|
Size: 2185
Comment:
|
Size: 13793
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Tutorial 12: Explore the average response = | = Tutorial 17: Visual exploration = |
| Line 4: | Line 4: |
| This tutorial illustrates the options Brainstorm offers to represent graphically and explore interactively the evoked responses we computed in the previous tutorial. It shows how to produce spatial maps of the sensors, temporal averages, save screen captures and movies. |
|
| Line 6: | Line 8: |
| From continuous tutorials: == Online filter == As introduced in the previous tutorials, you can add an online visualization filter to make the traces and the topographies look smoother. We suggest in this case a low-pass filter at 120Hz, because it shows very smooth traces, but all the waves we are interested in are still clearly visible in the MEG time series figure. {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawAvg?action=AttachFile&do=get&target=onlineFilter.gif|onlineFilter.gif|class="attachment"}} == Explore the average == Open the time series for the "'''Avg: left'''". Then press '''Control+T''', to see on the side a spatial topography at the current time point. Then observe what is happening between 0ms and 100ms. Start at 0ms and play it like a movie using the arrow keys left and right, to follow the brain activity millisecond by millisecond: * '''16 ms''': First response, the sensory information reaches the right somatosensory cortex (S1) * '''30 ms''': Stronger and more focal activity in the same region, but with a source oriented in the opposite direction (S1) * '''60 ms''': Activity appearing progressively in a more lateral region (S1 + S2) * '''70 ms''': Activity in the same area in the left hemisphere (S2 left + S2 right) {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawAvg?action=AttachFile&do=get&target=avg16.gif|avg16.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawAvg?action=AttachFile&do=get&target=avg30.gif|avg30.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawAvg?action=AttachFile&do=get&target=avg60.gif|avg60.gif|class="attachment"}} {{http://neuroimage.usc.edu/brainstorm/Tutorials/TutRawAvg?action=AttachFile&do=get&target=avg70.gif|avg70.gif|class="attachment"}} <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/Averaging&next=Tutorials/HeadModel")>> |
== 2D/3D topography == The sensor values at one time instant can be represented on a surface. Each amplitude value gets associated with a color using a colormap (described in the next tutorial). We call this type of representation "sensor topography", it shows the spatial distribution of the magnetic fields (or electric potentials). * Show the MEG signals for the '''standard average''' in '''Run#01''' (double-click on the file). This gives us a direct feedback of the current time instant and allows to jump quickly to a different time. * Right-click on the same file > MEG > select the menus: '''3D sensor cap''', '''2D sensor cap''', '''2D disc'''. <<BR>><<BR>> {{attachment:explore_topo.gif||height="281",width="669"}} * '''3D sensor cap''': Represents the real 3D positions of the sensors. * '''2D sensor cap''': The sensors are projected on a 2D plane. Realistic distribution of the sensors. * '''2D disc''': The sensors are projected on a sphere, then displayed as flat disc. Sometimes distorted... * In each of these views, you can add markers to indicate the sensors and their labels.<<BR>>Right-click on the figure > Channels > '''Display sensors/labels''' (or Ctrl+E). * You can change the number of contour lines: Right-click > '''Contour lines''' > 0-20.