|
Size: 26660
Comment:
|
Size: 27112
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Resting state MEG recordings = | = MEG resting state and phase-amplitude coupling = |
| Line 4: | Line 4: |
| This tutorial explains how to process continuous resting state MEG recordings. It is based on a eyes open resting recordings of one subject recorded at the Montreal Neurological Institute in 2012 with a CTF MEG 275 system. The segmentation of the T1 MRI of the subject was performed using !FreeSurfer. This tutorial features a few pre-processing steps and the calculation of phase-amplitude coupling measures. More methods will be added soon at the end of this tutorial. | This tutorial features a few pre-processing steps and the calculation of phase-amplitude coupling measures on resting state MEG recordings. It is based on a eyes open resting recordings of one subject recorded at the Montreal Neurological Institute in 2012 with a CTF MEG 275 system. |
| Line 8: | Line 8: |
| <<Include(DatasetResting, , from="\<\<HTML\(\<!-- START-PAGE --\>\)\>\>", to="\<\<HTML\(\<!-- STOP-SHORT --\>\)\>\>")>> |
|
| Line 9: | Line 11: |
| This tutorial dataset (MEG and MRI data) remains a property of the MEG Lab, !McConnell Brain Imaging Center, Montreal Neurological Institute, !McGill University, Canada. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the MEG Lab. | This tutorial dataset (MEG and MRI data) remains a property of the MEG Lab, McConnell Brain Imaging Center, Montreal Neurological Institute, McGill University, Canada. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the MEG Lab. |
| Line 15: | Line 17: |
| * Go to the [[http://neuroimage.usc.edu/brainstorm3_register/download.php|Download]] page of this website, and download the file: '''sample_resting.zip ''' | * Go to the [[http://neuroimage.usc.edu/bst/download.php|Download]] page of this website, and download the file: '''sample_resting.zip ''' |
| Line 18: | Line 20: |
| * Select the menu File > Create new protocol. Name it "'''!TutorialResting'''" and select the options: | * Select the menu File > Create new protocol. Name it "'''TutorialResting'''" and select the options: |
| Line 24: | Line 26: |
| * Right-click on the !TutorialResting folder > New subject > '''Subject02''' | * Right-click on the TutorialResting folder > New subject > '''Subject02''' |
| Line 27: | Line 29: |
| * Set the file format: "!FreeSurfer folder" | * Set the file format: "FreeSurfer folder" |
| Line 49: | Line 51: |
| * Ignore the misplaced MEG helmet you see after the file is link to the database. Those are noise measurements, with no subject in the MEG, so there is no head position in this file. We are not going to use the sensor positions from this file.<<BR>><<BR>> {{attachment:import_noise.gif||height="165",width="211"}} | * Ignore the misplaced MEG helmet you see after the file is link to the database. These are noise measurements, with no subject in the MEG, so there is no head position in this file. We are not going to use the sensor positions from this file.<<BR>><<BR>> {{attachment:import_noise.gif||height="165",width="211"}} |
| Line 56: | Line 58: |
| Some noisy segments have been marked manually. We are going to import those segments now: | Some noisy segments have been marked manually. We are going to import these segments now: |
| Line 61: | Line 63: |
| * Pick the file: '''sample_resting/Data/subj002_spontaneous_20111102_01_AUX.ds/events_BAD.mat''' | * Pick the file: '''Data/subj002_spontaneous_20111102_01_AUX.ds/events_BAD.mat''' |
| Line 71: | Line 73: |
| * '''Detect eye blinks:''' select channel '''EEG058''' (EOG channel), event name "blink".<<BR>><<BR>> {{attachment:blink.gif}} * Review the MISC channels and make sure the events detected make sense.<<BR>><<BR>> {{attachment:ssp_detect_after.gif}} * From the same menu, run the following processes: * '''Remove simulateneous''': Remove events "'''cardiac'''" that are less than '''250ms''' away from a "'''blink'''". * '''Compute SSP: Eyeblinks''': Enter event name "blink", sensor types="MEG". |
* '''Detect eye blinks:''' select channel '''EEG058''' (EOG channel), event name "blink".<<BR>><<BR>> {{attachment:blink.gif||height="209",width="547"}} * Review the MISC channels and make sure the events detected make sense.<<BR>><<BR>> {{attachment:ssp_detect_after.gif||height="257",width="557"}} * From the same SSP menu, run the following processes: * '''Remove simultaneous''': Remove events "'''cardiac'''" that are less than '''250ms''' from a "'''blink'''" |
| Line 78: | Line 79: |
| * The first component of each category should be selected automatically. Leave the default selection as it is but make sure that the corresponding topographies are really representative of blinks and heartbeats.<<BR>><<BR>> {{attachment:ssp_compute.gif||height="146",width="555"}} | * '''Compute SSP: Eyeblinks''': Enter event name "blink", sensor types="MEG".<<BR>>Here you can '''UNCHECK''' the option "Computing using existing SSP", it gives better results. * The first component of each category should be automatically selected. Leave the default selection as it is but make sure that the corresponding topographies are really representative of blinks and heartbeats.<<BR>><<BR>> {{attachment:ssp_compute.gif||height="146",width="555"}} |
| Line 105: | Line 107: |
| * Double-click on the source file and review it briefly with the keyboard shortcut indicated for the [>>] button in the time panel, just to get a sense of what those continuous resting state recordings look like. | * Double-click on the source file and review it briefly with the keyboard shortcut indicated for the [>>] button in the time panel, just to get a sense of what these continuous resting state recordings look like. |
