Synchronization with eye tracker (event transfer)
Authors: Martin Völker, Francois Tadel, Raymundo Cassani
This tutorial introduces how to transfer events from one data set to a second data set of a different type, which was recorded at the same time but with a different sampling rate. We will explain this on the example of synchronizing ocular data recorded with an EyeLink eye tracker (SR Research, Canada) with a second raw recording, e.g. EEG.
Introduction
Eye tracking is useful to detect blinks, saccades, microsaccades and fixations during an experiment. Typically, the eye tracker software will itself mark the most common artifact types like blinks and saccades.
These artifact markers are very useful for the analysis of EEG / MEG data, since specific time periods which are contaminated with artifacts can be marked and excluded easily. To do so, we need to synchronize the eye tracker data with our EEG / MEG recordings.
This tutorial focuses on how to synchronize EyeLink data (.edf) with any other raw data type within brainstorm. Information about the EyeLink can be found at http://www.sr-research.com/index.html.
Requirements
For this function to work, you need specific markers, which are stored within both EEG/MEG and eye tracker data at the same time. The exact same number of this marker type has to be available in both data sets. They can, however, have different names.
Data import
First of all, the continuous MEG/EEG recording has to be linked to your Brainstorm database, if not already done (Review continuous recordings).
Then link the EyeLink .edf file in the same subject:
- Switch to the "functional data" view.
Right-click on the subject folder > Review raw file
Select the file format: "EyeLink eye tracker (*.edf)"
Select the EyeLink .edf file(s)
- Click [open]. Due to the file format, the loading process can take a while.
Event transfer
Now click on the "Process2" tab.
In FilesA: drag and drop the files from which you want to copy the events (the EyeLink files).
In FilesB: drag and drop the recordings into which you want to paste the events (the EEG files).
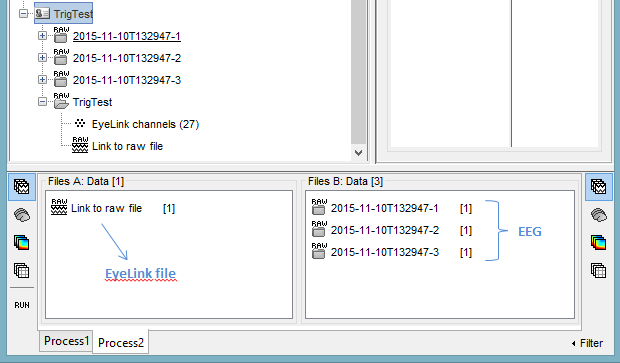
- It does not matter if the number of files in A and B is different. However, the total number of sync markers as to match exactly (number of markers in A = total number of markers in B).
Click on [RUN]. Select process "Synchronize > Transfer events (from A to B)"
In this example, the synchronization event is called "5" in the EyeLink file and "E5" in the EEG file.
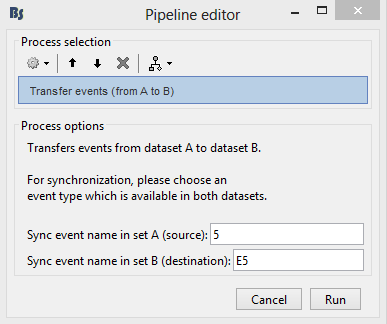
- Click on [Run]. The function will calculate the offsets between the datasets with the help of the Sync events, which are known to be occuring at the exact same time.
In the Matlab command window, the variance and standard deviations of the sample offset is displayed per file of the destination data set.
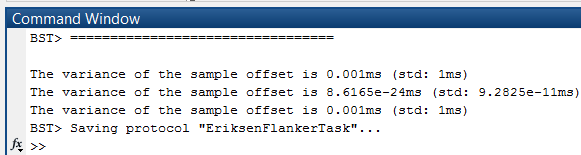
- If the variance is too high, something is wrong with the sync events. For example, when the files in "B" are in the wrong order, the offsets will differ a lot and thus have a high variance. Depending on the timing of your recording and marker system, small variations are however perfectly normal.
Verification
It is important to check if the event transfer was executed properly:
Events transferred from database A to database B will have the "sync" prefix
For no raw data files, their name will be update adding " | sync_events" at the end.
If your recordings contain EOG channels, you can e.g. check if the blinks are marked correctly.
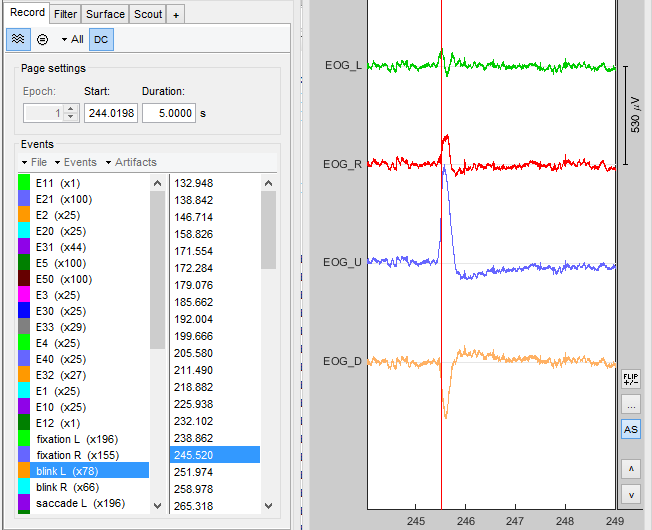
- If the two data sets share another event (besides the sync event you specified before), you can check if those are always marked at the same times.
- Control if the number of events in each file meets your expectations.