<<BR>><<BR>> {{attachment:explore_contours.gif||height="145",width="647"}} * In the 3D view, you can notice a hole in the right-occipital area. It corresponds to a damaged sensor in the MEG system we used for collecting this dataset. * For '''EEG/sEEG/ECoG''' recordings, there is an additional representation mode available: <<BR>>"'''3D Electrode'''". This will be detailed in the advanced tutorials corresponding to these modalities. <<BR>><<BR>> {{attachment:explore_3delectrode.gif||height="180",width="584"}} <<TAG(Advanced)>> == Magnetic interpolation == By default, some of the views re-interpolate the fields that are recorded by the MEG sensors to get smoother displays. A simple inverse problem and forward problem are solved to reconstruct the magnetic fields on a high-resolution surface of virtual magnetometers (function channel_extrapm.m). On Elekta-Neuromag systems, this interpolation has the effect of converting the topographies of the planar gradiometers into topographies of magnetometers, which deeply affects the display. The menu '''"No magnetic interpolation"''' offers the same views, but without using this reconstruction of the magnetic field. A spatial interpolation of the values between the sensors is performed instead. . {{attachment:explore_nointerp.gif||height="251",width="351"}} <<TAG(Advanced)>> == 2D Layout == The menu 2D Layout represents, in the same figure, the '''spatial information''' (the values for each channel is represented where the sensor is actually located) and the '''temporal information''' (instead of just one single value, we represent the signal around the current time). <<BR>>The light gray lines represent the zero amplitude (horizontal) and the current time (vertical lines). Only a part of the full time window is displayed for each channel, before and after the current time. The length of this time window can be modified either with the mouse shortcut '''Ctrl+mouse wheel''', or with the 2D Layout options, in the figure popup menu. The amplitude of the signal can be controlled with the shortcut '''Shift+mouse wheel'''. . {{attachment:explore_2dlayout.gif||height="272",width="546"}} <<TAG(Advanced)>> == Display as image == The menu "Display as image" shows the same information as the "time series" view, but the values for each sensor are represented with a color instead of a line. . {{attachment:explore_image.gif||height="200",width="660"}} <<TAG(Advanced)>> == Time selection == Click somewhere on the white part of the time series figure, hold the mouse button, and drag your mouse left or right: A transparent blue rectangle appears to represent the time selection. If you right-click on the figure, new options become available in the popup menu: * '''Set current time''': Move the time cursor where the right-click occurred. The shortcut '''Shift+Click''' can be useful when trying to move in time on dense displays in columns view. * '''Set selection manually''': Type the beginning and end of the selected window (in milliseconds). * '''Average time''': Average over the selected time window and save it as a new file in the database. <<BR>>Note that the best way to do this is to run the process "'''Average > Average time'''". * '''Export to database''': Extract the recordings and save them in a new file in the database.<<BR>>If some sensors are selected, only their values are extracted, all the others are set to zero.<<BR>>Note that the best way to do this is to run the process "'''Extract > Extract time'''". * '''Export to file''': Same, but in a user-defined file (not in the database). * '''Export to Matlab''': Same, but export the selection as a variable in the current Matlab workspace. <<BR>><<BR>> {{attachment:explore_timeselect.gif||height="200",width="434"}} <<TAG(Advanced)>> == Snapshots == Using Brainstorm, you will quickly feel like saving the beautiful images you produce. Your operating system already provides some nice tools for doing this. Many other options are available in the "Snapshot" menu, accessible with a right-click on any Brainstorm figure. ==== Operating system ==== * '''Windows/Linux''': Press the '''PrintScreen''' key on your keyboard and paste the copied screen in your favorite image or text editor. The combination '''Alt+PrintScreen''' only copies the figure that is currently selected. * '''MacOS''': Many more options available, Google for the best ones ([[http://graphicssoft.about.com/od/screencapturemac/ht/macscreenshot.htm|see example]]). ==== Snapshot menu ==== * The options available in the Snapshot menu depend on the type of data