|
Size: 17534
Comment:
|
Size: 17544
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 3: | Line 3: |
'''' |
Neuromaps plugin
Author: Le Thuy Duong Nguyen
' note: this tutorial page is currently under construction. The Neuromaps Brainstorm plugin integrates curated annotations and tools to further expand the accessibility and inclusivity of brain-mapping tools, as part of an Open Science initiative. We extend these pioneering tools to the MATLAB environment for users without any prior computer programming experience, providing an intuitive graphic user interface. The present tutorial will demonstrate the plugin’s functionality within the Brainstorm interface; for a detailed breakdown of the algorithm, please refer to the neuromaps plugin Github. Contents
Neuromaps is a toolbox designed for assessing, transforming, and analyzing structural and functional brain annotations (Markello, Hansen et al., 2022). It marks a significant stride towards integrative analytics in multimodal, multiscale neuroscience by bridging the gap between various neuroimaging modalities and proposing analytical tools to establish connections among different brain features. The toolbox comprises a curated repository of brain maps in their native space, methods for generating transformations across multiple coordinate systems, and offers a systematic workflow for comprehensive structural and functional annotation enrichment analysis of the human brain. For the first iteration of the implementation, we focused on the neurotransmitter receptors and transporters. Thirty different maps from the neuromaps toolbox were selected, covering nine different neurotransmitter systems: dopamine, norepinephrine, serotonin, acetylcholine, glutamate, GABA, histamine, cannabinoid, and opioid. These maps are sourced from open-access repositories, addressing the need for a comprehensive tool that integrates standardized analytic workflows for both surface and volumetric data.
FsAverage for surface maps- the default system used by the FreeSurfer software. MNI152 for volumetric maps- 152 normative MRI scans developed by the Montreal Neurological Institute (to be added soon). Repository of All the information for these maps, including the appropriate citations to use can be found in this spreadsheet. To obtain the surfaces, the original maps offered in MNI152 space were transformed to FsAverage using the registration fusion framework proposed in neuromaps (Buckner et al., 2011; Wu et al., 2018).
From the main window go to A message will appear saying, "Plugin 'Neuromaps' is not installed on your computer. Download the latest version of Neuromaps now?" click 'Yes'. The plugin will be downloaded and installed automatically. Once installed, you will see a confirmation message. By following these steps, you will successfully install the Neuromaps plugin.
Click on the When the Go to The brain annotations will be listed right under, in the name format '[neurotransmitter]:[subtype]_[tracer]_[source]_[size]_[age]' (e.g., 'Acetylcholine: VaCht_feobv_aghourian2017_N18_Age67'). Select all the maps you want to import and click
Once you have fetched the brain annotations, a new 'Neuromaps' node which contains the FSAverage anatomy will appear in the Anatomy view. All the imported brain annotations will be visible in the Functional data (sorted by subjects) view. To visualize the surfaces, you can either
Implementing neuromaps directly into Brainstorm further enhances the overall user experience by providing greater flexibility and control over the visualization and analysis of brain annotations. Brainstorm's advanced functionalities allow users to effortlessly The ‘Surface’ tab of the Brainstorm window contains buttons and sliders to control the display of these surfaces. Additional relevant functionalities, as outlined in Tutorial 3: Display the Anatomy, include:
The default colormap upon opening the brain annotation is royal_gramma, but you have the option to switch to other sequential, diverging, or rainbow colormaps. Simply right-click on the brain annotation window, navigate to If you modify a colormap, the changes will be applied to all the figures, saved in your user preferences and available the next time you start Brainstorm. If you wish to undo the changes and reset the colormap to its default values, click on You also have the opportunity to customize the brightness, contrast, and decide the color mapping, among other settings. For a comprehensive list of available options with detailed instructions, please refer to Tutorial 18: Colormaps.
