|
Size: 15397
Comment:
|
Size: 19653
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 38: | Line 38: |
| * Go to the [[http://neuroimage.usc.edu/brainstorm3_register/download.php|Download]] page of this website, and download the file: '''sample_epilepsy.zip ''' | * Go to the [[http://neuroimage.usc.edu/brainstorm3_register/download.php|Download]] page of this website, and download the file: '''sample_epilepsy.zip''' |
| Line 107: | Line 107: |
| * Click on "OK" to save the modifications. {{attachment:channel_align.gif|channel_type.gif}} |
* Click on "OK" to save the modifications.<<BR>><<BR>> {{attachment:channel_align.gif|channel_type.gif}} |
| Line 113: | Line 112: |
| * Right-click on the "Link to raw file" > EEG > Display time series. * In the Record tab, select the first button in the toolbar (Display mode for time series) to view the signals in columns. * In the same toolbar, select the "'''Longitudinal 1'''" in the drop-down list. This menu allows you to set a predefined montages to linearly re-arranging the waveforms. |
* Open a time series figure with the "'''Longitudinal 1'''"montage * Right-click on the "Link to raw file" > EEG > Display time series. * In the Record tab, select the first button in the toolbar (Display mode for time series) to view the signals in columns. * In the same toolbar, select the "Longitudinal 1" in the drop-down list. This menu allows you to set a predefined montages to linearly re-arranging the waveforms. |
| Line 123: | Line 123: |
| * Open another view on the same data: | * Open another view on the same data with the "'''Transversal 1'''" montage: |
| Line 126: | Line 126: |
| * Set the montage for this new figure to "'''Transversal 1'''" (note the keyboard shortcuts in the menus) * Right-click on "Link to raw file" again > EEG > 2D Sensor cap * Re-arrange the figures in a convenient way, for example as illustrated below ''=> Display the ECG/EMG/EOG at the same time? '' ''=> Screen captures'' |
* Set the montage for this new figure to "Transversal 1" (note the keyboard shortcuts in the menus) * Open a 2D map of the sensor values as well: * Right-click on "Link to raw file" again > EEG > 2D Sensor cap * In the Record tab, set the Montage to this view to "'''Average reference'''" * Re-arrange the figures in a convenient way, for example as illustrated below, and disable the automatic positioning of the figures you the figure arrangement doesn't get lost when you open a new figure (Window layout menu at the top-right of the Brainstorm figure > None) => ECG/EMG/EOG ??? => SCREEN CAPTURE ??? |
| Line 135: | Line 137: |
| Go to the Filter tab to enable some display frequency filters. General recommendations are to bandpass filter as: * High-pass filter: '''0.5 Hz ''' * Low-pass filter: '''70 Hz''' |
Go to the Filter tab to enable some display frequency filters. General recommendations are: * High-pass filter: '''0.5 Hz''' * Low-pass filter: '''70 Hz'''<<BR>><<BR>> {{attachment:filters.gif}} |
| Line 141: | Line 143: |
| The resolution of the time and amplitude axis has a lot of importance for the visual detection of epileptic spikes. * Time axis: 30 mm/sec (~100 pixels/sec?) * Amplitude: 10 μV/mm (~3 μV/pixels?) ''[ADDITIONAL EXPLANATIONS]'' |
The resolutions of the time and amplitude axes has a lot of importance for the visual detection of epileptic spikes. The shapes we are looking for are altered by the horizontal and vertical scaling. The distance unit on a screen is the pixel, we can set how much time is represented by one pixel horizontally and how much amplitude is represented by one pixel vertically. In the Brainstorm interface, this resolution is usually set implicitly: you can set the size of the window, the duration or recordings reviewed at once (text box "duration" in tab Record) and the maximum amplitude to show in the figure (buttons [...] and [AS] on the right of the time series figure). From there, you can also zoom in time ([<], [>], mouse wheel) or amplitude (['''^'''], [v], Shift+mouse wheel). Those parameters are convenient to explore the recordings interactively but don't allow us to have reproducible displays with constant time and amplitude resolutions. To set the figure resolutions: right-click on the figure > Figure > Set axes resolution. Note that this interface does not store the input values, it just modifies the other parameters (figure size, time window, max amplitude) to fit the resolution objectives. Then if you modify those parameters (resize the figure, leave the button [AS] selected and scroll in time...) the resolution is lost, you have to set it again manually. Recommendations for this dataset are: * Time axis: 90 pixels/sec (~30 mm/sec) * Amplitude: 4 μV/pixels (~12 μV/mm) => CHECK THE VALUES ??? => SCREEN CAPTURE ??? |
