|
Size: 3039
Comment:
|
Size: 4280
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| The case from the tutorial is also published in this article: Dümpelmann M, Ball T, Schulze-Bonhage A<<BR>>[[http://www.ncbi.nlm.nih.gov/pubmed/21618659|LORETA allows reliable distributed source reconstruction based on subdural strip and grid recordings]], Hum Brain Mapp. 2012 | The case from this tutorial is also published in this article: Dümpelmann M, Ball T, Schulze-Bonhage A<<BR>>[[http://www.ncbi.nlm.nih.gov/pubmed/21618659|LORETA allows reliable distributed source reconstruction based on subdural strip and grid recordings]], Hum Brain Mapp. 2012 |
| Line 7: | Line 7: |
| This tutorial dataset (EEG and MRI data) remains proprietary of the Epilepsy Centre, University Hospital Freiburg, Germany. Its use and transfer outside the brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the Epilepsy Centre in Freiburg. <<BR>>For questions please contact A. Schulze-Bonhage, MD, PhD (andreas.schulze-bonhage@uniklinik-freiburg.de). | This tutorial dataset (EEG and MRI data) remains proprietary of the Epilepsy Centre, University Hospital Freiburg, Germany. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the Epilepsy Centre in Freiburg. For questions please contact A. Schulze-Bonhage, MD, PhD: andreas.schulze-bonhage@uniklinik-freiburg.de |
| Line 12: | Line 12: |
| Type of epilepsy, supposed location, etc. | Type of epilepsy, supposed location, clinical conclusions, etc. |
| Line 15: | Line 15: |
| * Requirements: You have already followed all the basic tutorials and you have a working copy of Brainstorm installed on your computer. | * Requirements: You have already followed all the introduction tutorials and you have a working copy of Brainstorm installed on your computer. |
| Line 17: | Line 17: |
| * Unzip it in a folder that is not in any of the Brainstorm folders (program folder or database folder). This is really important that you always keep your original data files in a separate folder: the program folder can be deleted when updating the software, and the contents of the database folder is supposed to be manipulated only by the program itself. | * Unzip it in a folder that is __not__ in any of the Brainstorm folders (program folder or database folder) |
| Line 19: | Line 19: |
| * Select the menu File > Create new protocol. Name it "'''TutorialEpilepsy'''" and select the options: | * Select the menu File > Create new protocol. Name it "'''!TutorialEpilepsy'''" and select the options: |
| Line 24: | Line 24: |