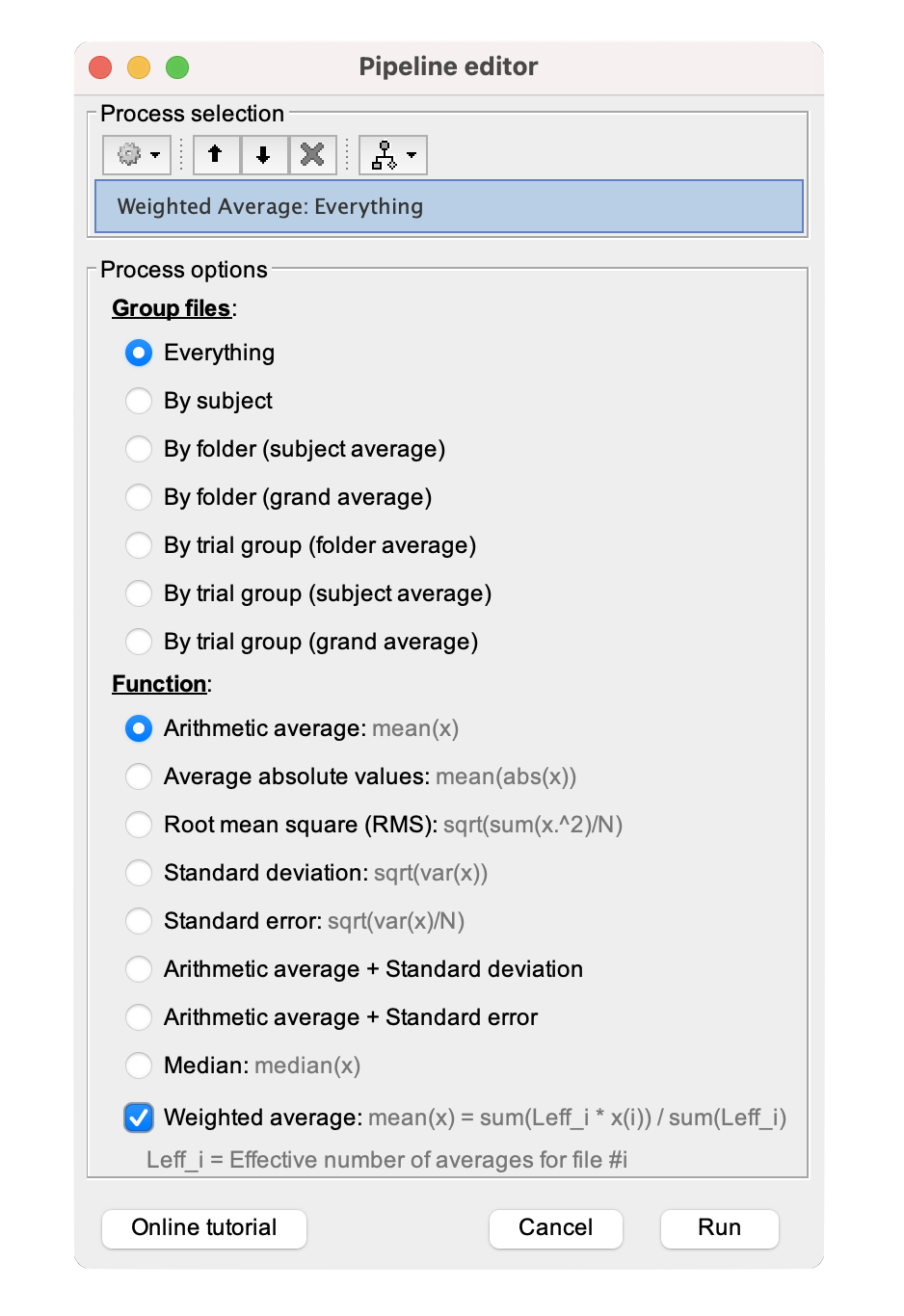
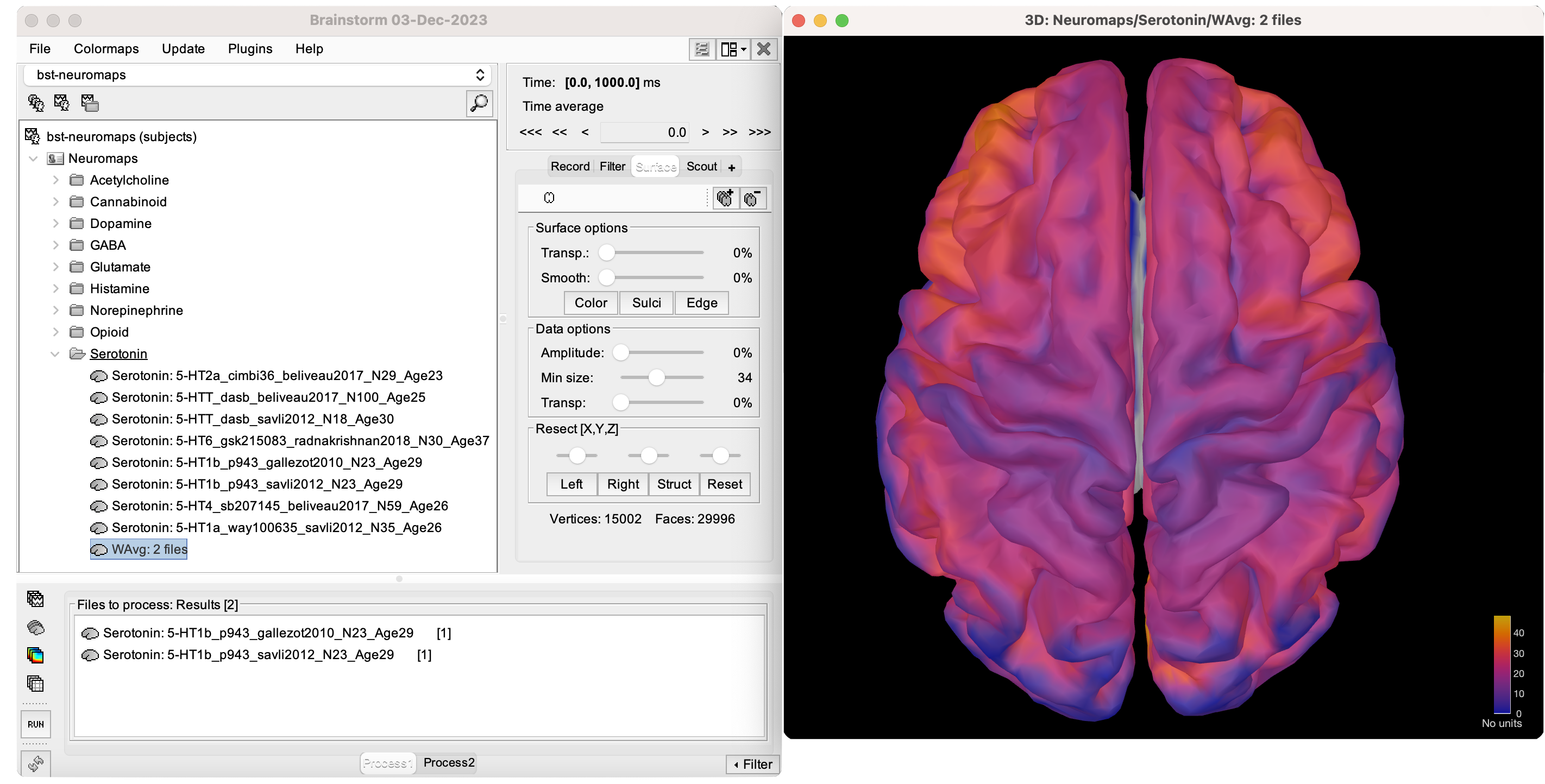
Neurotransmitter annotations that have the same tracer (e.g., Serotonin: 5-HT1b_p943_gallezot2010_N23_Age29 and Serotonin: 5-HT1b_p943_savli2012_N23_Age29 have the same p943 tracer) can be combined to obtain a Click on the Once the Select the process Under 'Group files,' select Check the 'Weighted average' box, as shown below. Click To visualize this new weighted-averaged annotation, follow the same steps as with any brain annotation. Either
By default, the new weighted-averaged file is named 'WAvg:' followed by the number of files averaged. If you wish to rename this or any other files from the Neuromaps plugin to provide more informative labels for your analysis, follow these steps: Alternatively, you can right-click > File > Rename.