represented. Examples: <<BR>><<BR>> {{attachment:explore_snapshot.gif||height="262",width="614"}} * '''Save as image''': Save the figure in a file, without the title bar and borders. Many formats available. * '''Open as image''': Capture the figure and open it in as an image. This can be useful if you want to visually compare the selected figure with another one that you cannot display at the same time (because they have different time or frequency definitions). * '''Open as figure''': Similar, but copies the figure as a new Matlab figure with some interactivity. * '''Contact sheet and movies''': See next section. * '''Export to database''': Save the recordings in the figure as a new entry in the database. <<BR>>If there are selected channels, only their values will be saved, the others being set to zero. * '''Export to file''': Extract the time series displayed in this figure (or only the selected sensors), and save them in a file. Several exchange file formats available for exporting to another program. * '''Export to Matlab''': Same thing, but exports the structure in a variable of the Matlab workspace. * '''Save as SSP projector''': Create an SSP projector that removes the current topography. * '''Save surface''': Save the surface in a file, with the current modifiers applied (smooth, resect). <<TAG(Advanced)>> == Movie studio == * '''Movie (horizontal/vertical)''': Rotate spatially the 3D scene. * '''Movie (time): Selected figure''': Create .avi movies to show the evolution of the selected figure. * The dimensions of the movie depend on the actual size of the figure on the screen. <<BR>>Resize the figure to the appropriate dimensions for the movie before using this menu. * Zoom in/out (mouse wheel) and move the image (middle click+move) to give enough space to the time stamp that is added at the bottom-left of the rendered movie. * Don't do anything else while rendering: the captured figure must be visible all the time. * '''Movie (time): All figures''': Instead of capturing one figure only, it captures them all. Arrange your figures the way you want and create a movie of all your workspace at once.<<BR>><<BR>> {{attachment:movie_options.gif||height="285",width="139"}} {{attachment:movie.gif}} <<TAG(Advanced)>> == Contact sheets == A contact sheet is a large image representing many time frames of the same figure. * Same recommendations as for movies: if you don't want the final image to be too big, reduce the size of the figure, zoom in, move, hide the colorbar. Keep the figure visible during the capture. * At the end, the image is displayed in an image viewer with which you can zoom (menu or wheel), move (click+move) and save the image (File > Save as). * Example for the '''standard average''', run#01: <<BR>><<BR>> {{attachment:contact_options.gif||height="220",width="135"}} {{attachment:contact_standard.gif||height="283",width="340"}} <<TAG(Advanced)>> == Edit the figures == All the figures can be edited with the Figure popup menu: . {{attachment:explore_figure.gif||height="319",width="403"}} If you select both "'''Matlab controls'''" and "'''Plot edit toolbar'''", you will get all the tools available in the Matlab environment to explore the data and edit the figure. Select the button "'''Edit plot'''" to edit the graphic properties of an object (eg. select a signal, then right-click on it to edit its properties) or unselect it to get back to the regular Brainstorm figure interactivity. . {{attachment:figure_edit.gif||height="215",width="326"}} <<TAG(Advanced)>> == Mouse shortcuts == '''Scroll''' * '''Mouse wheel''': Zoom in / zoom out * '''Control + mouse wheel''': Change the length of the displayed time window (2D Layout) * '''Control + mouse wheel''': Vertical zoom (time series) '''Click''' * '''Left click + move''': Rotate (3D) or select (time) * '''Middle click + move''': Move in zoomed figure (ie. panning) * '''Left click + right click + move''': Move in zoomed figure (ie. panning) * '''Shift + left click''': Force setting the current time, ignoring if a line was clicked (time series) * '''Right click + move''': Vertical zoom (time series) * '''Right click + move''': Select sensors (2D topography) * '''Right click''': Popup menu * '''Double-click''': Restore initial view '''Click on something''' * '''Click on a line''': Select a sensor * '''Shift + click on a line''': Select a sensor and unselect all the others (2D topography) * '''Click on the colorbar + move''': Change contrast (up/down) and brightness (left/right) <<TAG(Advanced)>> == Keyboard shortcuts == Here is a memo of all the keyboard shortcuts for time series and topography figures. If you don't remember them, you can find most of them in the figure popup menus. * '''Arrows: '''Left, right, PageUp, PageDown: Move in time * '''Delete''': Mark selected sensors as ''bad'' * '''Shift + ''''''Delete''': Mark non-selected sensors as ''bad'' (=keeps ony the selected sensors) * '''Enter''': View time series for the selected sensors * '''Escape''': Unselect all the selected sensors * '''Shift + Escape''': Set all the bad sensors as good (=brings back all the channels in the display) * '''Ctrl + A''': Show axis on 3D figures (X,Y,Z) * '''Ctrl + B''': Set trial as bad * '''Ctrl + D''': Dock/undock figure in Matlab's figures list * '''Ctrl + E''': Show sensors markers and labels (E=''Electrode'') or add an event marker (E=''Event'') * '''Ctrl + F''': Copy figure, removes all the callbacks and detach from Brainstorm figure management * '''Ctrl + I''': Save figure as image * '''Ctrl + J''': Open figure as an image * '''Ctrl + R''': Open ''Time series'' view (R=''Recordings'') * '''Ctrl + S''': Open ''Sources'' view (S=''Sources'') * '''Ctrl + T''': Open ''2D sensor cap'' view (T=''Topography'') * '''Shift + letter''': Change selected montage * '''F1, F2, F3''': with or without '''Shift''', calls the database navigator (F1=subject, F2=condition, F3=file) * '''1, 2, 3, 4, 5, 6, 7, 8, 9, 0''': Set a pre-defined 3D view * '''+ / -''': Increase/decrease the channel gain (vertical zoom for time series) * '''=''': Apply view to all figures * '''*''': Apply montage to all figures * Notes for Mac users: * PageDown = '''Fn''' + DOWN * PageUp = '''Fn''' + UP * F1 = '''Fn''' + F1 * Mouse wheel = Two finger up/down on the MacBook pad <<TAG(Advanced)>> == Graphic bugs == If you observe any graphic problem with these displays, there might be an issue with the OpenGL drivers. You may try the following options. * Update the drivers for your graphics card. * Upgrade your version of Matlab. * Run the compiled version of Brainstorm (see [[http://neuroimage.usc.edu/brainstorm/Installation|Installation]]). * Turn off the OpenGL hardware acceleration: Menu File > Edit preferences > ''Software'' or ''Disabled''. * Send a bug report to the Mathworks. <<HTML(<!-- END-PAGE -->)>> <<EmbedContent("http://neuroimage.usc.edu/bst/get_prevnext.php?prev=Tutorials/Averaging&next=Tutorials/Colormaps")>> |
Tutorial 17: Visual exploration
Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet
This tutorial illustrates the options Brainstorm offers to represent graphically and explore interactively the evoked responses we computed in the previous tutorial. It shows how to produce spatial maps of the sensors, temporal averages, save screen captures and movies.
Contents
2D/3D topography
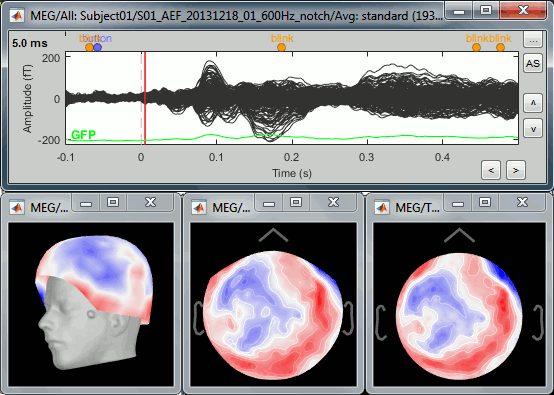
The sensor values at one time instant can be represented on a surface. Each amplitude value gets associated with a color using a colormap (described in the next tutorial). We call this type of representation "sensor topography", it shows the spatial distribution of the magnetic fields (or electric potentials).
Show the MEG signals for the standard average in Run#01 (double-click on the file). This gives us a direct feedback of the current time instant and allows to jump quickly to a different time.
Right-click on the same file > MEG > select the menus: 3D sensor cap, 2D sensor cap, 2D disc.
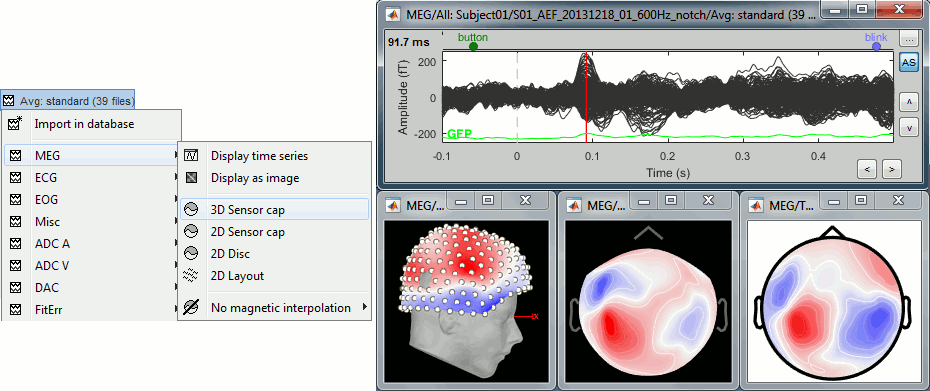
3D sensor cap: Represents the real 3D positions of the sensors.
2D sensor cap: The sensors are projected on a 2D plane. Realistic distribution of the sensors.
