|
Size: 7691
Comment:
|
Size: 10215
Comment: Events for object images are 49-60 and not 49-59
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Decoding conditions = ''Authors: Seyed-Mahdi Khaligh-Razavi, Francois Tadel, Dimitrios Pantazis'' |
= Machine learning: Decoding / MVPA = ''Authors: Dimitrios Pantazis''', '''Seyed-Mahdi Khaligh-Razavi, Francois Tadel, '' |
| Line 4: | Line 4: |
| This tutorial illustrates how to use the functions developed at Aude Oliva's lab (MIT, CSAIL), and McGovern's MEG lab (Dimitrios Pantazis's lab) to run support vector machine (SVM) and linear discriminant analysis (LDA) classification on your MEG data across time. | This tutorial illustrates how to run MEG decoding (a type of multivariate pattern analysis / MVPA) using support vector machines (SVM). |
| Line 8: | Line 8: |
| <<Include(DatasetDecoding, , from="\<\<HTML\(\<!-- START-PAGE --\>\)\>\>", to="\<\<HTML\(\<!-- STOP-SHORT --\>\)\>\>")>> | == License == To reference this dataset in your publications, please cite Cichy et al. (2014). |
| Line 13: | Line 14: |
| * Decoding > Classification with cross-validation (process_decoding_crossval.m) * Decoding > Classification with permutation (process_decoding_permutation.m) |
* Decoding -> SVM classifier decoding * Decoding -> max-correlation classifier decoding |
| Line 16: | Line 17: |
| These two processes work in a similar way: | These two processes work in a similar way, but they use a different classifier, so only SVM is demonstrated here. |
| Line 18: | Line 19: |
| * '''Input''': the input is the channel data from two conditions (e.g. condA and condB) across time. Number of samples per condition should be the same for both condA and condB. Each of them should at least contain two samples. * '''Output''': the output is a decoding curve across time, showing your decoding accuracy (decoding condA vs. condB) at time point 't'. * '''Classifier''': Two methods are offered for the classification of MEG recordings across time: Support vector machine (SVM) and Linear discriminant analysis (LDA). |
* '''Input''': the input is the time recording data from two conditions (e.g. condA and condB) across time. Number of samples per condition do not have to be the same for both condA and condB, but each of them should have enough samples to create k-folds (see parameter below). * '''Output''': the output is a decoding time course, or a temporal generalization matrix (train time x test time). * '''Classifier''': Two methods are offered for the classification of MEG recordings across time: support vector machine (SVM) and max-correlation classifier. |
| Line 22: | Line 23: |
| In the context of this tutorial, we have two condition types: faces, and scenes. We want to decode faces vs. scenes using 306 MEG channels. In the data, the faces are named as condition ‘201’; and the scenes are named as condition ‘203’. | In the context of this tutorial, we have two condition types: faces, and objects. The participant was shown different types of images and we want to decode the face images vs. the object images using 306 MEG channels. |
| Line 25: | Line 26: |
| * Go to the [[http://neuroimage.usc.edu/bst/download.php|Download]] page of this website, and download the file: '''sample_decoding.zip ''' * Unzip it in a folder that is not in any of the Brainstorm folders (program folder or database folder). This is really important that you always keep your original data files in a separate folder: the program folder can be deleted when updating the software, and the contents of the database folder is supposed to be manipulated only by the program itself. |
* From the [[http://neuroimage.usc.edu/bst/download.php|Download]] page of this website, download the file 'sample_decoding.zip', and unzip to get the MEG file: '''subj04NN_sess01-0_tsss.fif''' * SVM decoding requires the LibSVM toolbox. The SVM decoding process tries to automatically install it, but should it fail here's how you install it manually: * [[https://www.csie.ntu.edu.tw/~cjlin/libsvm/#download|Download]] the libsvm ZIP file here. * Extract the content of the ZIP file to a permanent folder. * On Linux or Mac, you need to compile the library. Mac users can use XCode to do so. * On Windows, you can use windows subfolder of the zip file. * Add the path to the LibSVM folder in Matlab using addpath. |
| Line 28: | Line 34: |
| * Start Brainstorm (Matlab scripts or stand-alone version). * Select the menu File > Create new protocol. Name it "'''TutorialDecoding'''" and select the options: |
* Start Brainstorm. * Select the menu File -> Create new protocol. Name it "'''TutorialDecoding'''" and select the options: |
| Line 31: | Line 37: |
| * "'''No, use one channel file per condition'''". <<BR>><<BR>> {{attachment:decoding_protocol.gif||height="370",width="344"}} | * "'''No, use one channel file per acquisition run'''". <<BR>><<BR>> {{attachment:1_create_new_protocol.jpg||width="400"}} |
| Line 34: | Line 40: |
| * Go to the "functional data" view (sorted by subjects). * Right-click on the TutorialDecoding folder > New subject > '''Subject01''' <<BR>>Leave the default options you defined for the protocol. * Right click on the subject node (Subject01) > '''Review raw file'''''.'' <<BR>>Select the file format: "'''MEG/EEG: Neuromag FIFF (*.fif)'''"<<BR>>Select the file: sample_decoding/'''mem6-0_tsss_mc.fif''' <<BR>><<BR>> {{attachment:decoding_link.gif||height="169",width="547"}} * Select "Event channels" to read the triggers from the stimulus channel. <<BR>><<BR>> {{attachment:decoding_events.gif||height="162",width="289"}} * We will not pay much attention to MEG/MRI registration because we are not going to compute any source models, the decoding is done on the sensor data. * Right-click on the "Link to raw file" > '''Import in database'''. <<BR>><<BR>> {{attachment:decoding_import.gif||height="317",width="571"}} * Select only two events: 201 (faces) and 203(scenes) * Epoch time: [-100, 1000] ms * Remove DC offset: Time range: [-100, 0] ms * Do not create separate folders for each event type * You will get a message saying "some epochs are shorter than the others". Answer '''yes'''. |
