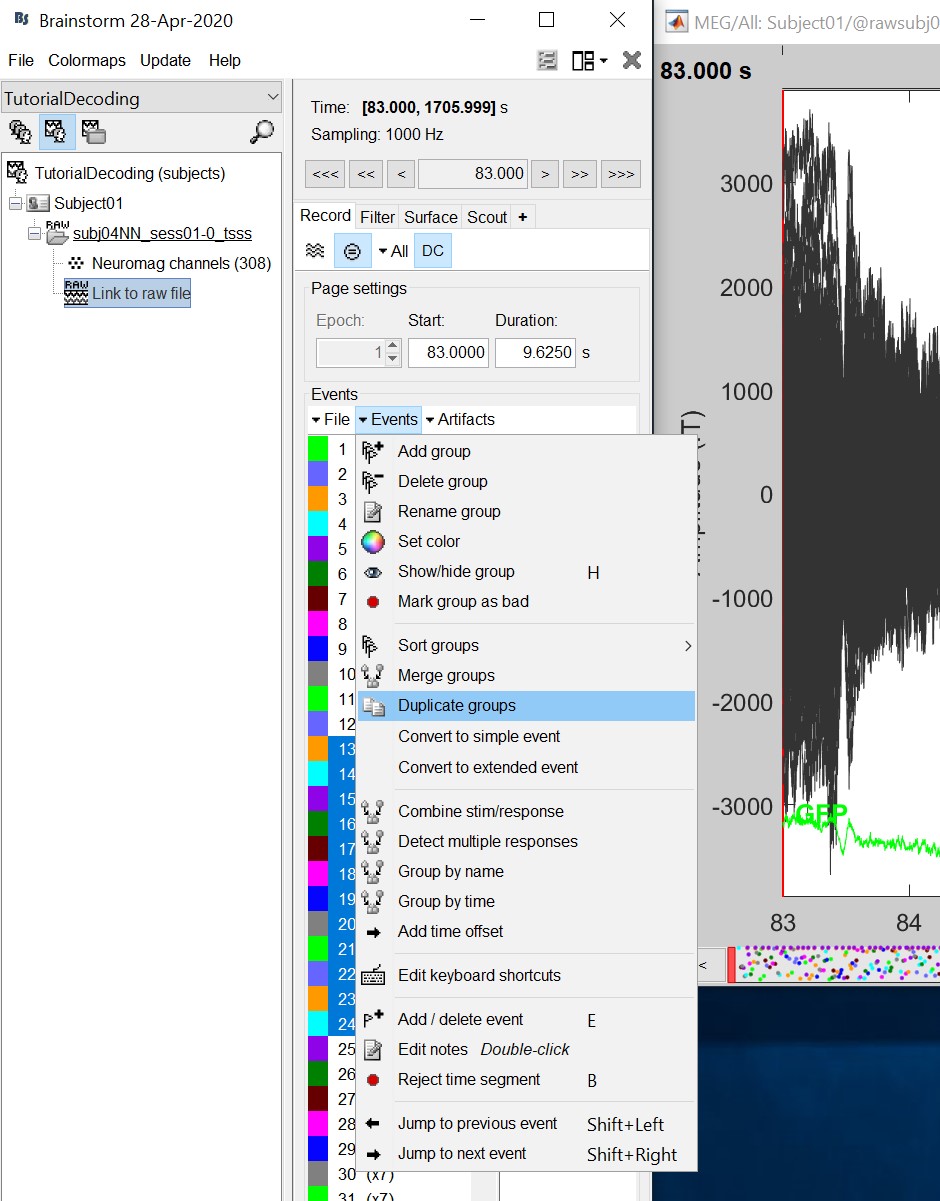
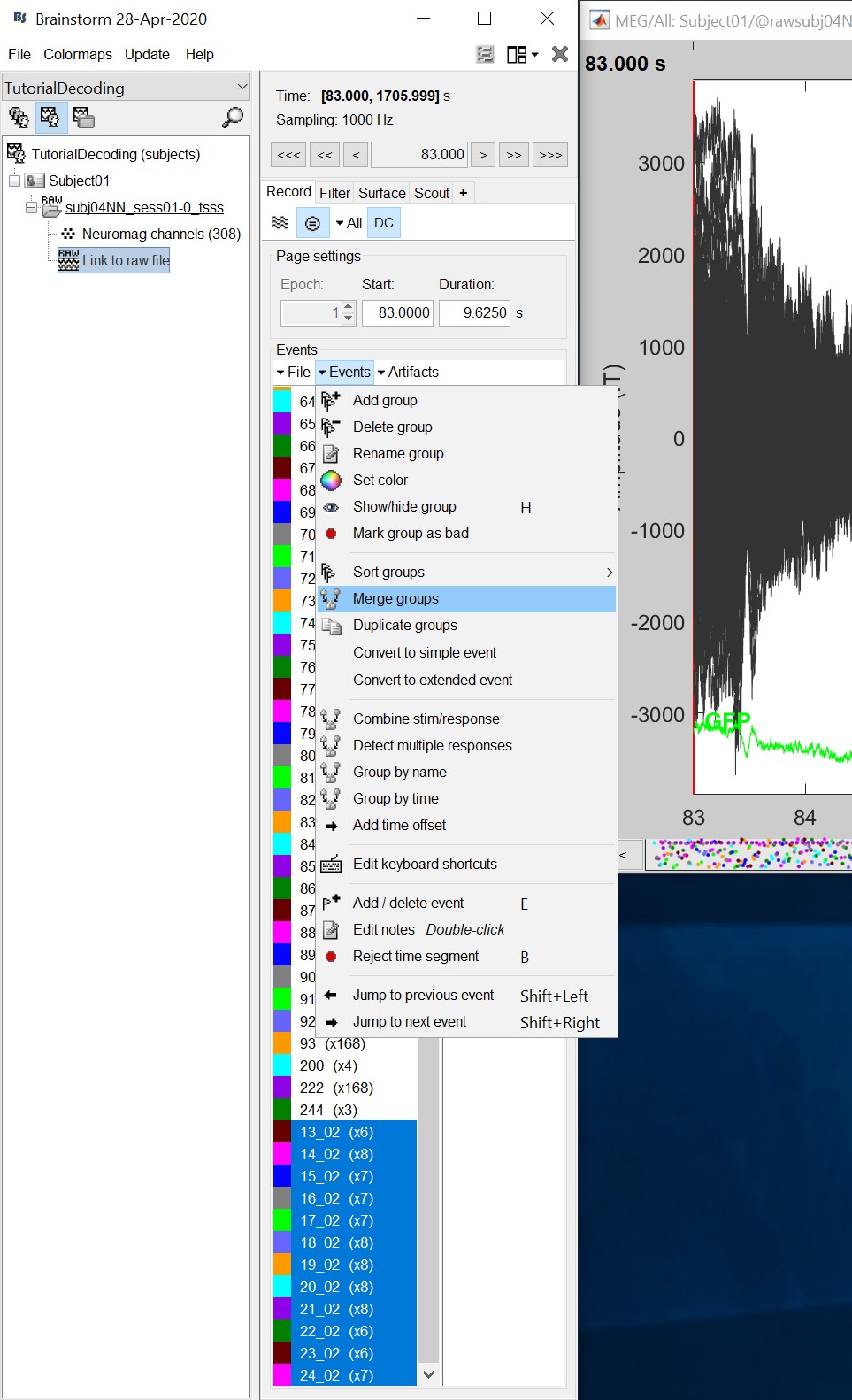
- We will not pay attention to MEG/MRI registration because we are not going to compute any source models. The decoding is done on the sensor data.
Right-click on the "Link to raw file" > Import in database.
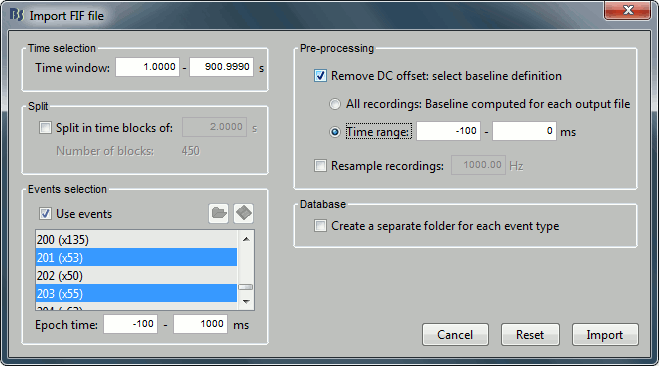
- Select only two events: 201 (faces) and 203(scenes)
- Epoch time: [-100, 1000] ms
- Remove DC offset: Time range: [-100, 0] ms
- Do not create separate folders for each event type
You will get a message saying "some epochs are shorter than the others". Answer yes.
Select files
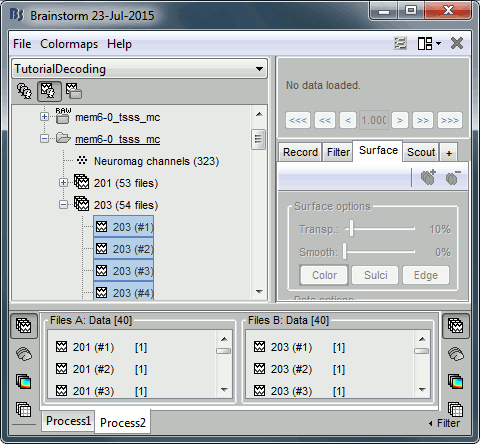
Select the Process2 tab at the bottom of the Brainstorm window.