2D disc: The sensors are projected on a sphere, then displayed as flat disc. Sometimes distorted...
In each of these views, you can add markers to indicate the sensors and their labels.
Right-click on the figure > Channels > Display sensors/labels (or Ctrl+E).You can change the number of contour lines: Right-click > Contour lines > 0-20.
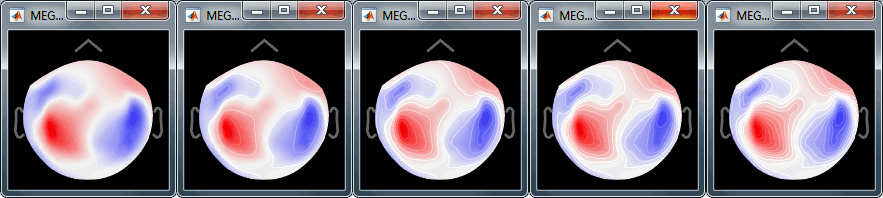
- In the 3D view, you can notice a hole in the right-occipital area. It corresponds to a damaged sensor in the MEG system we used for collecting this dataset.
For EEG/sEEG/ECoG recordings, there is an additional representation mode available:
"3D Electrode". This will be detailed in the advanced tutorials corresponding to these modalities.
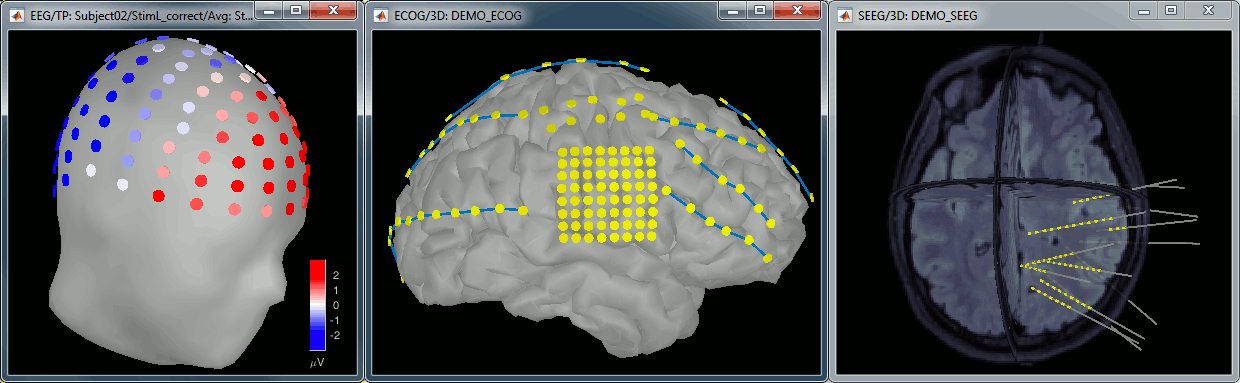
Magnetic interpolation
By default, some of the views re-interpolate the fields that are recorded by the MEG sensors to get smoother displays. A simple inverse problem and forward problem are solved to reconstruct the magnetic fields on a high-resolution surface of virtual magnetometers (function channel_extrapm.m).
On Elekta-Neuromag systems, this interpolation has the effect of converting the topographies of the planar gradiometers into topographies of magnetometers, which deeply affects the display.
The menu "No magnetic interpolation" offers the same views, but without using this reconstruction of the magnetic field. A spatial interpolation of the values between the sensors is performed instead.
2D Layout
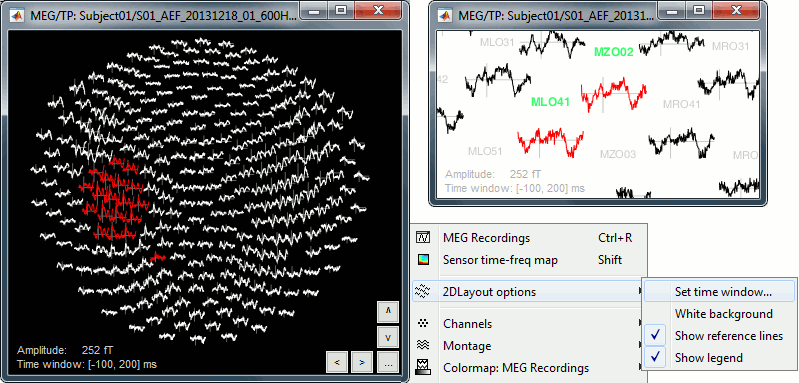
The menu 2D Layout represents, in the same figure, the spatial information (the values for each channel is represented where the sensor is actually located) and the temporal information (instead of just one single value, we represent the signal around the current time).