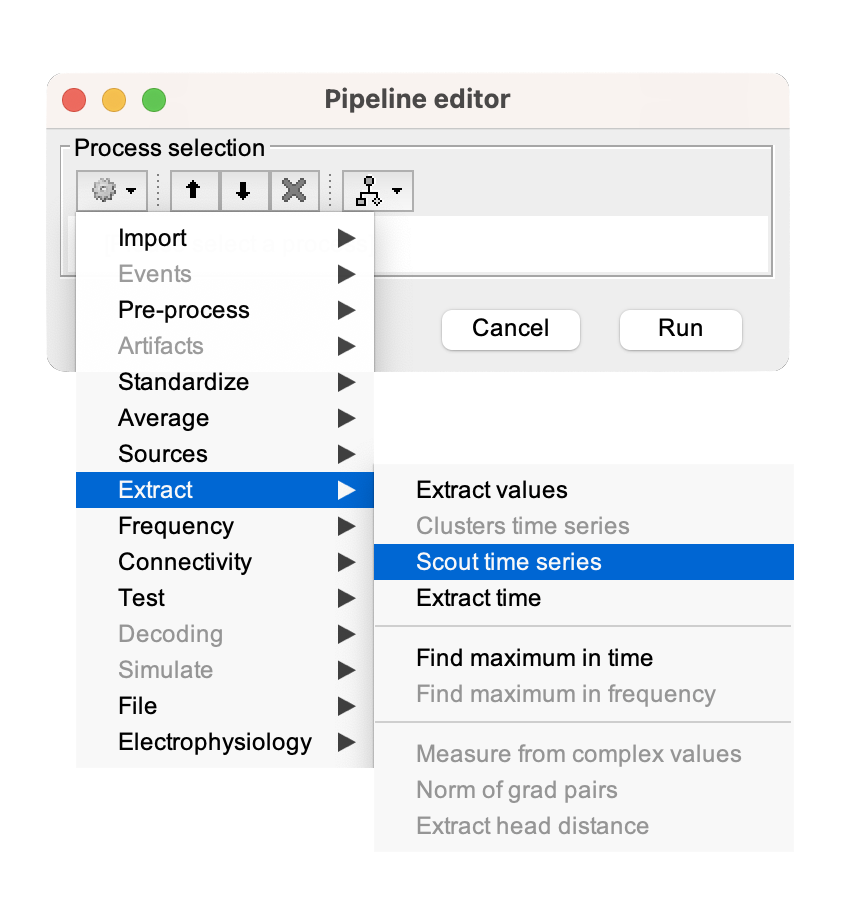
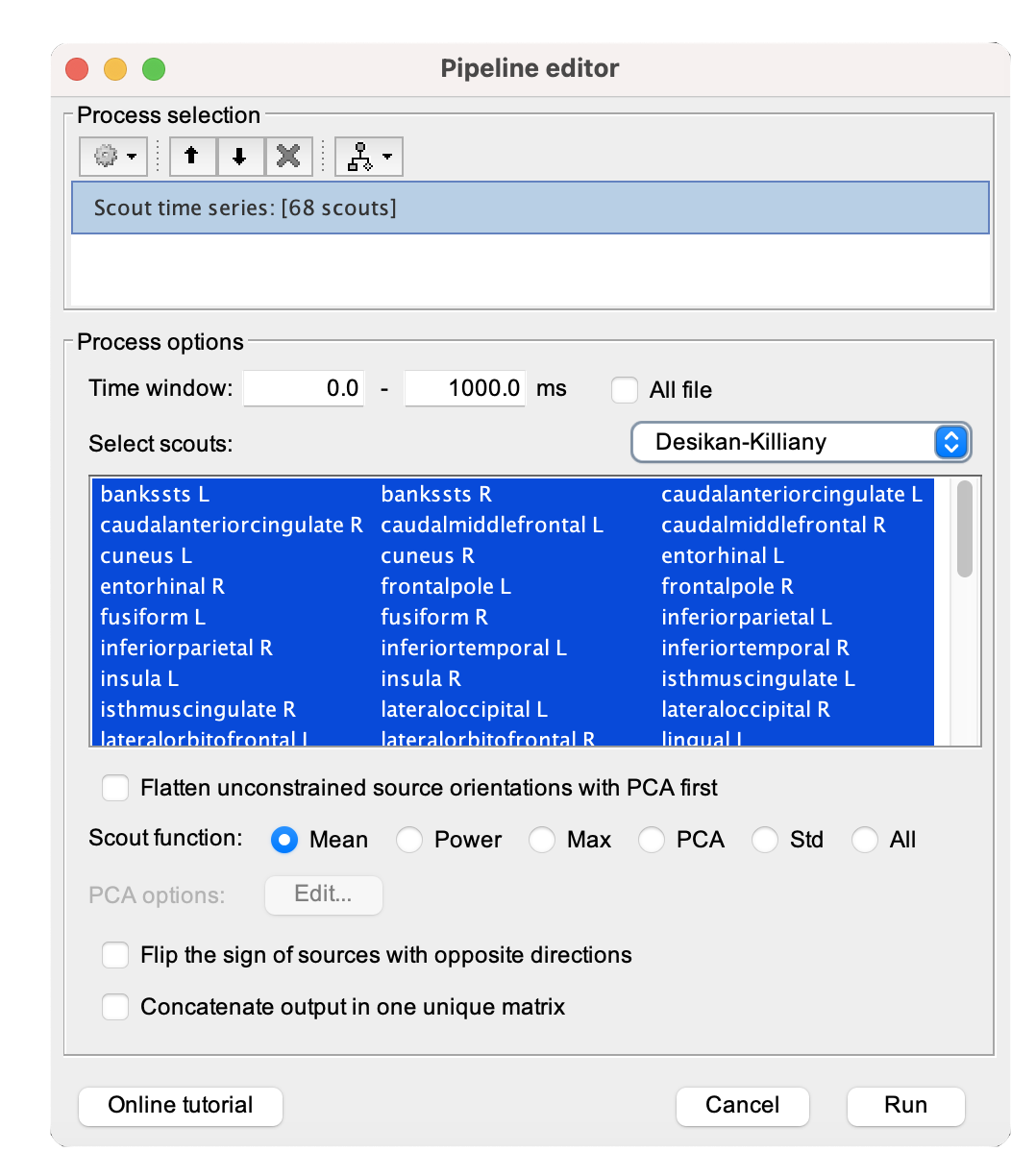
[...] Please refer to Tutorial 23: Scouts for a more in-depth guide in creating, manipulating, and analyzing scouts for effective exploration and comparison of brain activity across different experimental conditions and regions of interest.