| Line 149: | Line 159: |
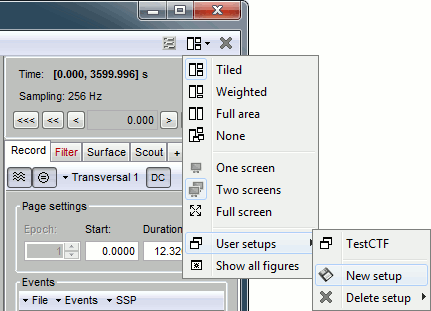
| This preparation of the reviewing environment requires a large number of operations, and would become quickly annoying if you have to repeat it everytime you open a file. This is a good time to use the menu "User setups" to save this window configuration, to reload it in one click later. In the menu "Window layout", at the top-right of the Brainstorm window, select User setup > New setup > "Spike search" |
This preparation of the reviewing environment requires a large number of operations, and would become quickly annoying if you have to repeat it everytime you open a file. This is a good time to use the menu "User setups" to save this window configuration, to reload it in one click later. In the menu "Window layout", at the top-right of the Brainstorm window, select User setup > New setup. Enter a name of your choice for this particular window arrangement. This operation will also automatically disable the automatic window arrangement (Window layout > None). To reload it later, open one figure on the dataset you want to review and then select your new entry in the User setup menu. . {{attachment:usersetup.gif}} |
| Line 154: | Line 166: |
| ''HOW TO PROCEED?'' ''SCREEN CAPTURE?'' ==== Import recordings ==== We have a set of events already marked for this data set. * High-pass filter the recording at 0.5Hz * Import 177 spikes: * '''[-100, +300]ms''' around the spike events * The DC offset will have been removed with the high-pass filter, therefore there is no need to remove it at this point. |
In the dataset, the spikes have already been identified by experts at the University Hospital of Freiburg. You can see that 177 SPIKE events are available in the Record tab. Click on a few of them a try to identify the shape of the spike. => SCREEN CAPTURE ??? The procedure if you are marking the events by yourself is the following: * Open a continuous file viewer, the load your reviewing environment (menu User setups) * Start by creating a group of events (in the Record tab: Events > Add group), and select it in the list of events. * Make sure that the time and amplitude resolutions are what you are used to (right-click on the figure > Figure > Set axes resolution) * Scroll through the recordings using the [<<<] and [>>>] buttons or the associated shortcuts such as F3 or F4 (complete list and descriptions available when you leave your mouse over those buttons). * You can adjust the gain of the electrodes to observe better an event with the buttons ['''^'''] and [v] or the shortcut [Shift+mouse wheel]. * When you identify a spike, click in a white area of the figure in order to place the time cursor at the peak of the spike. If you click on the signal itself, it selects the corresponding channel, but you can use the shortcut Shift+Click to prevent this behavior and force the time cursor to be moved instead. * Press Ctrl+E to add a marker where the time cursor is. * If you are marking multiple types of events, it is convenient to set up some additional keyboard shortcuts. Using the menu '''Events > Edit keyboard shortcuts''', you can associate custom events to the key 1 to 9 of the keyboard. Define the name of the event type to create for each key, and then simply press the corresponding key to add/delete a marker at the current time position. * More information on the data viewer, see tutorial: [[TutRawViewer|Review continuous recordings]]. => CONFIRM THE PROCEDURE ??? == Pre-process recordings == * High-pass filter the recording at 0.5Hz => APPLY 0.5 Hz FILTER TO THE CONTINUOUS RECORDINGS BEFORE IMPORTING ??? => (NOT POSSIBLE WITH THIS FILE FORMAT) == Import recordings == * Right-click on the "Link to raw file" > Import in database * Select the option "Use events" and select the category '''SPIKE''', with 177 events. * Epoch time: '''[-100, +300]ms''' around the spike events * Uncheck the option "Remove DC offset" because the DC offset has already been removed with the high-pass filter<<BR>>{{attachment:import.gif}} |