The light gray lines represent the zero amplitude (horizontal) and the current time (vertical lines).
Only a part of the full time window is displayed for each channel, before and after the current time. The length of this time window can be modified either with the mouse shortcut Ctrl+mouse wheel, or with the 2D Layout options, in the figure popup menu. The amplitude of the signal can be controlled with the shortcut Shift+mouse wheel.
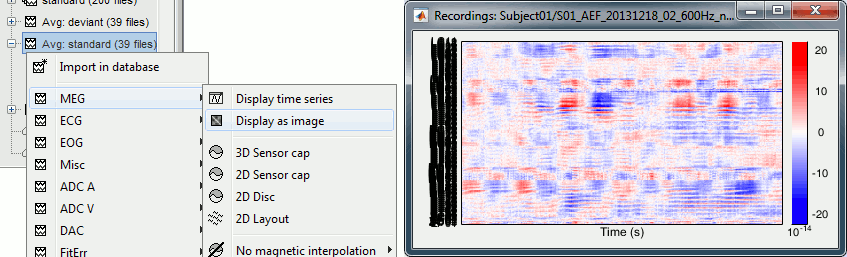
Display as image
The menu "Display as image" shows the same information as the "time series" view, but the values for each sensor are represented with a color instead of a line.
Time selection
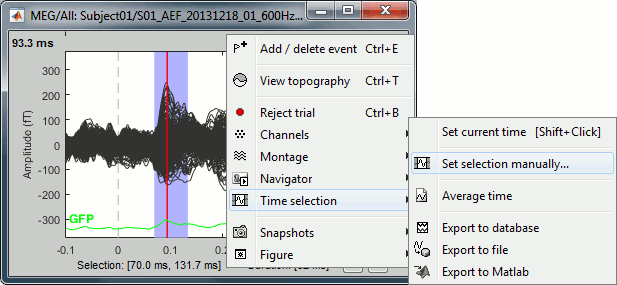
Click somewhere on the white part of the time series figure, hold the mouse button, and drag your mouse left or right: A transparent blue rectangle appears to represent the time selection. If you right-click on the figure, new options become available in the popup menu:
Set current time: Move the time cursor where the right-click occurred. The shortcut Shift+Click can be useful when trying to move in time on dense displays in columns view.
Set selection manually: Type the beginning and end of the selected window (in milliseconds).
Average time: Average over the selected time window and save it as a new file in the database.
Note that the best way to do this is to run the process "Average > Average time".Export to database: Extract the recordings and save them in a new file in the database.
If some sensors are selected, only their values are extracted, all the others are set to zero.
Note that the best way to do this is to run the process "Extract > Extract time".Export to file: Same, but in a user-defined file (not in the database).
Export to Matlab: Same, but export the selection as a variable in the current Matlab workspace.
Snapshots
Using Brainstorm, you will quickly feel like saving the beautiful images you produce. Your operating system already provides some nice tools for doing this. Many other options are available in the "Snapshot" menu, accessible with a right-click on any Brainstorm figure.
Operating system
Windows/Linux: Press the PrintScreen key on your keyboard and paste the copied screen in your favorite image or text editor. The combination Alt+PrintScreen only copies the figure that is currently selected.
MacOS: Many more options available, Google for the best ones (see example).
Snapshot menu
The options available in the Snapshot menu depend on the type of data represented. Examples:
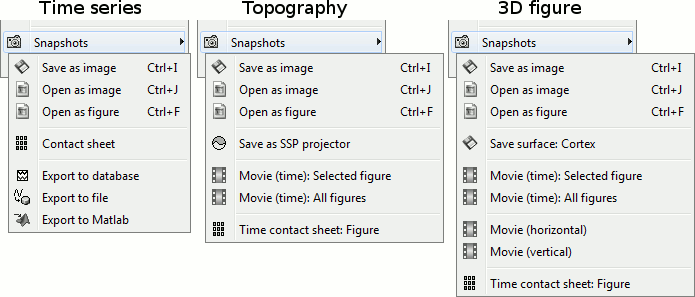
Save as image: Save the figure in a file, without the title bar and borders. Many formats available.