Drag and drop 40 files from group 201 to the left (Files A).
Drag and drop 40 files from group 203 to the right (Files B).
You can select more than 40 or less. The important thing is that both ‘A’ and ‘B’ should have the same number of files.
Cross-validation
Cross-validation is a model validation technique for assessing how the results of our decoding analysis will generalize to an independent data set.
Select process "Decoding > Classification with cross-validation":
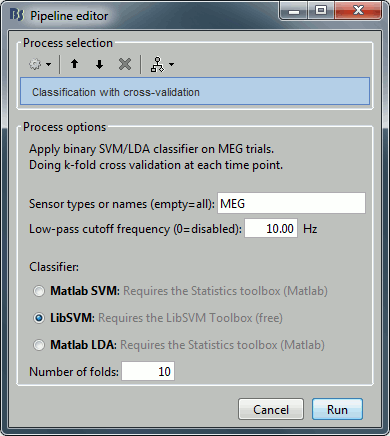
Low-pass cutoff frequency: If set, it will apply a low-pass filter to all the input recordings.
Matlab SVM/LDA: Require Matlab's Statistics and Machine Learning Toolbox.
These methods do a k-fold stratified cross-validation for you, meaning that each fold will contain the same proportions of the two types of class labels (option "Number of folds").LibSVM: Requires the LibSVM toolbox (download and add to your path).
The LibSVM cross-validation may be faster but will not be stratified.
The process will take some time. The results are then saved in a file in the new decoding folder.
If you double click on it you will see a decoding curve across time.
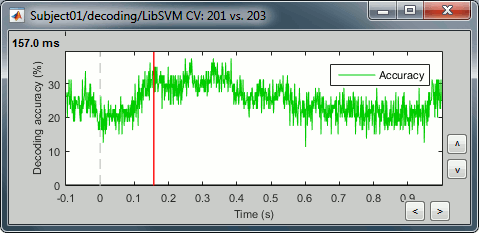
Permutation
This is an iterative procedure. The training and test data for the SVM/LDA classifier are selected in each iteration by randomly permuting the samples and grouping them into bins of size n (you can select the trial bin sizes). In each iteration two samples (one from each condition) are left out for test. The rest of the data are used to train the classifier with.
Select process "Decoding > Classification with cross-validation". Set options as below:
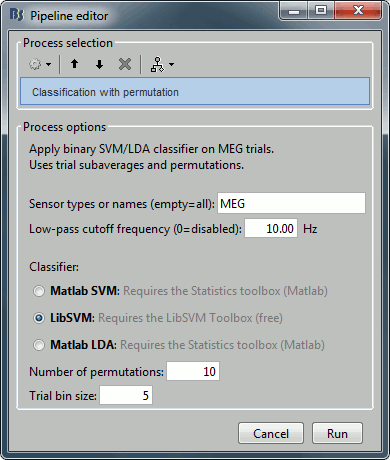
Trial bin size: If greater than 1, the training data will be randomly grouped into bins of the size you determine here. The samples within each bin are then averaged (we refer to this as sub-averaging); the classifier is then trained using the averaged samples. For example, if you have 40 faces and 40 scenes, and you set the trial bin size to 5; then for each condition you will have 8 bins each containing 5 samples. Seven bins from each condition will be used for training, and the two left-out bins (one face bin, one scene bin) will be used for testing the classifier performance.
The results are saved in a file in the new decoding folder.
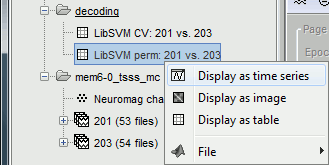
Right-click > Display as time series (or double click).
Acknowledgment
This work was supported by the McGovern Institute Neurotechnology Program to PIs: Aude Oliva and Dimitrios Pantazis. http://mcgovern.mit.edu/technology/neurotechnology-program
References
Khaligh-Razavi SM, Bainbridge W, Pantazis D, Oliva A (2016)
From what we perceive to what we remember: Characterizing representational dynamics of visual memorability. bioRxiv, 049700.Cichy RM, Pantazis D, Oliva A (2014)
Resolving human object recognition in space and time, Nature Neuroscience, 17:455–462.
Additional documentation
Forum: Decoding in source space: http://neuroimage.usc.edu/forums/showthread.php?2719