One of the main advantages of having these brain annotations in a common coordinate system is that we can statistically compare their spatial topographies. However, most statistical analyses (e.g., Pearson correlation) assume that the values of observations in each sample are independent of one another. Spatial autocorrelation violates this assumption because samples taken from nearby areas are related to each other and are not independent. Spatially-naive null models—both parametric and non-parametric—yield inflated p-values and are inappropriate for significance testing of neuroimaging brain maps. When applied to spatially-autocorrelated brain maps typical of most neuroimaging data, these models approach a false positive rate of >75% and their usage in the field is discouraged (Markello, R. D., & Misic, B., 2021).
According to the literature, null models converge quickly, reaching stable statistical estimates after approximately 100–500 nulls (Markello, R. D., & Misic, B. 2021). Researchers can use this information to balance accuracy and computational feasibility when determining the number of nulls for their analysis.
to be added
The Neuromaps plugin opens up new and exciting avenues for research and can help address questions that depend crucially on anatomical localization. Below, we explore some potential uses of the plugin in advancing integrative research, supported by examples from the literature.
Hansen, J. Y., Shafiei, G., Voigt, K., Liang, E. X., Cox, S. M., Leyton, M., ... & Misic, B. (2023). Integrating multimodal and multiscale connectivity blueprints of the human cerebral cortex in health and disease. Plos Biology, 21(9), e3002314.
da Silva Castanheira, J., Wiesman, A. I., Hansen, J. Y., Misic, B., Baillet, S., Network Quebec Parkinson, & PREVENT-AD Research Group. (2023). Neurophysiological brain-fingerprints of motor and cognitive decline in Parkinson’s disease. medRxiv. Wiesman, A. I., da Silva Castanheira, J., Degroot, C., Fon, E. A., Baillet, S., Network, Q. P., & Prevent-Ad Research Group. (2023). Adverse and compensatory neurophysiological slowing in Parkinson’s disease. Progress in Neurobiology, 231, 102538.
All credit for the conception of the original neuromaps algorithm is due to Ross D. Markello, Justine Y. Hansen, Zhen-Qi Liu, Vincent Bazinet, Golia Shafiei, Laura E. Suárez, Nadia Blostein, Jakob Seidlitz, Sylvain Baillet, Theodore D. Satterthwaite, M. Mallar Chakravarty, Armin Raznahan & Bratislav Misic. The appropriate citation for neuromaps is as follows: Markello, R.D., Hansen, J.Y., Liu, ZQ. et al. neuromaps: structural and functional interpretation of brain maps. Nat Methods 19, 1472–1479 (2022). If you used any of the included maps, please also cite the original papers that publish the data. References for each map can be found in this spreadsheet. Wu, J. et al. Accurate nonlinear mapping between MNI volumetric and FreeSurfer surface coordinate systems. Hum. Brain Mapp. 39, 3793–3808 (2018).
We welcome contributions from the community to help improve and expand the functionality of the Neuromaps plugin. Feel free to submit pull requests on the Github repository, report issues, or provide suggestions below! Your feedback is invaluable in ensuring a user-friendly experience for researchers worldwide. We believe that open science is most impactful when the countless everyone is provided with equal access to the newest and greatest resources in the field.
Introduction
Plugin Key Features
Flexible framework designed to accommodate future expansions and updates. Installing and Running the Neuromaps Plugin
Plugins > Anatomy > neuromaps > Install.