Open as image: Capture the figure and open it in as an image. This can be useful if you want to visually compare the selected figure with another one that you cannot display at the same time (because they have different time or frequency definitions).
Open as figure: Similar, but copies the figure as a new Matlab figure with some interactivity.
Contact sheet and movies: See next section.
Export to database: Save the recordings in the figure as a new entry in the database.
If there are selected channels, only their values will be saved, the others being set to zero.Export to file: Extract the time series displayed in this figure (or only the selected sensors), and save them in a file. Several exchange file formats available for exporting to another program.
Export to Matlab: Same thing, but exports the structure in a variable of the Matlab workspace.
Save as SSP projector: Create an SSP projector that removes the current topography.
Save surface: Save the surface in a file, with the current modifiers applied (smooth, resect).
Movie studio
Movie (horizontal/vertical): Rotate spatially the 3D scene.
Movie (time): Selected figure: Create .avi movies to show the evolution of the selected figure.
The dimensions of the movie depend on the actual size of the figure on the screen.
Resize the figure to the appropriate dimensions for the movie before using this menu.- Zoom in/out (mouse wheel) and move the image (middle click+move) to give enough space to the time stamp that is added at the bottom-left of the rendered movie.
- Don't do anything else while rendering: the captured figure must be visible all the time.
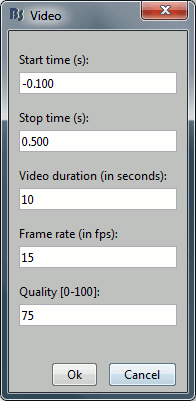
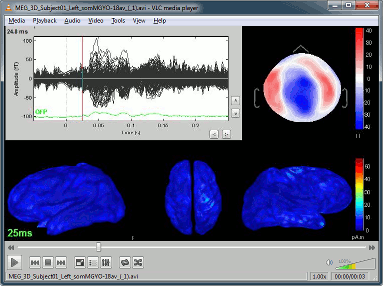
Movie (time): All figures: Instead of capturing one figure only, it captures them all. Arrange your figures the way you want and create a movie of all your workspace at once.
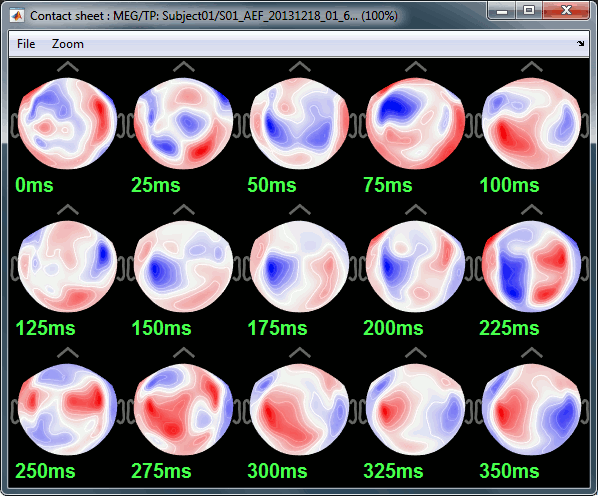
Contact sheets
A contact sheet is a large image representing many time frames of the same figure.
- Same recommendations as for movies: if you don't want the final image to be too big, reduce the size of the figure, zoom in, move, hide the colorbar. Keep the figure visible during the capture.
At the end, the image is displayed in an image viewer with which you can zoom (menu or wheel), move (click+move) and save the image (File > Save as).