* Go to the "functional data" view of the database (sorted by subjects). * Right-click on the TutorialDecoding folder -> New subject -> '''Subject01''' <<BR>>Leave the default options you defined for the protocol. * Right click on the subject node (Subject01) -> '''Review raw file'''''.'' <<BR>>Select the file format: "'''MEG/EEG: Neuromag FIFF (*.fif)'''"<<BR>>Select the file: '''subj04NN-sess01-0_tsss.fif''' <<BR>> * {{attachment:2_review_raw_file.jpg||width="440"}} <<BR>><<BR>> {{attachment:2_review_raw_file2.jpg||width="440"}} <<BR>><<BR>> * Select "Event channels" to read the triggers from the stimulus channel. <<BR>><<BR>> {{attachment:3_event_channel.jpg||width="320"}} * We will not pay attention to MEG/MRI registration because we are not going to compute any source models. The decoding is done on the sensor data. * Double click on the 'Link to raw file' to visualize the raw recordings. Event codes 13-24 indicate responses to face images, and we will combine them to a single group called 'faces'. To do so, select events 13-24 using SHIFT + click and from the menu select "'''Events -> Duplicate groups'''". Then select "'''Events -> Merge groups'''". The event codes are duplicated first so we do not lose the original 13-24 event codes. {{attachment:4_duplicate_groups_faces.jpg||width="500"}} <<BR>><<BR>> {{attachment:5_merge_groups_faces.jpg||width="500"}} * Event codes 49-60 indicate responses to object images, and we will combine them to a single group called 'objects'. To do so, select events 49-60 (SHIFT + click) and from the menu select Events -> Duplicate groups. Then select Events -> Merge groups. No screenshots are shown since this is similar to above. * We will now import the 'faces' and 'objects' responses to the database. Select "'''File -> Import in database"'''. <<BR>><<BR>> {{attachment:10_import_in_database.jpg||width="500"}} <<BR>><<BR>> * Select only two events: 'faces' and 'objects' * Epoch time: [-200, 800] ms * Remove DC offset: Time range: [-200, 0] ms * Do not create separate folders for each event type<<BR>> {{attachment:11_import_in_database_window.jpg||width="500"}} |
| Line 47: | Line 55: |
| Select the Process2 tab at the bottom of the Brainstorm window. | * Drag and drop all the face and object trials to the Process1 tab at the bottom of the Brainstorm window. * Intuitively, you might have expected to use the Process2 tab to decode faces vs. objects. But the decoding process is designed to also handle pairwise decoding of multiple classes (not just two classes) for computational efficiency, so more that two categories can be entered in the Process1 tab.<<BR>> {{attachment:12_select_files.jpg||width="400"}} |
| Line 49: | Line 58: |
| * Drag and drop '''40 files''' from group '''201''' to the left (Files A). * Drag and drop '''40 files''' from group '''203''' to the right (Files B). * You can select more than 40 or less. The important thing is that both ‘A’ and ‘B’ should have the same number of files. <<BR>><<BR>> {{attachment:decoding_selectfiles.gif||height="369",width="398"}} == Cross-validation == |
== Decoding with cross-validation == |
| Line 56: | Line 61: |
| * Select process "'''Decoding > Classification with cross-validation'''": <<BR>><<BR>> {{attachment:cv_process.gif||height="388",width="346"}} * '''Low-pass cutoff frequency''': If set, it will apply a low-pass filter to all the input recordings. * '''Matlab SVM/LDA''': Require Matlab's Statistics and Machine Learning Toolbox. <<BR>>These methods do a k-fold stratified cross-validation for you, meaning that each fold will contain the same proportions of the two types of class labels (option "Number of folds"). * '''LibSVM''': Requires the LibSVM toolbox ([[http://www.csie.ntu.edu.tw/~cjlin/libsvm/#download|download]] and add to your path).<<BR>>The LibSVM cross-validation may be faster but will not be stratified. |
* Select process "'''Decoding > SVM decoding'''"<<BR>>'''Note:''' the SVM process requires the LibSVM toolbox, see installation instructions above. {{attachment:13_pipeline_editor_select_decoding.jpg||width="400"}} * Select 'MEG' for sensor types * Set 30 Hz for low-pass cutoff frequency. Equivalently, one could have applied a low-pass filters to the recordings and then run the decoding process. But this is a shortcut to apply a low-pass filter just for decoding without permanently altering the input recordings. * Select 100 for number of permutations. Alternatively use a smaller number for faster results. * Select 5 for number of k-folds * Select 'Pairwise' for decoding. Hint: if more that two classes were input to the Process1 tab, the decoding process will perform decoding separately for each possible pair of classes. It will return them in the same form as Matlab's 'squareform' function (i.e. lower triangular elements in columnwise order) * The decoding process follows a similar procedure as Pantazis et al. (2018). Namely, to reduce computational load and improve signal-to-noise ratio, we first randomly assign all trials (from each class) into k folds, and then subaverage all trials within each fold into a single trial, thus yielding a total of k subaveraged trials per class. Decoding then follows with a leave-one-out cross-validation procedure on the subavaraged trials. * For example, if we have two classes with 100 trials each, selecting 5 number of folds will randomly assign the 100 trials in 5 folds with 20 trials each. The process than will subaverage the 20 trials yielding 5 subaveraged trials for each class. <<BR>> {{attachment:14_svm_decoding_pairwise.jpg||width="380"}} |
| Line 61: | Line 70: |
| * The process will take some time. The results are then saved in a file in the new decoding folder. <<BR>><<BR>> {{attachment:cv_file.gif||height="167",width="268"}} * If you double click on it you will see a decoding curve across time. <<BR>><<BR>> {{attachment:cv_plot.gif||height="172",width="355"}} |
* The process will take some time. The results are then saved in a file in the 'decoding' folder<<BR>> {{attachment:15_svm_decoding_pairwise_results.jpg||width="800"}} |
| Line 64: | Line 72: |
| == Permutation == This is an iterative procedure. The training and test data for the SVM/LDA classifier are selected in each iteration by randomly permuting the samples and grouping them into bins of size n (you can select the trial bin sizes). In each iteration two samples (one from each condition) are left out for test. The rest of the data are used to train the classifier with. |
* The resulting matrix gives you the decoding accuracy over time. Notice that there is no significant accuracy before about 80 ms, after which there is a very high accuracy (> 90%) between 100 to 300 ms. This tells us that the neural representation of the face and object images vary significantly, and if we wanted to analyse this further we could narrow our analysis at this time period. |
| Line 67: | Line 74: |