| Line 167: | Line 198: |
| * The pial surface will be used for this modeling. * Compute head model: OpenMEEG * Noise covariance matrix: select 10 sec time window from the raw file which would represent the subject's resting state with no apparent epileptic activity - use this for all runs<<BR>>''[=> WHAT WOULD IT BE?]'' * Source model: wMNE, constrained orientations * Compute z-score * Discuss advantage of distributed source models in the evaluation of epilepsy data compared with ECD (Kobayashi K, Epilepsia 2005) |
* The pial surface will be used for this modeling. * Compute head model: OpenMEEG * Noise covariance matrix: select 10 sec time window from the raw file which would represent the subject's resting state with no apparent epileptic activity - use this for all runs * Source model: wMNE, constrained orientations * Compute z-score * Discuss advantage of distributed source models in the evaluation of epilepsy data compared with ECD (Kobayashi K, Epilepsia 2005) => WHAT SEGMENT OF 10s CAN WE USE FOR THE NOISE COVARIANCE ??? |
| Line 175: | Line 208: |
| Illustrate John/Beth's tools for calculating and displaying dipoles. {{attachment:SL_MNE_tutorial_patient.jpg||height="403",width="645"}} |
Illustrate John/Beth's tools for calculating and displaying dipoles. => DOES THIS WORK ??? . {{attachment:SL_MNE_tutorial_patient.jpg||height="403",width="645"}} |
EEG and Epilepsy
This tutorial introduces some concepts that are specific to the management of EEG recordings in the Brainstorm environment. It also describes a standard pipeline for analyzing epilepsy recordings. It is based on a clinical case from the Epilepsy Center at the University Hospital of Freiburg, Germany. The anonymized dataset can be downloaded directly from the Brainstorm download page.
License
This tutorial dataset (EEG and MRI data) remains proprietary of the Epilepsy Centre, University Hospital Freiburg, Germany. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the Epilepsy Center in Freiburg. For questions please contact A. Schulze-Bonhage, MD, PhD: andreas.schulze-bonhage@uniklinik-freiburg.de
Presentation of the clinical case
This tutorial dataset was acquired in a patient who suffered from focal epilepsy with focal sensory, dyscognitive and secondarily generalized seizures since the age of eight years. He does not have any typical risk factors for epilepsy. The high resolution 3T epilepsy MRI including postprocessing was found to be normal. FDG-PET of the brain did not show any pathological changes in the glucose metabolism. Non-invasive telemetry revealed left fronto-central sharp waves, polyspikes and bursts of beta band activity (max. amplitude FC1, Cz) especially during sleep. The tutorial dataset was acquired during one night of the non-invasive telemetry recording at the Epilepsy Center Freiburg, Germany.
Afterwards the patient underwent invasive EEG to identify the epileptogenic area and to map functionally important cortex. Details about invasive EEG and source localization from invasive EEG in this patient are reported in Dümpelmann, et al. (2011). Subsequently a left frontal tailored resection was performed. The histological analysis revealed a focal cortical dysplasia type IIB according to the classification of Palmini, et al. (2004). The postsurgical outcome is Engel 1A with a follow-up of 5 years.
The EEG data distributed here was recorded at 1024Hz, using a Neurofile NT digital video-EEG system with 128 channels and a 16-bit A/D converter. The signal was filtered in the recording system with a high-pass filter with a time constant of 1 second and a low-pass filter with a cutoff frequency of 344 Hz. The spikes were marked in the Deltamed Coherence Viewer.