Example for the standard average, run#01:
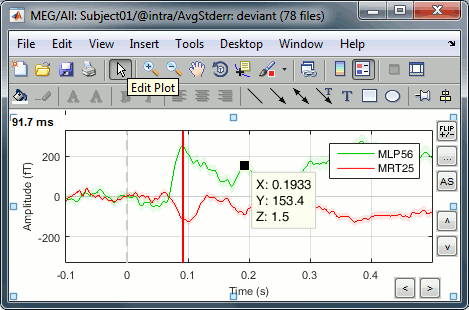
Edit the figures
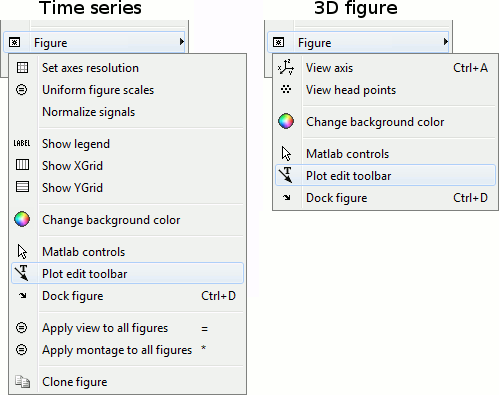
All the figures can be edited with the Figure popup menu:
If you select both "Matlab controls" and "Plot edit toolbar", you will get all the tools available in the Matlab environment to explore the data and edit the figure. Select the button "Edit plot" to edit the graphic properties of an object (eg. select a signal, then right-click on it to edit its properties) or unselect it to get back to the regular Brainstorm figure interactivity.
Mouse shortcuts
Scroll
Mouse wheel: Zoom in / zoom out
Control + mouse wheel: Change the length of the displayed time window (2D Layout)
Control + mouse wheel: Vertical zoom (time series)
Click
Left click + move: Rotate (3D) or select (time)
Middle click + move: Move in zoomed figure (ie. panning)
Left click + right click + move: Move in zoomed figure (ie. panning)
Shift + left click: Force setting the current time, ignoring if a line was clicked (time series)
Right click + move: Vertical zoom (time series)
Right click + move: Select sensors (2D topography)
Right click: Popup menu
Double-click: Restore initial view
Click on something
Click on a line: Select a sensor
Shift + click on a line: Select a sensor and unselect all the others (2D topography)
Click on the colorbar + move: Change contrast (up/down) and brightness (left/right)
Keyboard shortcuts
Here is a memo of all the keyboard shortcuts for time series and topography figures. If you don't remember them, you can find most of them in the figure popup menus.
Arrows: Left, right, PageUp, PageDown: Move in time
Delete: Mark selected sensors as bad
Shift + Delete: Mark non-selected sensors as bad (=keeps ony the selected sensors)
Enter: View time series for the selected sensors
Escape: Unselect all the selected sensors
Shift + Escape: Set all the bad sensors as good (=brings back all the channels in the display)
Ctrl + A: Show axis on 3D figures (X,Y,Z)
Ctrl + B: Set trial as bad
Ctrl + D: Dock/undock figure in Matlab's figures list
Ctrl + E: Show sensors markers and labels (E=Electrode) or add an event marker (E=Event)
Ctrl + F: Copy figure, removes all the callbacks and detach from Brainstorm figure management
Ctrl + I: Save figure as image
Ctrl + J: Open figure as an image
Ctrl + R: Open Time series view (R=Recordings)
Ctrl + S: Open Sources view (S=Sources)
Ctrl + T: Open 2D sensor cap view (T=Topography)
Shift + letter: Change selected montage
F1, F2, F3: with or without Shift, calls the database navigator (F1=subject, F2=condition, F3=file)
1, 2, 3, 4, 5, 6, 7, 8, 9, 0: Set a pre-defined 3D view
+ / -: Increase/decrease the channel gain (vertical zoom for time series)
=: Apply view to all figures
*: Apply montage to all figures
- Notes for Mac users:
PageDown = Fn + DOWN
PageUp = Fn + UP
F1 = Fn + F1
Mouse wheel = Two finger up/down on the MacBook pad
Graphic bugs
If you observe any graphic problem with these displays, there might be an issue with the OpenGL drivers. You may try the following options.
- Update the drivers for your graphics card.
- Upgrade your version of Matlab.
Run the compiled version of Brainstorm (see Installation).
Turn off the OpenGL hardware acceleration: Menu File > Edit preferences > Software or Disabled.
- Send a bug report to the Mathworks.