| * Select process "'''Decoding > Classification with cross-validation'''". Set options as below: <<BR>><<BR>> {{attachment:perm_process.gif||height="366",width="311"}} * '''Trial bin size''': If greater than 1, the training data will be randomly grouped into bins of the size you determine here. The samples within each bin are then averaged (we refer to this as sub-averaging); the classifier is then trained using the averaged samples. For example, if you have 40 faces and 40 scenes, and you set the trial bin size to 5; then for each condition you will have 8 bins each containing 5 samples. Seven bins from each condition will be used for training, and the two left-out bins (one face bin, one scene bin) will be used for testing the classifier performance. * The results are saved in a file in the new decoding folder. <<BR>><<BR>> {{attachment:perm_file.gif||height="142",width="284"}} * |
* For temporal generalization, repeat the above process but select 'Temporal Generalization'. * To evaluate the persistence of neural representations over time, the decoding procedure can be generalized across time by training the SVM classifier at a given time point t, as before, but testing across all other time points (Cichy et al., 2014; King and Dehaene, 2014; Isik et al., 2014). Intuitively, if representations are stable over time, the classifier should successfully discriminate signals not only at the trained time t, but also over extended periods of time.<<BR>> {{attachment:16_svm_decoding_temporalgeneralization.jpg||width="380"}} |
| Line 72: | Line 77: |
| Right-click > Display as time series (or double click). <<BR>><<BR>> {{attachment:perm_plot.gif||height="169",width="346"}} | * The process will take some time. The results are then saved in a file in the 'decoding' folder {{attachment:17_svm_decoding_temporalgeneralization_results.jpg||width="800"}} |
| Line 74: | Line 79: |
| == Acknowledgment == This work was supported by the ''McGovern Institute Neurotechnology Program'' to PIs: Aude Oliva and Dimitrios Pantazis. ''http://mcgovern.mit.edu/technology/neurotechnology-program'' |
* This representation gives us a decoding (approx. symmetric) matrix highlighting time periods that are similar in neural representation in red. For example, early signals up until 200ms have a narrow diagonal, indicating highly transient visual representations. On the other hand, the extended diagonal (red blob) between 200 and 400 ms indicates a stable neural representation at this time. We notice a similar albeit slightly attenuated phenomenon between 400 and 800 ms. In sum, this visualization can help us identify time periods with transient and sustained neural representations. |
| Line 78: | Line 82: |
| 1. Khaligh-Razavi SM, Bainbridge W, Pantazis D, Oliva A (2016)<<BR>>[[http://biorxiv.org/content/early/2016/04/22/049700.abstract|From what we perceive to what we remember: Characterizing representational dynamics of visual memorability]]. ''bioRxiv'', 049700. 1. Cichy RM, Pantazis D, Oliva A (2014)<<BR>>[[http://www.nature.com/neuro/journal/v17/n3/full/nn.3635.html|Resolving human object recognition in space and time]], Nature Neuroscience, 17:455–462. |
1. Cichy RM, Pantazis D, Oliva A (2014), [[http://www.nature.com/neuro/journal/v17/n3/full/nn.3635.html|Resolving human object recognition in space and time]], Nature Neuroscience, 17:455–462. 1. Guggenmos M, Sterzer P, Cichy RM (2018), [[https://doi.org/10.1016/j.neuroimage.2018.02.044|Multivariate pattern analysis for MEG: A comparison of dissimilarity measures]], NeuroImage, 173:434-447. 1. King JR, Dehaene S (2014), [[https://doi.org/10.1016/j.tics.2014.01.002|Characterizing the dynamics of mental representations: the temporal generalization method]], Trends in Cognitive Sciences, 18(4): 203-210 1. Isik L, Meyers EM, Leibo JZ, Poggio T, [[https://doi.org/10.1152/jn.00394.2013|The dynamics of invariant object recognition in the human visual system]], Journal of Neurophysiology, 111(1): 91-102 == Additional documentation == * Forum: Decoding in source space: http://neuroimage.usc.edu/forums/showthread.php?2719 |
Machine learning: Decoding / MVPA
Authors: Dimitrios Pantazis, Seyed-Mahdi Khaligh-Razavi, Francois Tadel,
This tutorial illustrates how to run MEG decoding (a type of multivariate pattern analysis / MVPA) using support vector machines (SVM).
Contents
License
To reference this dataset in your publications, please cite Cichy et al. (2014).
Description of the decoding functions
Two decoding processes are available in Brainstorm:
Decoding -> SVM classifier decoding
Decoding -> max-correlation classifier decoding
These two processes work in a similar way, but they use a different classifier, so only SVM is demonstrated here.
Input: the input is the time recording data from two conditions (e.g. condA and condB) across time. Number of samples per condition do not have to be the same for both condA and condB, but each of them should have enough samples to create k-folds (see parameter below).
Output: the output is a decoding time course, or a temporal generalization matrix (train time x test time).
Classifier: Two methods are offered for the classification of MEG recordings across time: support vector machine (SVM) and max-correlation classifier.
In the context of this tutorial, we have two condition types: faces, and objects. The participant was shown different types of images and we want to decode the face images vs. the object images using 306 MEG channels.
Download and installation
From the Download page of this website, download the file 'sample_decoding.zip', and unzip to get the MEG file: subj04NN_sess01-0_tsss.fif
- SVM decoding requires the LibSVM toolbox. The SVM decoding process tries to automatically install it, but should it fail here's how you install it manually:
Download the libsvm ZIP file here.
- Extract the content of the ZIP file to a permanent folder.
- On Linux or Mac, you need to compile the library. Mac users can use XCode to do so.
- On Windows, you can use windows subfolder of the zip file.
- Add the path to the LibSVM folder in Matlab using addpath.
- Start Brainstorm.
Select the menu File -> Create new protocol. Name it "TutorialDecoding" and select the options:
"Yes, use protocol's default anatomy",
"No, use one channel file per acquisition run".
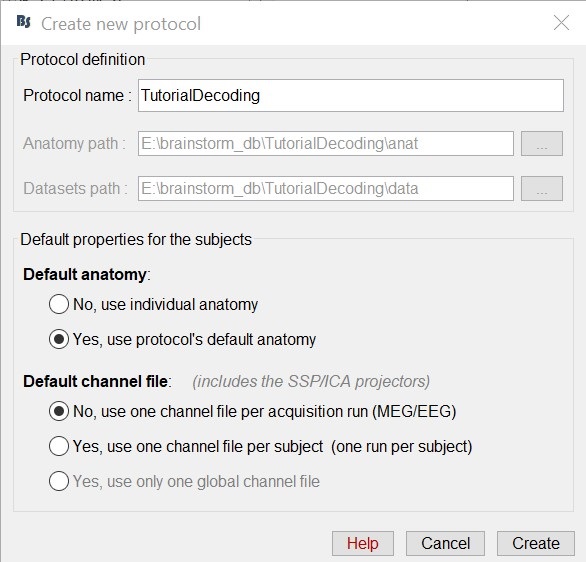
Import the recordings
- Go to the "functional data" view of the database (sorted by subjects).
Right-click on the TutorialDecoding folder -> New subject -> Subject01
Leave the default options you defined for the protocol.Right click on the subject node (Subject01) -> Review raw file.