Overview of the data processing
The proper identification of epileptiform discharges ("spikes") is a complicated topic beyond the scope of this tutorial. Neurology students train intensively on how to identify true interictal and ictal discharges, particularly as distinct from so-called "normal variants" that can be abundant. Sleep and drowsy states of the brain can generate "vertex waves," "K-complexes," "positive occipital sharp transients of sleep" (POSTS), and "wickets," to name a few variants, none of which are epileptic. A good overview of terminology and application can be found, for example, on the Medscape website, Epileptiform Discharges.
With this caveat, we nonetheless give an overview of the processing approach. Although automation has been proposed for decades, "spike hunting" is often done by manual inspection of the recorded EEG waveforms (whether scalp, subdural, or depths). The classic scalp sensor arrangement is the International 10-20 System (Wikipedia Site, Bioelectromagnetism Book, Chapter 13.3) arranged at 10 percent and 20 percent distances about the circumference of the head. Each of these 21 electrodes is acquired with respect to some reference (as all potentials must be). In reviewing the recordings, however, several "montages" are traditionally recommended that digitally "re-reference" the original recordings into new linear combinations of electrodes.
A classic montage is the "double banana" (Google Search) which emphasizes local changes in the scalp EEG by forming sequential bipolar pairs, such as "Fp1-F3", "F3-C3." Because F3 is found twice in this montage and of opposite sign, then an epileptic spike centered under F3 will appear as a reversed polarity in these two channels, a visual cue the trained epileptologist seeks when rapidly scanning through hours of recordings. This montage is more formally known as "LB-18.3" or "Longitudinal Bipolar 3" in the nomenclature of the American Clinical Neurophysiology Society Guidelines (see References Below).
ACNS guidelines suggest using both "longitudinal" and "transverse" bipolar montages to survey your data. For temporal epilepsy cases, you may also add a "temporal ring" montage. Brainstorm automatically provides several variations of these montages, and allows the users full flexibility in creating their own. Brainstorm allows multiple different montages to be run simultaneously in multiple windows, thus the user can step along through the data, look for abnormal brain activity, then use Brainstorm's event markers to tag suspect intervals.
With the suspected events marked and saved, the user can return later to perform source analyses on these intervals. With this brief overview, we detail below an exercise with the sample epilepsy data.
References
Dümpelmann M, Ball T, Schulze-Bonhage A (2011)
sLORETA allows reliable distributed source reconstruction based on subdural strip and grid recordings. Human Brain Mapping.
Palmini A, Najm I, Avanzini G, Babb T, Guerrini R, Foldvary-Schaefer N, Jackson G, Luders HO, Prayson R, Spreafico R, Vinters HV (2004) Terminology and classification of the cortical dysplasias.
Neurology, 62:S2-8.
Standard montages recommended by the American Clinical Neurophysiology Society:
Download and installation
- Requirements: You have already followed all the introduction tutorials and you have a working copy of Brainstorm installed on your computer.
Go to the Download page of this website, and download the file: sample_epilepsy.zip
Unzip it in a folder that is not in any of the Brainstorm folders (program folder or database folder)
- Start Brainstorm (Matlab scripts or stand-alone version)
Select the menu File > Create new protocol. Name it "TutorialEpilepsy" and select the options:
"No, use individual anatomy",
"Yes, use one channel file per subject".