| For now the MRI is processed with BrainVISA, but we would like to have it fully processed with FreeSurfer. | * Create one subject. * Right-click on the subject node > Import anatomy folder > sample_epilepsy/anatomy (format=!FreeSurfer) * If no anatomy available: [[Tutorials/TutWarping|Warping tutorial]] |
| Line 26: | Line 28: |
| == Access the raw file == ... |
== Access the recordings == * Switch to the "functional data" view. * Right-click on the subject folder > Review raw file > sample_epilepsy/data/tutorial_EEG.bin (format = "EEG: Deltamet Neurofile-Coherence (*.bin)") * Right-click on the subject folder > Import channel file > sample_epilepsy/data/XensorTest.elc (format = "EEG: ANT Xensor (*.elc)")<<BR>=> Contains the default electrodes positions from the ASA software (ANT). * Change the sensor type of the following electrodes to '''MISC''': SP1, SP2, RS, PHO, DELR, DELL, QR, QL * Run the automatic alignment manually. |
| Line 30: | Line 36: |
| How was it done initially? | * How was it done initially? * How would we do it in Brainstorm? * Import 177 spikes: * '''[-1, +1] seconds''' around the spike events (???) * Remove DC offset based on the whole trial (-1s to +1s) (???) |
| Line 32: | Line 42: |
| How would we do it in Brainstorm? | == Source analysis == * Compute head model: OpenMEEG or 3-shell sphere (???) * Noise covariance matrix: '''[-1000, -800] ms''' from all the imported trials (???) * Source model: (???) |
| Line 34: | Line 47: |
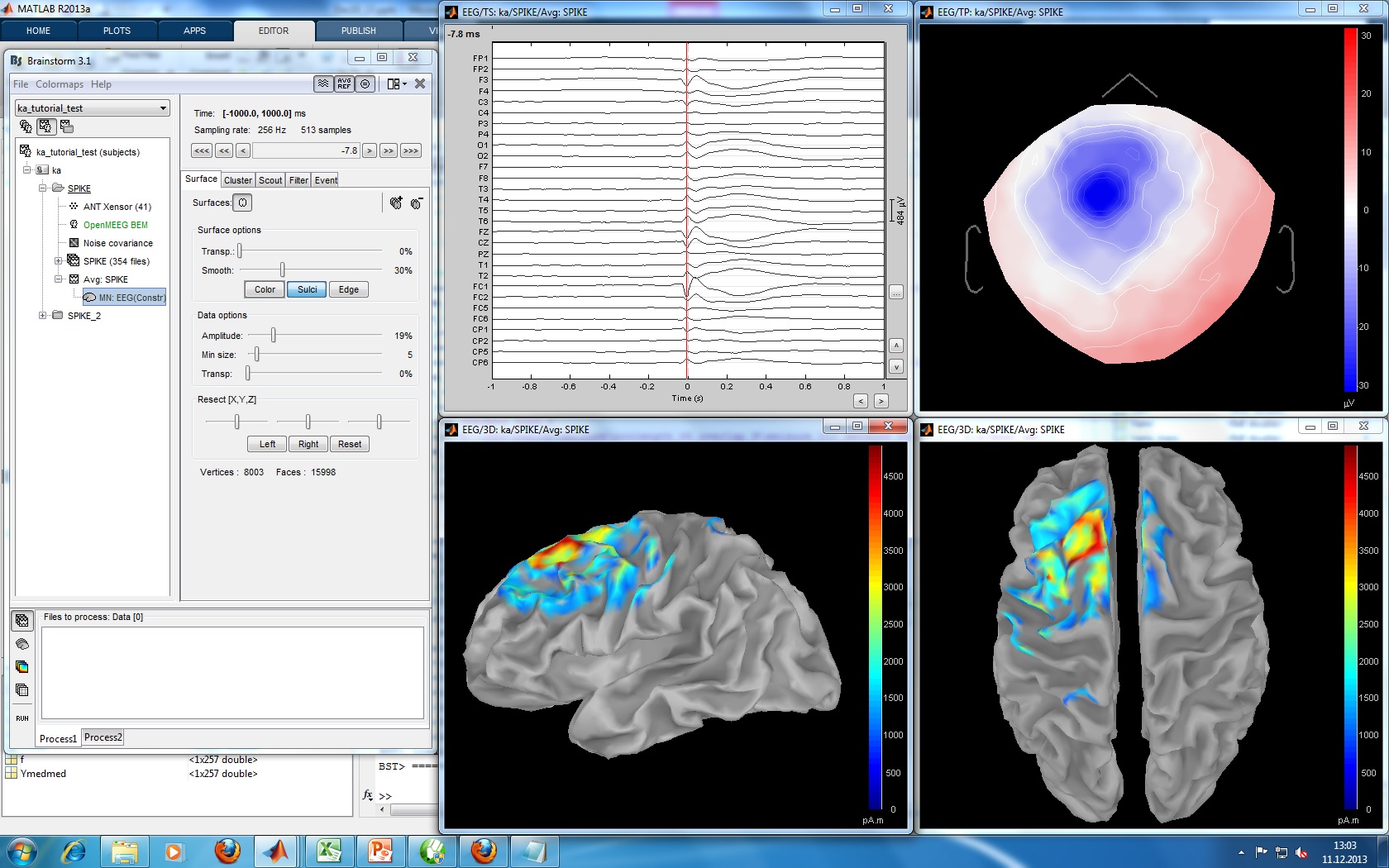
| Import 177 spikes: [-1, +1] seconds arounf the spike events | == Moving dipoles == Illustrate John/Beth's tools for calculating and displaying dipoles. == Tools to be developed for this tutorial == * Set the time resolution of the display figures in seconds per centimeter (s/cm): the length of the page changes automatically with the size of the window. * Flip +/- the Y axis (already done: check the brainstorm toolbar) * Refine a bit the display of the dipoles * Flexible montage interface (is this useful here ???) {{attachment:SL_MNE_tutorial_patient.jpg||height="403",width="645"}} |
EEG and epilepsy
This tutorial introduces some concepts that are specific to the management of EEG recordings in the Brainstorm environment. It also describes a standard pipeline for analyzing epilepsy recordings. It is based on clinical case from the Epilepsy Centre, at the University Hospital Freiburg, Germany. The EEG data was recorded at 1024Hz, using a Neurofile NT digital video-EEG system with 128 channels. The anonymized dataset can be downloaded directly from the Brainstorm download page.