Select the file format: "MEG/EEG: Neuromag FIFF (*.fif)"
Select the file: subj04NN-sess01-0_tsss.fif
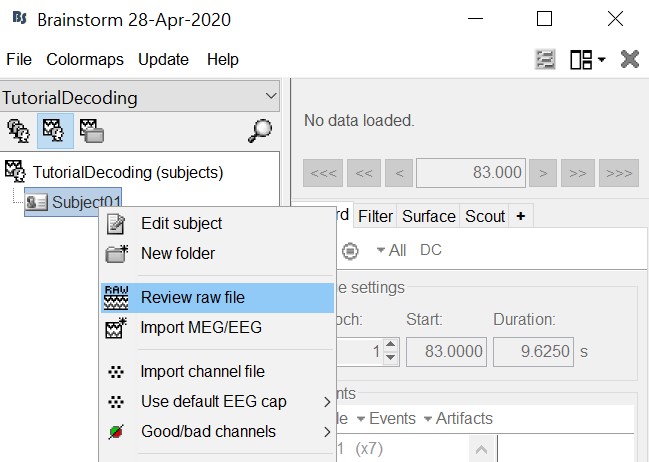
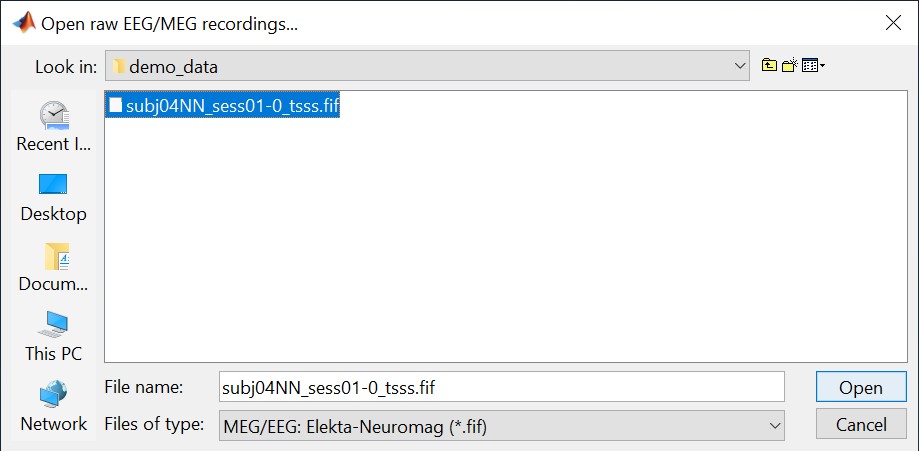
Select "Event channels" to read the triggers from the stimulus channel.
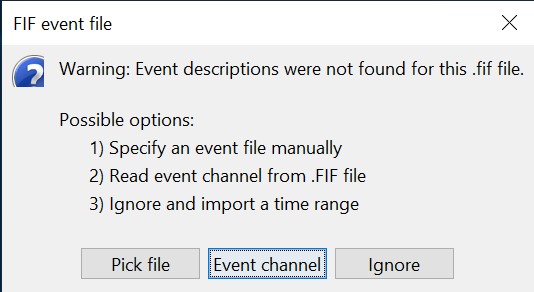
- We will not pay attention to MEG/MRI registration because we are not going to compute any source models. The decoding is done on the sensor data.
Double click on the 'Link to raw file' to visualize the raw recordings. Event codes 13-24 indicate responses to face images, and we will combine them to a single group called 'faces'. To do so, select events 13-24 using SHIFT + click and from the menu select "Events -> Duplicate groups". Then select "Events -> Merge groups". The event codes are duplicated first so we do not lose the original 13-24 event codes.
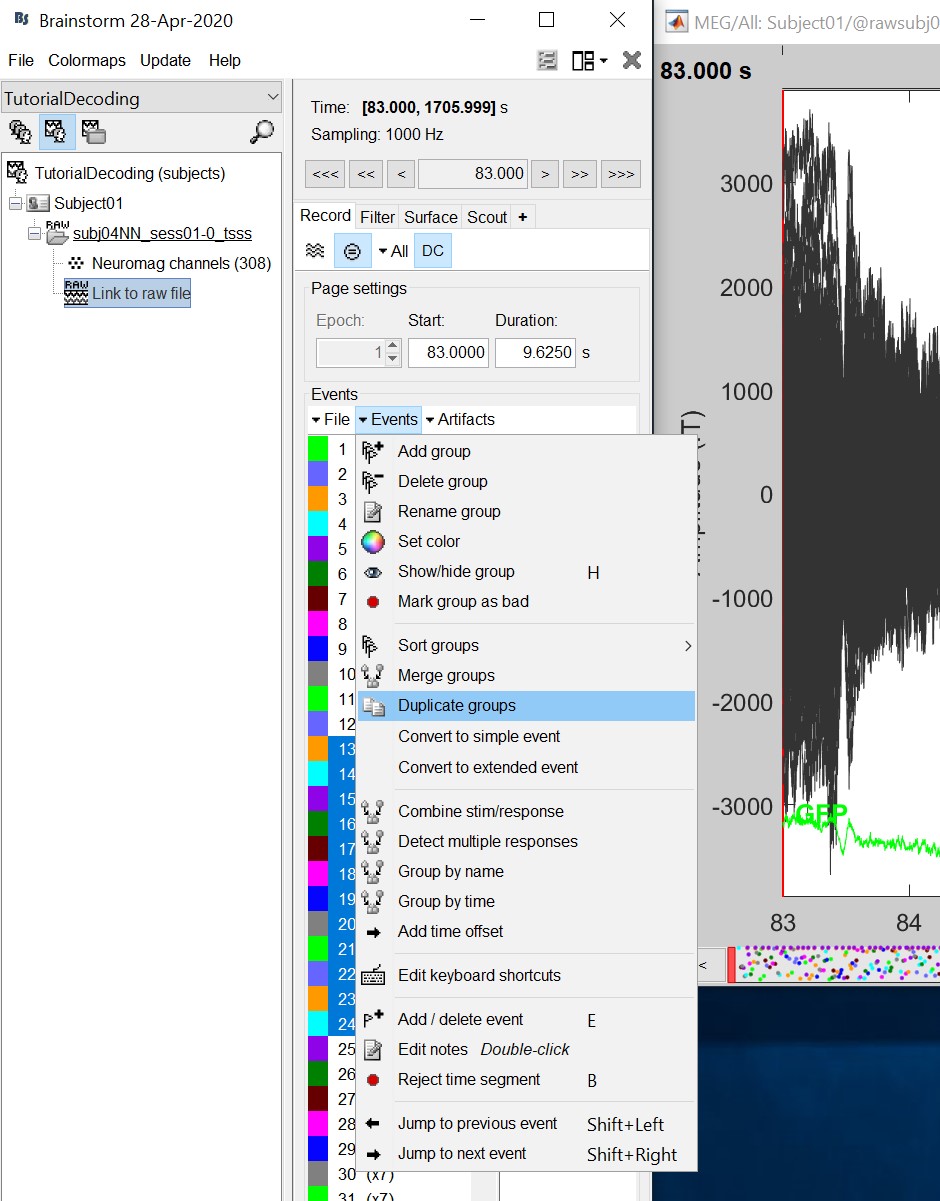