Import the anatomy
Right-click on the TutorialEpilepsy folder > New subject > sepi01
- Leave the default options you set for the protocol
Right-click on the subject node > Import anatomy folder:
Set the file format: "FreeSurfer folder"
Select the folder: sample_epilepsy/anatomy
- Number of vertices of the cortex surface: 15000 (default value)
- Set the 6 required fiducial points (indicated in MRI coordinates):
- NAS: x=135, y=222, z=75
- LPA: x=57, y=118, z=68
- RPA: x=204, y=119, z=76
- AC: x=131, y=145, z=110
- PC: x=130, y=119, z=111
- IH: x=128, y=134, z=170 (anywhere on the midsagittal plane)
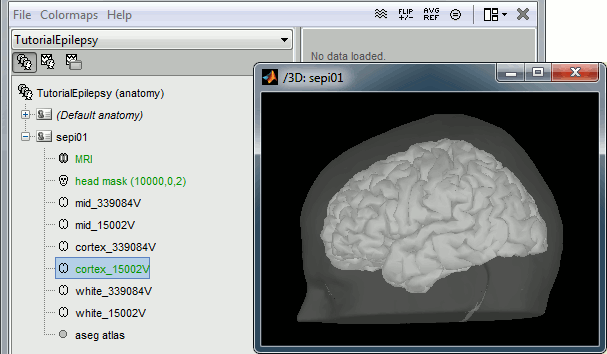
Without the individual MRI
If you do not have access to an individual MR scan of the subject (or if its quality is too low to be processed with FreeSurfer), but if you have digitized the head shape of the subject using a tracking system, you have an alternative: deform one of the Brainstorm templates (Colin27 or ICBM152) to match the shape of the subject's head.
For more information, read the following tutorial: Warping default anatomy
Access the recordings
Link the recordings
- Switch to the "functional data" view.
Right-click on the subject folder > Review raw file:
- Select the file format: "EEG: Deltamed Neurofile-Coherence (*.bin)"
Select the file: sample_epilepsy/data/tutorial_EEG.bin
- The new file "Link to raw file" lets you access directly the contents of the EEG recordings
- The channel file "Deltamed channels" in the (Common files) folder contains the name of the channels, but not their positions. We need to overwrite this file and import manually the positions of the electrodes (either a standard cap or accurate positions digitized with a Polhemus device).
Prepare the channel file
Right-click on the subject folder > Import channel file:
- Select the file format: "EEG: ANT Xensor (*.elc)"
Select the file: sample_epilepsy/data/tutorial.elc
- Confirm that you want to overwrite the exising channel file.
- This file contains the default electrodes positions from the ASA software (ANT)
- The recordings contain signals coming from different types of electrodes:
- 29 EEG electrodes
- EOG1, EOG2: Electrooculograms
- EMG, ECG: Electromyogram and electrocardiogram
- SP1, SP2: Sphenoidal electrodes
- RS: Electrode on the right shoulder
- PHO: Photo stimulation channel
- DELR, DELL, QR, QL: Additional
- The file format for the electrodes positions does not describe the type of the channels correctly, therefore all the signals saved in the files are classified as EEG. We need to redefine this manually to get correct groups of sensors, we want to have only real EEG electrodes in the "EEG" category and put everything that we are not going to use a "MISC" category.
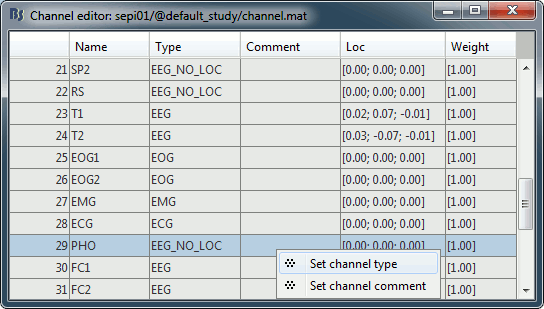
Right-click on the channel file > Edit channel file:
- Note that the EOG, EMG and ECG channels already have a different type
- Select all the other non-EEG channels: SP1, SP2, RS, PHO, DELR, DELL, QR, QL
Right-click in the window > Set channel type: type MISC
- Close the figure and accept to save the modifications
Register electrodes with MRI
- The channel file we imported contains generic electrodes positions, hence it cannot be properly aligned properly with the head surface coming from the MRI. We need to register manually these electrodes positions with the subject anatomy.
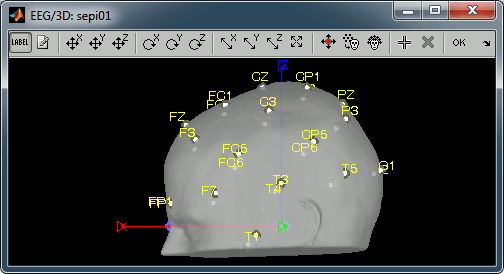
Right-click on the channel file > MRI registration > Edit...