The case from this tutorial is also published in this article: Dümpelmann M, Ball T, Schulze-Bonhage A
LORETA allows reliable distributed source reconstruction based on subdural strip and grid recordings, Hum Brain Mapp. 2012
License
This tutorial dataset (EEG and MRI data) remains proprietary of the Epilepsy Centre, University Hospital Freiburg, Germany. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the Epilepsy Centre in Freiburg. For questions please contact A. Schulze-Bonhage, MD, PhD: andreas.schulze-bonhage@uniklinik-freiburg.de
Presentation of the clinical case
The EEG data was recorded at 1024Hz, using a Neurofile NT digital video-EEG system with 128 channels and a 16-bit A/D converter. The signal was filtered in the recording system with a high-pass filter with a time constant of 1 second and a low-pass filter with a cutoff frequency of 344 Hz. The spikes were visually identified and averaged with the ASA package. The spike average showed prominent peaks in the grid contacts G_A2-4, G_B2-5, G_C1-3.
Type of epilepsy, supposed location, clinical conclusions, etc.
Download and installation
- Requirements: You have already followed all the introduction tutorials and you have a working copy of Brainstorm installed on your computer.
Go to the Download page of this website, and dowload the file: sample_epilepsy.zip
Unzip it in a folder that is not in any of the Brainstorm folders (program folder or database folder)
- Start Brainstorm (Matlab scripts or stand-alone version)
Select the menu File > Create new protocol. Name it "TutorialEpilepsy" and select the options:
"No, use individual anatomy",
"Yes, use one channel file per subject".
Import the anatomy
- Create one subject.
Right-click on the subject node > Import anatomy folder > sample_epilepsy/anatomy (format=FreeSurfer)
If no anatomy available: Warping tutorial
Access the recordings
- Switch to the "functional data" view.
Right-click on the subject folder > Review raw file > sample_epilepsy/data/tutorial_EEG.bin (format = "EEG: Deltamet Neurofile-Coherence (*.bin)")
Right-click on the subject folder > Import channel file > sample_epilepsy/data/XensorTest.elc (format = "EEG: ANT Xensor (*.elc)")<<BR>=> Contains the default electrodes positions from the ASA software (ANT).
Change the sensor type of the following electrodes to MISC: SP1, SP2, RS, PHO, DELR, DELL, QR, QL
- Run the automatic alignment manually.
Mark and review spikes
- How was it done initially?
- How would we do it in Brainstorm?
- Import 177 spikes:
[-1, +1] seconds around the spike events (???)
- Remove DC offset based on the whole trial (-1s to +1s) (???)
Source analysis
- Compute head model: OpenMEEG or 3-shell sphere (???)
Noise covariance matrix: [-1000, -800] ms from all the imported trials (???)
- Source model: (???)
Moving dipoles
Illustrate John/Beth's tools for calculating and displaying dipoles.
Tools to be developed for this tutorial
- Set the time resolution of the display figures in seconds per centimeter (s/cm): the length of the page changes automatically with the size of the window.
- Flip +/- the Y axis (already done: check the brainstorm toolbar)
- Refine a bit the display of the dipoles
- Flexible montage interface (is this useful here ???)