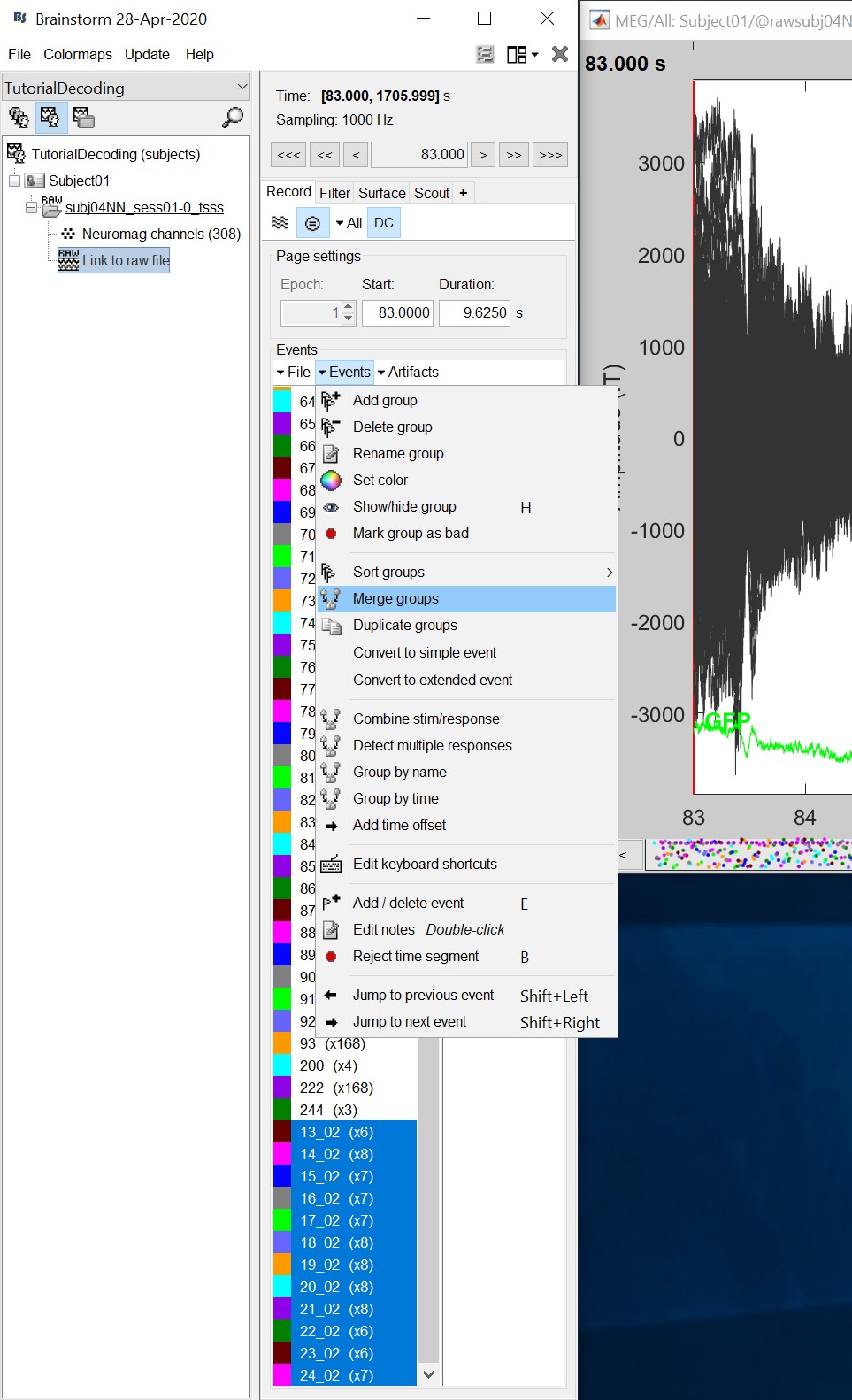
Event codes 49-60 indicate responses to object images, and we will combine them to a single group called 'objects'. To do so, select events 49-60 (SHIFT + click) and from the menu select Events -> Duplicate groups. Then select Events -> Merge groups. No screenshots are shown since this is similar to above.
We will now import the 'faces' and 'objects' responses to the database. Select "File -> Import in database".
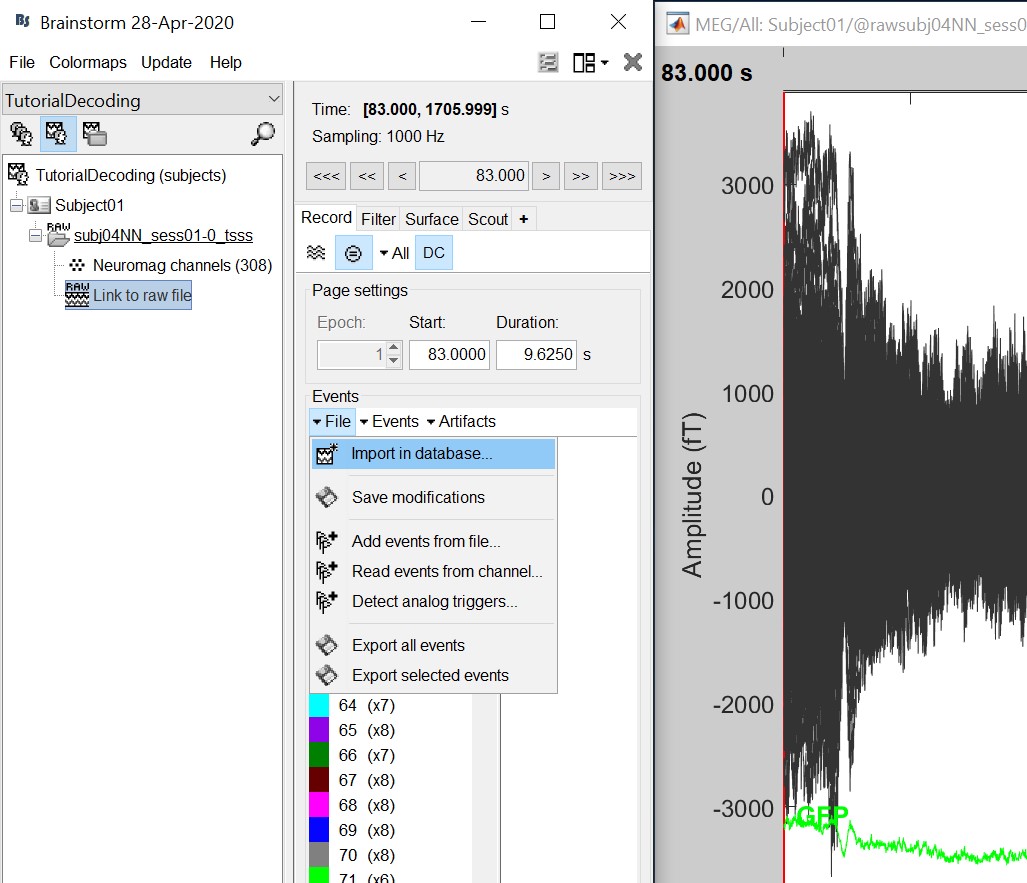
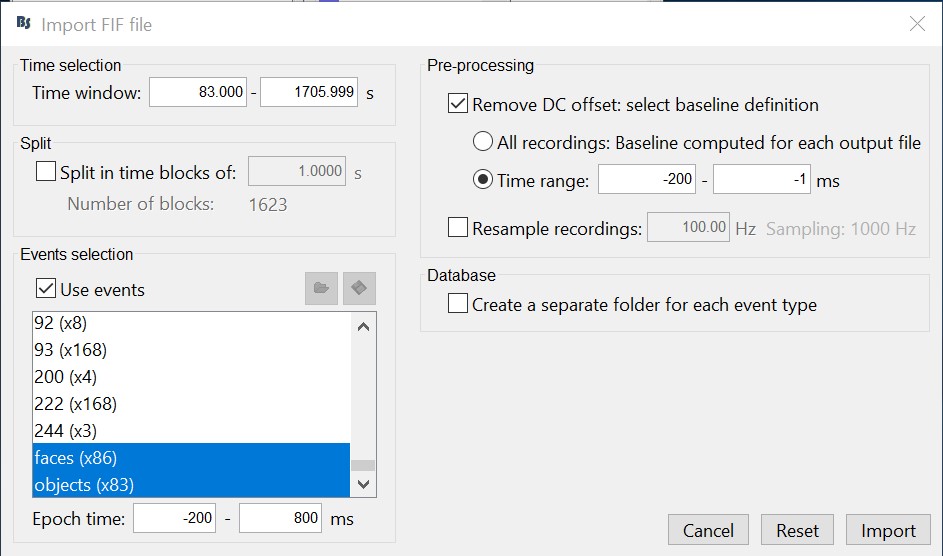
- Select only two events: 'faces' and 'objects'
- Epoch time: [-200, 800] ms
- Remove DC offset: Time range: [-200, 0] ms
Do not create separate folders for each event type
Select files
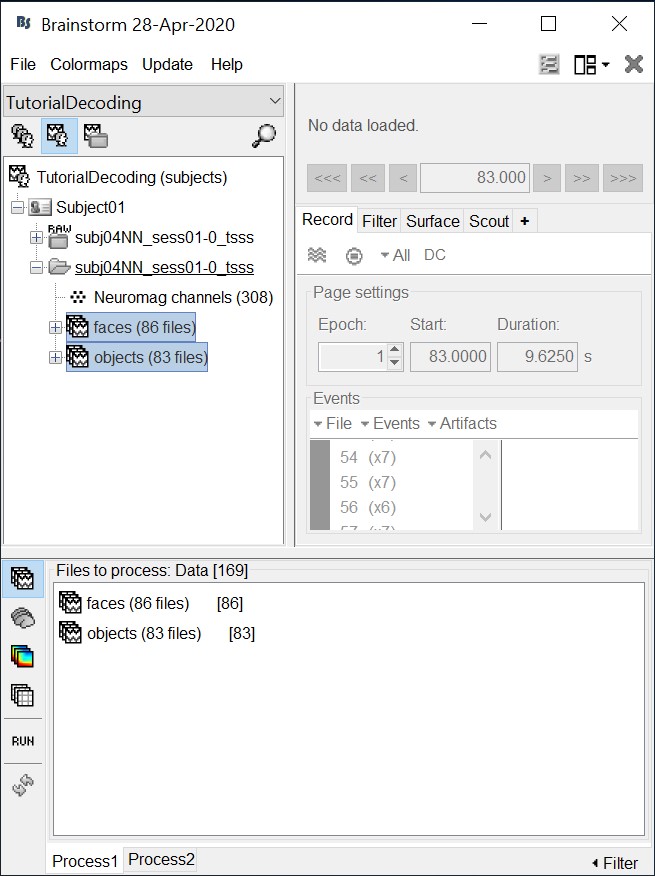
- Drag and drop all the face and object trials to the Process1 tab at the bottom of the Brainstorm window.