- The default positions are already quite good, and the head shape is correct, there not much manual registration to do. You can click on the Label button in the toolbar to show the electrodes names.
- Click on the button "Refine registration with the head points" to find a better registration between the head shape defined by the electrodes and the head surface coming from the MRI.
- Click on the button "Project electrodes on the scalp surfaces", to ensure all the electrodes touch the skin surface.
Click on "OK" to save the modifications.
Review recordings to find the spikes
Display the recordings in one or more montages
For basic information on how to manipulate the montages in Brainstorm: read this tutorial.
Open a time series figure with the "Longitudinal 1"montage
Right-click on the "Link to raw file" > EEG > Display time series.
- In the Record tab, select the first button in the toolbar (Display mode for time series) to view the signals in columns.
- In the same toolbar, select the "Longitudinal 1" in the drop-down list. This menu allows you to set a predefined montages to linearly re-arranging the waveforms.
- If you don't see the "Longitudinal 1" menu, it is because you ran Brainstorm for the first time before those predefined montages were made available in Brainstorm. To add them manually:
- In the Record tab, select "Edit montages" in the drop-down menu
- Click on the "Load montage" button
- Go to the folder "brainstorm3/toolbox/sensors/private/", and select the first file
- Note that a new entry (probably "Longitudinal 1") is added to the list of available montages
- Repeat the operation with all the files in the folder "brainstorm3/toolbox/sensors/private/"
- Click on "Save" to close the montage editor and now select "Longitudinal 1"
Open another view on the same data with the "Transversal 1" montage:
Right-click on the "Link to raw file" again > EEG > Display time series
Alternatively, you can right-click on the existing figure > Figure > Clone figure.
- Set the montage for this new figure to "Transversal 1" (note the keyboard shortcuts in the menus)
- Open a 2D map of the sensor values as well:
Right-click on "Link to raw file" again > EEG > 2D Sensor cap
In the Record tab, set the Montage to this view to "Average reference"
Re-arrange the figures in a convenient way, for example as illustrated below, and disable the automatic positioning of the figures you the figure arrangement doesn't get lost when you open a new figure (Window layout menu at the top-right of the Brainstorm figure > None)
=> ECG/EMG/EOG ???
=> SCREEN CAPTURE ???
Frequency filters
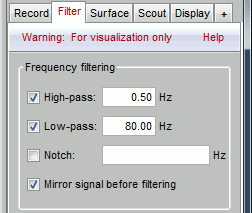
Go to the Filter tab to enable some display frequency filters. General recommendations are:
High-pass filter: 0.5 Hz
Low-pass filter: 70 Hz
Time and amplitude resolution
The resolutions of the time and amplitude axes has a lot of importance for the visual detection of epileptic spikes. The shapes we are looking for are altered by the horizontal and vertical scaling. The distance unit on a screen is the pixel, we can set how much time is represented by one pixel horizontally and how much amplitude is represented by one pixel vertically.
In the Brainstorm interface, this resolution is usually set implicitly: you can set the size of the window, the duration or recordings reviewed at once (text box "duration" in tab Record) and the maximum amplitude to show in the figure (buttons [...] and [AS] on the right of the time series figure). From there, you can also zoom in time ([<], [>], mouse wheel) or amplitude ([^], [v], Shift+mouse wheel). Those parameters are convenient to explore the recordings interactively but don't allow us to have reproducible displays with constant time and amplitude resolutions.
To set the figure resolutions: right-click on the figure > Figure > Set axes resolution. Note that this interface does not store the input values, it just modifies the other parameters (figure size, time window, max amplitude) to fit the resolution objectives. Then if you modify those parameters (resize the figure, leave the button [AS] selected and scroll in time...) the resolution is lost, you have to set it again manually.
Recommendations for this dataset are:
- Time axis: 90 pixels/sec (~30 mm/sec)
- Amplitude: 4 μV/pixels (~12 μV/mm)
=> CHECK THE VALUES ???
=> SCREEN CAPTURE ???