| Line 119: | Line 121: |
| * '''After''': Applies the scout function after the process. First the full PAC comodulograms are calculated for all the individual sources within the scout, then those comodulograms are grouped together using the scout function. Only one resulting averaged PAC map is evaluated for maxPAC and/or saved in the database. | * '''After''': Applies the scout function after the process. First the full PAC comodulograms are calculated for all the individual sources within the scout, then these comodulograms are grouped together using the scout function. Only one resulting averaged PAC map is evaluated for maxPAC and/or saved in the database. |
| Line 126: | Line 128: |
| * By default it shows the PAC map for the first signal only. This file contains one map for each of the 20 sources, because we selected the scout function "All" in the process options. To switch to a different source, use the drop-down menu in the Display tab, or the '''up and down arrows''' of your keyboard. The name of each map is "!ScoutName.!VertexIndex".<<BR>><<BR>> {{attachment:pac_change_vertex.gif||height="172",width="508"}} | * By default it shows the PAC map for the first signal only. This file contains one map for each of the 20 sources, because we selected the scout function "All" in the process options. To switch to a different source, use the drop-down menu in the Display tab, or the '''up and down arrows''' of your keyboard. The name of each map is "ScoutName.VertexIndex".<<BR>><<BR>> {{attachment:pac_change_vertex.gif||height="172",width="508"}} |
| Line 133: | Line 135: |
| * As a general observation we notice that many sources seem to show a strong coupling around 10Hz/90Hz, others seem to show coupling around 12Hz/110Hz. Both could be relative to relation between the alpha rhythms and the high-frequency activity in the primary visual areas of the cortex (V1). However, it is difficult from this first computation to estimate whether those two observations are relative to the same brain process, or due to two different mechanisms. | * As a general observation we notice that many sources seem to show a strong coupling around 10Hz/90Hz, others seem to show coupling around 12Hz/110Hz. Both could be relative to relation between the alpha rhythms and the high-frequency activity in the primary visual areas of the cortex (V1). However, it is difficult from this first computation to estimate whether these two observations are relative to the same brain process, or due to two different mechanisms. |
| Line 141: | Line 143: |
| * Open all those files, and observe that there is now only one comodulogram saved per file, for the RO1 scout. The first one (before) shows only the most robust effects that are still present in the signal after the averaging of all the sources. The two others (after) show a combination of all the coupling effects that were identified individually for each source, the resulting maps are a lot noisier, with more intermediate values. | * Open all these files, and observe that there is now only one comodulogram saved per file, for the RO1 scout. The first one (before) shows only the most robust effects that are still present in the signal after the averaging of all the sources. The two others (after) show a combination of all the coupling effects that were identified individually for each source, the resulting maps are a lot noisier, with more intermediate values. |
| Line 158: | Line 160: |
| * A small time window is extracted around each of those marked troughs. * A time-frequency decomposition of those short epochs is performed using a collection of narrow band-pass filters. |
* A small time window is extracted around each of these marked troughs. * A time-frequency decomposition of these short epochs is performed using a collection of narrow band-pass filters. |
| Line 166: | Line 168: |
| * '''Time window''': Create those maps for the segment [400s-600s] (same as before). | * '''Time window''': Create these maps for the segment [400s-600s] (same as before). |
| Line 196: | Line 198: |
| * In the previous example, we have identified visually the optimal low frequency on the PAC comodulograms, and we have entered this value manually in the options of the "Canolty map" process. This methodology is probably the best way to proceed when exploring manually some specific brain regions, but it will not be adapted to process automatically a large number of signals. In the case of a full brain analysis, we cannot reproduce the same steps individually for the 15000 sources. | * In the previous example, we have identified visually the optimal low frequency on the PAC comodulograms, and we have entered this value manually in the options of the "Canolty map" process. This methodology is probably the best way to proceed when exploring manually some specific brain regions, but it will not be adapted to process a large number of signals. In the case of a full brain analysis, we cannot reproduce the same steps individually for the 15000 sources. |
| Line 199: | Line 201: |