Intuitively, you might have expected to use the Process2 tab to decode faces vs. objects. But the decoding process is designed to also handle pairwise decoding of multiple classes (not just two classes) for computational efficiency, so more that two categories can be entered in the Process1 tab.
Decoding with cross-validation
Cross-validation is a model validation technique for assessing how the results of our decoding analysis will generalize to an independent data set.
Select process "Decoding > SVM decoding"
Note: the SVM process requires the LibSVM toolbox, see installation instructions above.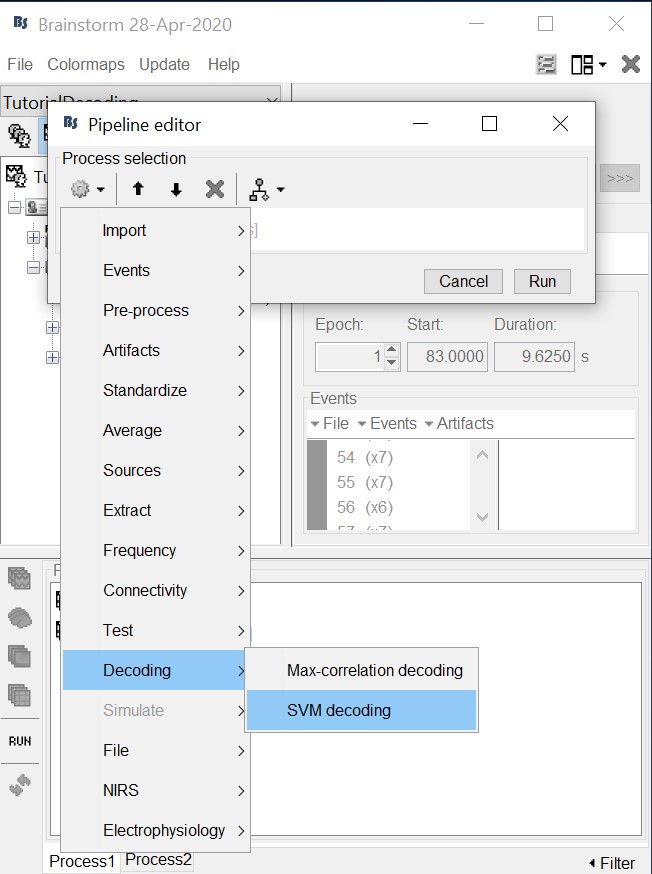
- Select 'MEG' for sensor types
- Set 30 Hz for low-pass cutoff frequency. Equivalently, one could have applied a low-pass filters to the recordings and then run the decoding process. But this is a shortcut to apply a low-pass filter just for decoding without permanently altering the input recordings.