User setups
This preparation of the reviewing environment requires a large number of operations, and would become quickly annoying if you have to repeat it everytime you open a file. This is a good time to use the menu "User setups" to save this window configuration, to reload it in one click later. In the menu "Window layout", at the top-right of the Brainstorm window, select User setup > New setup. Enter a name of your choice for this particular window arrangement.
This operation will also automatically disable the automatic window arrangement (Window layout > None). To reload it later, open one figure on the dataset you want to review and then select your new entry in the User setup menu.
Mark spikes
In the dataset, the spikes have already been identified by experts at the University Hospital of Freiburg. You can see that 177 SPIKE events are available in the Record tab. Click on a few of them a try to identify the shape of the spike.
=> SCREEN CAPTURE ???
The procedure if you are marking the events by yourself is the following:
- Open a continuous file viewer, the load your reviewing environment (menu User setups)
Start by creating a group of events (in the Record tab: Events > Add group), and select it in the list of events.
Make sure that the time and amplitude resolutions are what you are used to (right-click on the figure > Figure > Set axes resolution)
Scroll through the recordings using the [<<<] and [>>>] buttons or the associated shortcuts such as F3 or F4 (complete list and descriptions available when you leave your mouse over those buttons).
You can adjust the gain of the electrodes to observe better an event with the buttons [^] and [v] or the shortcut [Shift+mouse wheel].
- When you identify a spike, click in a white area of the figure in order to place the time cursor at the peak of the spike. If you click on the signal itself, it selects the corresponding channel, but you can use the shortcut Shift+Click to prevent this behavior and force the time cursor to be moved instead.
- Press Ctrl+E to add a marker where the time cursor is.
If you are marking multiple types of events, it is convenient to set up some additional keyboard shortcuts. Using the menu Events > Edit keyboard shortcuts, you can associate custom events to the key 1 to 9 of the keyboard. Define the name of the event type to create for each key, and then simply press the corresponding key to add/delete a marker at the current time position.
More information on the data viewer, see tutorial: ?Review continuous recordings.
=> CONFIRM THE PROCEDURE ???
Pre-process recordings
- High-pass filter the recording at 0.5Hz
=> APPLY 0.5 Hz FILTER TO THE CONTINUOUS RECORDINGS BEFORE IMPORTING ???
=> (NOT POSSIBLE WITH THIS FILE FORMAT)
Import recordings
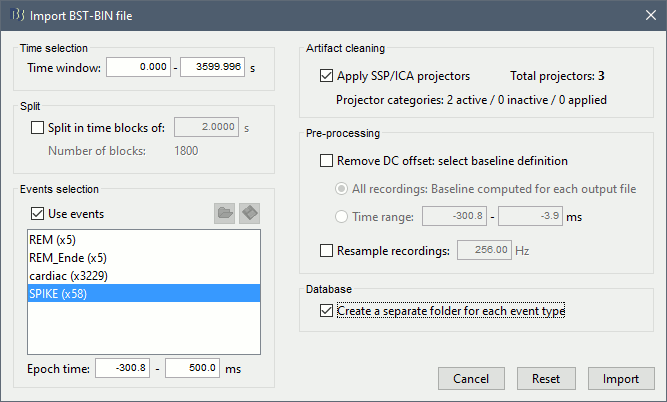
Right-click on the "Link to raw file" > Import in database
Select the option "Use events" and select the category SPIKE, with 177 events.
Epoch time: [-100, +300]ms around the spike events
Uncheck the option "Remove DC offset" because the DC offset has already been removed with the high-pass filter
Source analysis
- The pial surface will be used for this modeling.
- Compute head model: OpenMEEG
- Noise covariance matrix: select 10 sec time window from the raw file which would represent the subject's resting state with no apparent epileptic activity - use this for all runs
- Source model: wMNE, constrained orientations
- Compute z-score
- Discuss advantage of distributed source models in the evaluation of epilepsy data compared with ECD (Kobayashi K, Epilepsia 2005)
=> WHAT SEGMENT OF 10s CAN WE USE FOR THE NOISE COVARIANCE ???