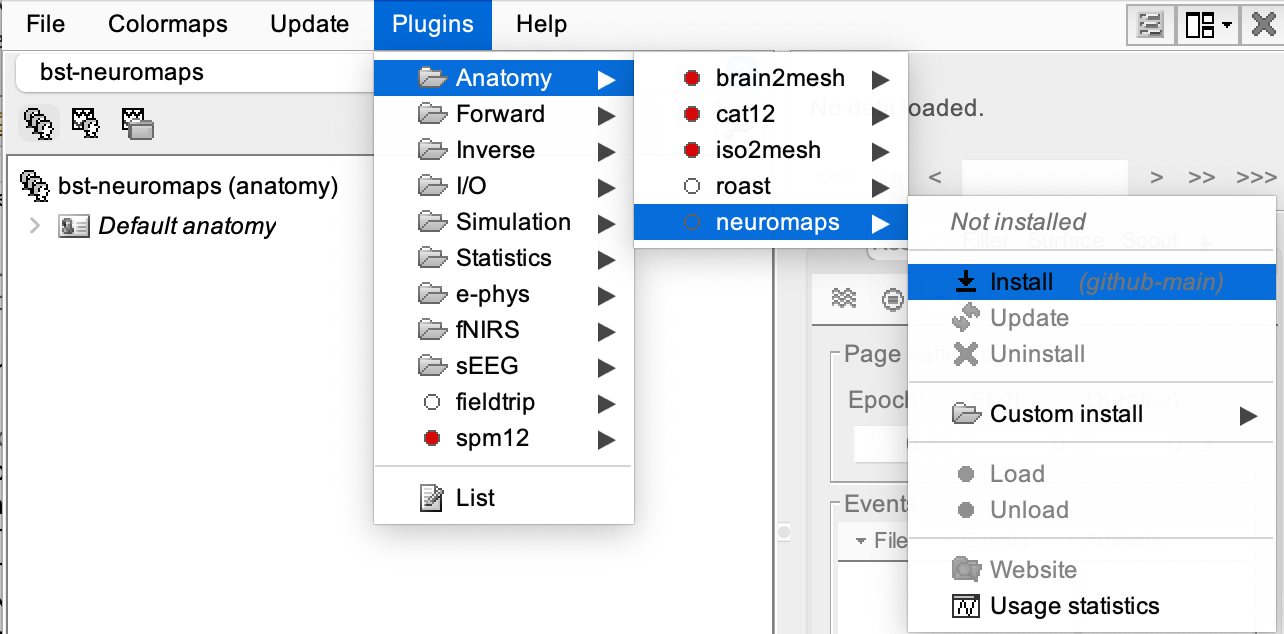
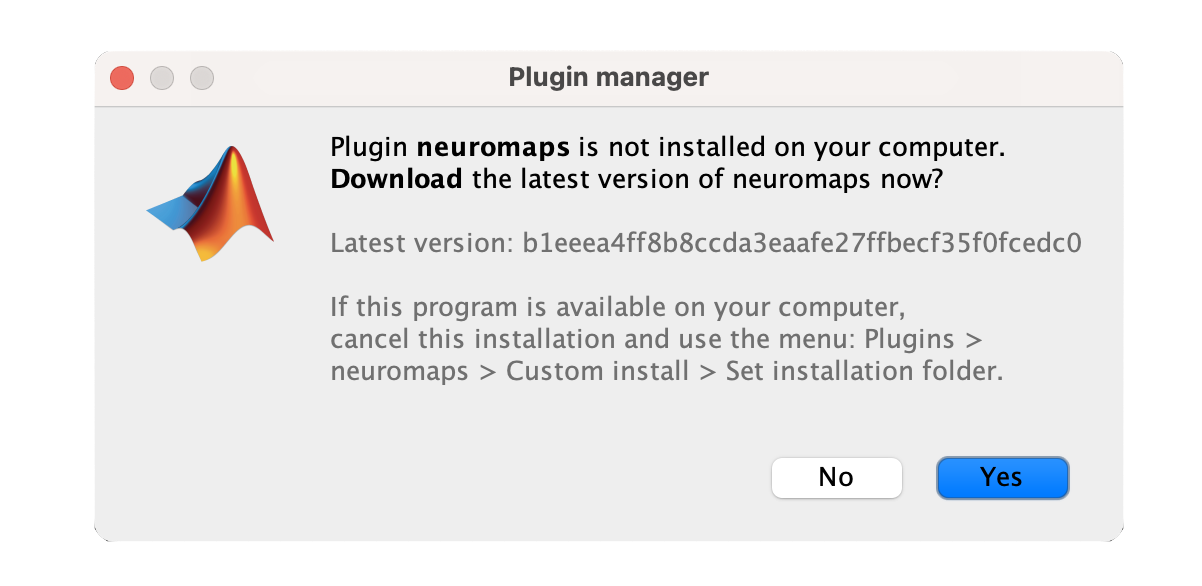
Importing the Brain Annotations
[Run] button at the bottom-left corner of the Process1 tab.