- Select 100 for number of permutations. Alternatively use a smaller number for faster results.
- Select 5 for number of k-folds
- Select 'Pairwise' for decoding. Hint: if more that two classes were input to the Process1 tab, the decoding process will perform decoding separately for each possible pair of classes. It will return them in the same form as Matlab's 'squareform' function (i.e. lower triangular elements in columnwise order)
- The decoding process follows a similar procedure as Pantazis et al. (2018). Namely, to reduce computational load and improve signal-to-noise ratio, we first randomly assign all trials (from each class) into k folds, and then subaverage all trials within each fold into a single trial, thus yielding a total of k subaveraged trials per class. Decoding then follows with a leave-one-out cross-validation procedure on the subavaraged trials.
For example, if we have two classes with 100 trials each, selecting 5 number of folds will randomly assign the 100 trials in 5 folds with 20 trials each. The process than will subaverage the 20 trials yielding 5 subaveraged trials for each class.
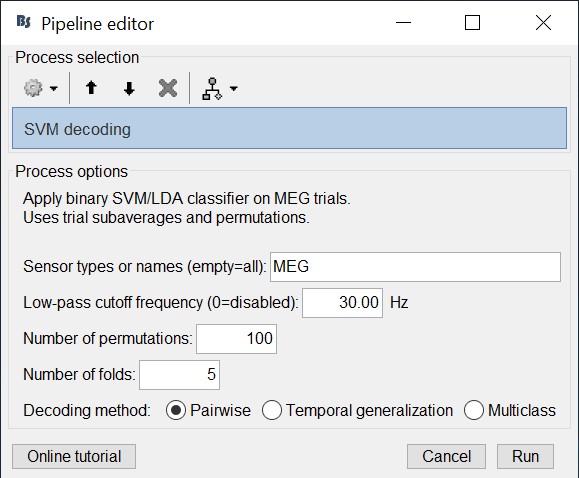
The process will take some time. The results are then saved in a file in the 'decoding' folder
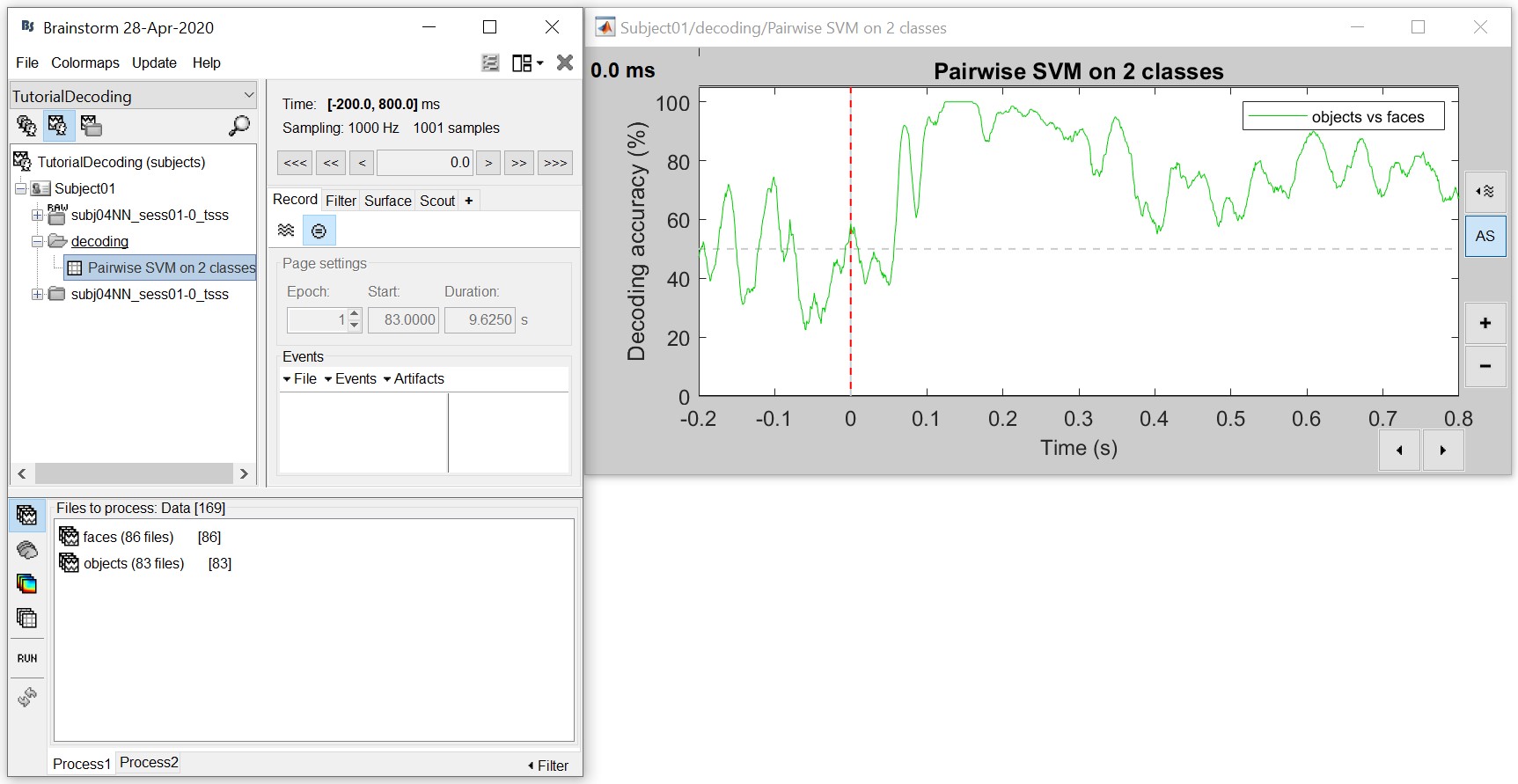
The resulting matrix gives you the decoding accuracy over time. Notice that there is no significant accuracy before about 80 ms, after which there is a very high accuracy (> 90%) between 100 to 300 ms. This tells us that the neural representation of the face and object images vary significantly, and if we wanted to analyse this further we could narrow our analysis at this time period.