Moving dipoles
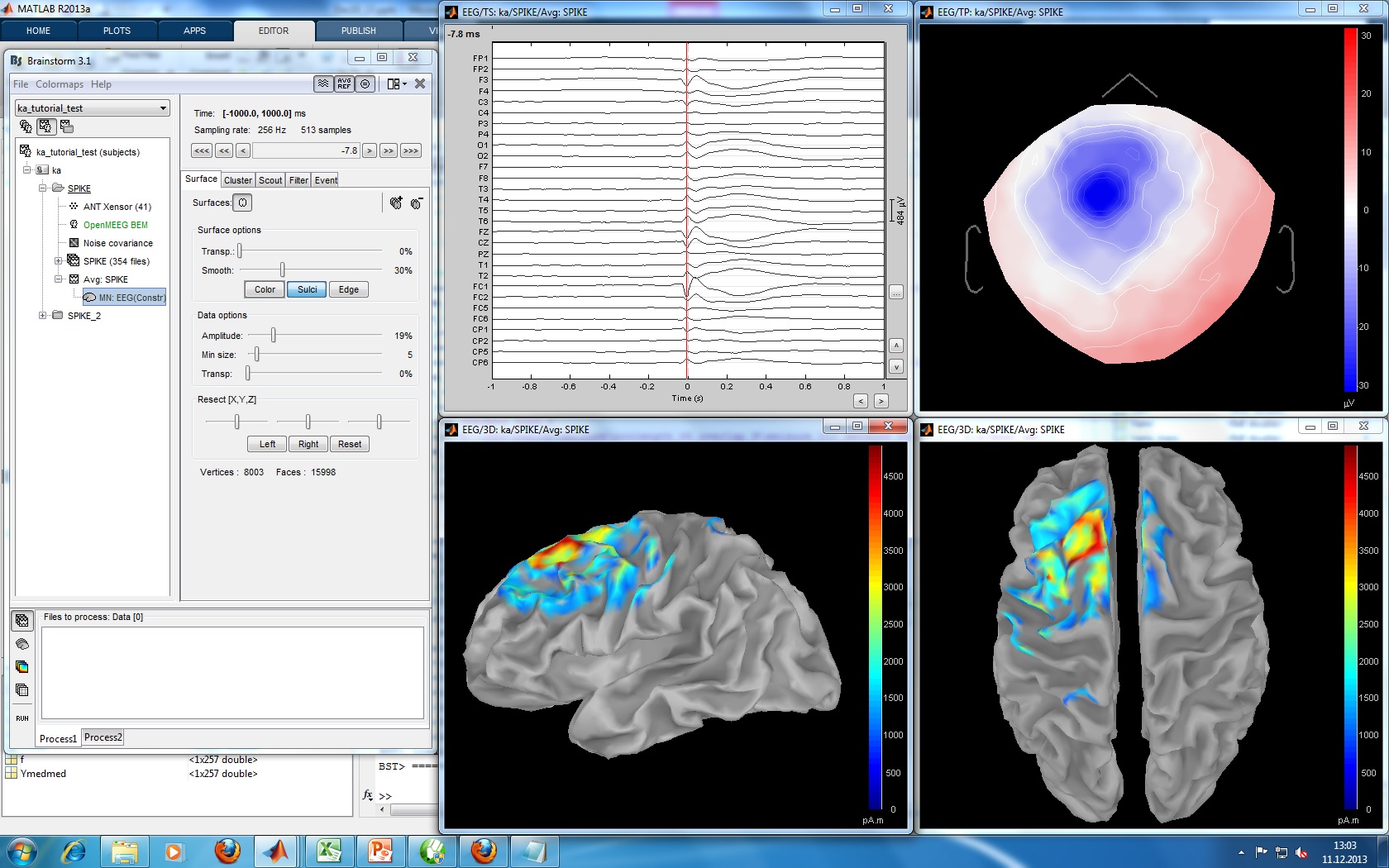
Illustrate John/Beth's tools for calculating and displaying dipoles.
=> DOES THIS WORK ???