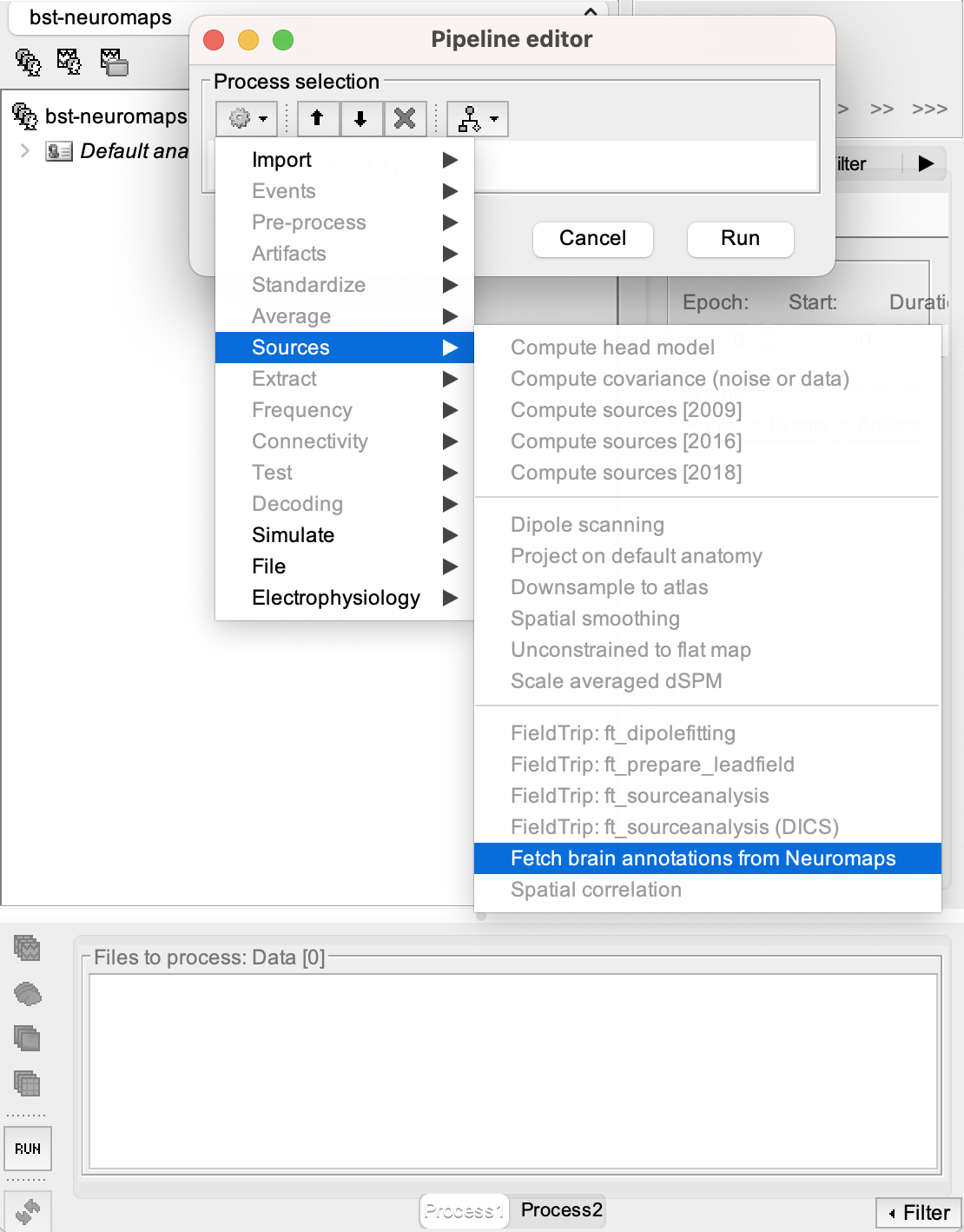
Visualizing and Accessing Annotation Parameters
double-click on the file or right-click > Cortical activations > Display on cortex. Visualization capabilities
Smooth: Inflates the surface to make all the parts of the cortex envelope visible.This is just a display option, it does not actually modify the surface. Colormap
Restore defaults or double-click on the color bar. Creating Weighted-Averaged Annotations
[Run] button at the bottom-left corner of the Process1 tab.
Renaming Files
Scouting values of annotations
Statistical Analyses for Significance Testing
We therefore use spatially informed null models to benchmark the statistical unexpectedness of specific features of interest.