- For temporal generalization, repeat the above process but select 'Temporal Generalization'.
To evaluate the persistence of neural representations over time, the decoding procedure can be generalized across time by training the SVM classifier at a given time point t, as before, but testing across all other time points (Cichy et al., 2014; King and Dehaene, 2014; Isik et al., 2014). Intuitively, if representations are stable over time, the classifier should successfully discriminate signals not only at the trained time t, but also over extended periods of time.
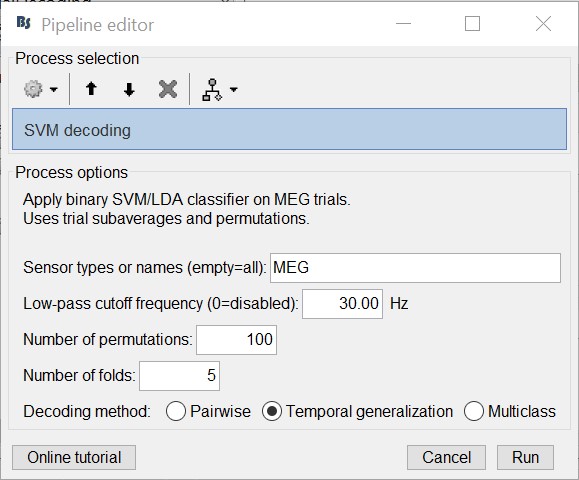
The process will take some time. The results are then saved in a file in the 'decoding' folder
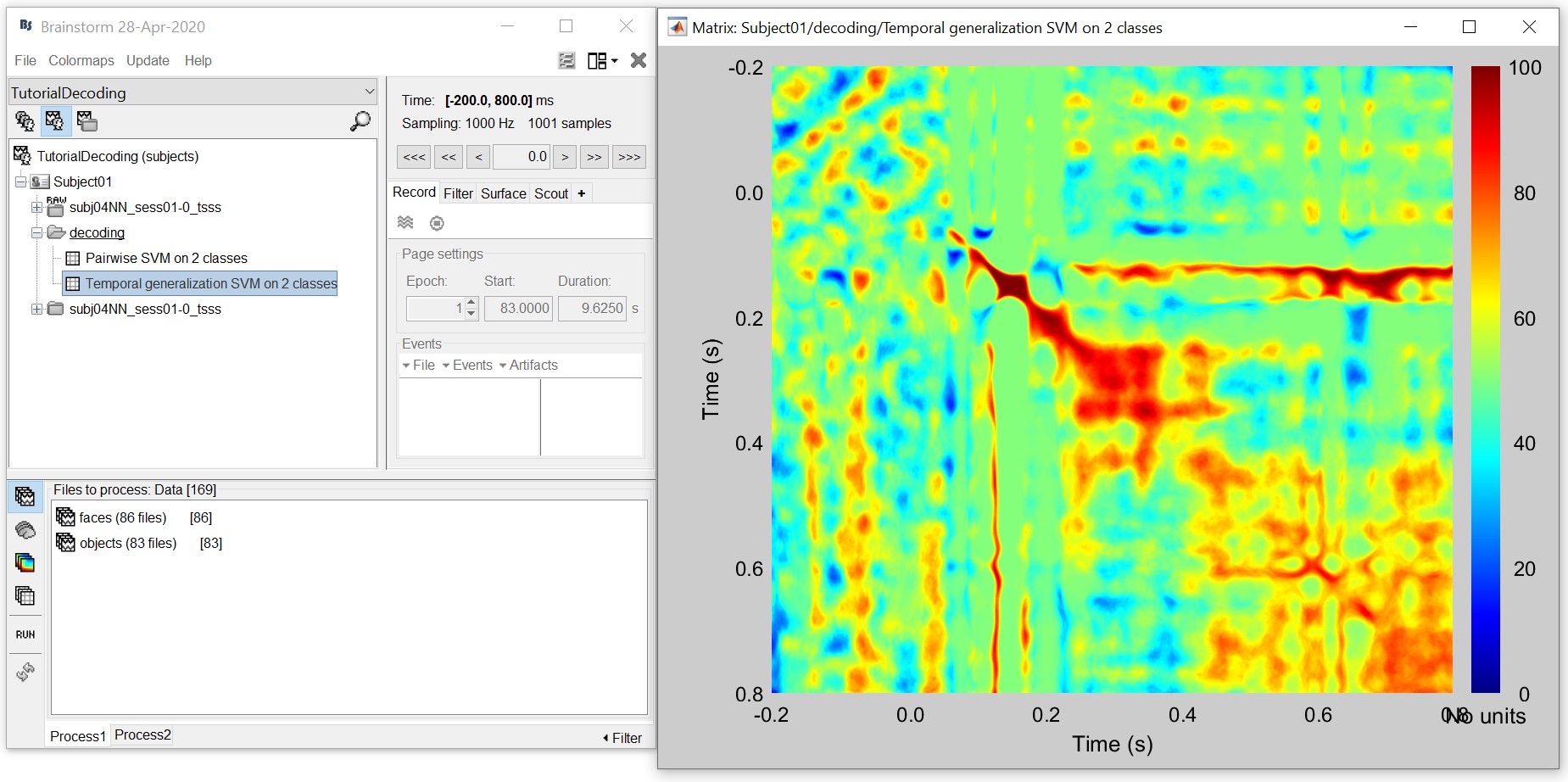
- This representation gives us a decoding (approx. symmetric) matrix highlighting time periods that are similar in neural representation in red. For example, early signals up until 200ms have a narrow diagonal, indicating highly transient visual representations. On the other hand, the extended diagonal (red blob) between 200 and 400 ms indicates a stable neural representation at this time. We notice a similar albeit slightly attenuated phenomenon between 400 and 800 ms. In sum, this visualization can help us identify time periods with transient and sustained neural representations.
References
Cichy RM, Pantazis D, Oliva A (2014), Resolving human object recognition in space and time, Nature Neuroscience, 17:455–462.
Guggenmos M, Sterzer P, Cichy RM (2018), Multivariate pattern analysis for MEG: A comparison of dissimilarity measures, NeuroImage, 173:434-447.
King JR, Dehaene S (2014), Characterizing the dynamics of mental representations: the temporal generalization method, Trends in Cognitive Sciences, 18(4): 203-210
Isik L, Meyers EM, Leibo JZ, Poggio T, The dynamics of invariant object recognition in the human visual system, Journal of Neurophysiology, 111(1): 91-102
Additional documentation
Forum: Decoding in source space: http://neuroimage.usc.edu/forums/showthread.php?2719