The original neuromaps workflow integrates multiple methods of performing spatial permutations for significance testing. For the plugin, we implemented the most widely adopted of these approaches—the spin method, which involves randomizing the anatomical alignment between two cortical surface maps through spherical rotation by a random angle (Alexander-Bloch et al., 2018). Spatial correlation with brain annotations
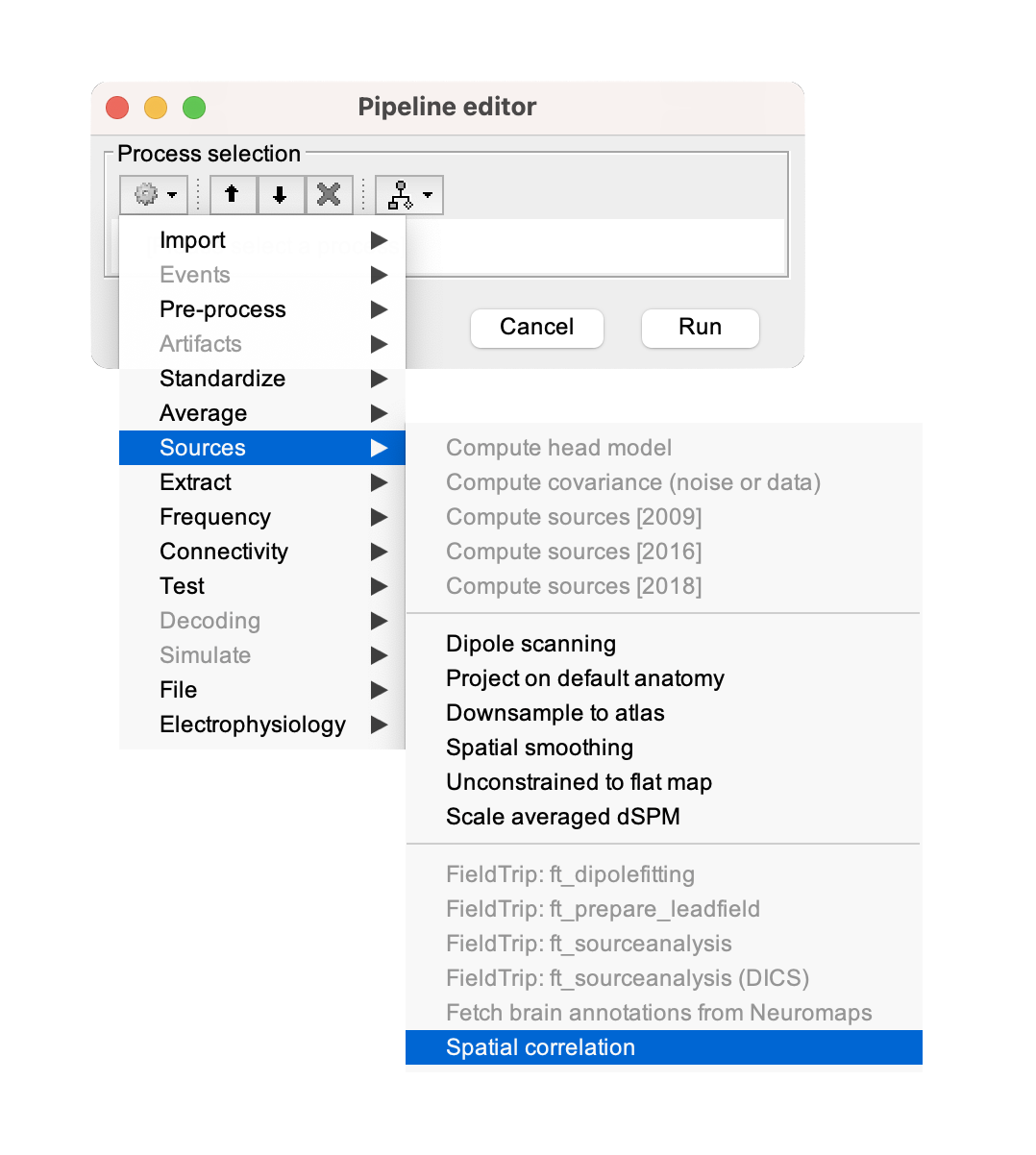
Spatial correlation with any files
Number of spins
Our recommendation is to generate On the Hard Drive
Applications of Neuromaps for Integrative Research
Annotating Structural Connectomes
Identification of Specific Neurochemical Targets for Potential Future Clinical Treatments
Acknowledgements
Contributing