| * At the current moment, the problem with this approach is that the metric used for estimating the maxPAC is not very stable. It just takes the frequency pair that gives the maximum PAC value, and as we saw earlier in this tutorial, it is most of the time not matching the frequency we are really interested in. This automated version of the low-frequency detection is not recommended for now, but we are currently working on implementing more stable metrics. We will post the updates on those developments on this tutorial page. | * At the current moment, the problem with this approach is that the metric used for estimating the maxPAC is not very stable. It just takes the frequency pair that gives the maximum PAC value, and as we saw earlier in this tutorial, it is most of the time not matching the frequency we are really interested in. This automated version of the low-frequency detection is not recommended for now, but we are currently working on implementing more stable metrics. We will post the updates on these developments on this tutorial page. |
| Line 202: | Line 204: |
| The operations described in this tutorial can be reproduced from a Matlab script, available in the Brainstorm distribution: '''brainstorm3/toolbox/script/tutorial_resting.m ''' == Feedback == <<EmbedContent(http://neuroimage.usc.edu/brainstorm3_register/get_feedback.php?Tutorials/Resting)>> |
The following script from the Brainstorm distribution reproduces the analysis presented in this tutorial page: '''brainstorm3/toolbox/script/tutorial_''''''resting''''''.m''' <<HTML(<div style="border:1px solid black; background-color:#EEEEFF; width:720px; height:500px; overflow:scroll; padding:10px; font-family: Consolas,Menlo,Monaco,Lucida Console,Liberation Mono,DejaVu Sans Mono,Bitstream Vera Sans Mono,Courier New,monospace,sans-serif; font-size: 13px; white-space: pre;">)>><<EmbedContent("http://neuroimage.usc.edu/bst/viewcode.php?f=tutorial_resting.m")>><<HTML(</div >)>> <<EmbedContent(http://neuroimage.usc.edu/bst/get_feedback.php?Tutorials/Resting)>> |
MEG resting state and phase-amplitude coupling
Authors: Thomas Donoghue, Soheila Samiee, Elizabeth Bock, Esther Florin, Francois Tadel, Sylvain Baillet
This tutorial features a few pre-processing steps and the calculation of phase-amplitude coupling measures on resting state MEG recordings. It is based on a eyes open resting recordings of one subject recorded at the Montreal Neurological Institute in 2012 with a CTF MEG 275 system.
Contents
License
This tutorial dataset (MEG and MRI data) remains a property of the MEG Lab, McConnell Brain Imaging Center, Montreal Neurological Institute, McGill University, Canada. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the MEG Lab.
If you reference this dataset in your publications, please acknowledge its authors (Elizabeth Bock, Esther Florin, Francois Tadel and Sylvain Baillet) and cite Brainstorm as indicated on the website. For questions, please contact us through the forum.
Presentation of the experiment
Experiment
- One subject
- Two runs of 10 min of resting state recordings
- Eyes open
MEG acquisition
Acquisition at 2400Hz, with a CTF 275 system, subject in seating position
- Recorded at the Montreal Neurological Institute in 2012
- Anti-aliasing low-pass filter at 600Hz, files saved with the 3rd order gradient
- Recorded channels (301):
- 26 MEG reference sensors (#2-#27)
- 272 MEG axial gradiometers (#28-#299)
- 1 ECG bipolar (#300)
- 2 vertical EOG bipolar (#301)
- 1 Unused channels (#1)
- 3 datasets:
subj002_spontaneous_20111102_01_AUX.ds: Run #1, 10min resting state
subj002_spontaneous_20111102_02_AUX.ds: Run #2, 10min resting state
subj002_noise_20111104_02.ds: Empty room recordings, 100s
Head shape and fiducial points
3D digitization using a Polhemus Fastrak device (subj002_11022011.pos)
Position of the anatomical references used for these recordings:
The position of the CTF head localization coils, pictures available in file fiducials.jpg- Around 100 head points distributed on the hard parts of the head (no soft tissues)
Subject anatomy
- Subject with 1.5T MRI
Processed with FreeSurfer 5.2
License
This tutorial dataset (MEG and MRI data) remains a property of the MEG Lab, McConnell Brain Imaging Center, Montreal Neurological Institute, McGill University, Canada. Its use and transfer outside the Brainstorm tutorial, e.g. for research purposes, is prohibited without written consent from the MEG Lab.
If you reference this dataset in your publications, please acknowledge its authors (Elizabeth Bock, Esther Florin, Francois Tadel and Sylvain Baillet) and cite Brainstorm as indicated on the website. For questions, please contact us through the forum.
Download and installation
Requirements: You have already followed all the basic tutorials and you have a working copy of Brainstorm installed on your computer.
Go to the Download page of this website, and download the file: sample_resting.zip
Unzip it in a folder that is not in any of the Brainstorm folders (program or database folder).
- Start Brainstorm (Matlab scripts or stand-alone version).
Select the menu File > Create new protocol. Name it "TutorialResting" and select the options:
"No, use individual anatomy",
"No, use one channel file per condition".
Import the anatomy
- Switch to the "anatomy" view.
Right-click on the TutorialResting folder > New subject > Subject02
- Leave the default options you set for the protocol.
Right-click on the subject node > Import anatomy folder:
Set the file format: "FreeSurfer folder"
Select the folder: sample_resting/anatomy
- Number of vertices of the cortex surface: 15000 (default value)
- Set the 6 required fiducial points (indicated in MRI coordinates):
- NAS: x=128, y=225, z=135
- LPA: x=54, y=115, z=107
- RPA: x=204, y=115, z=99
- AC: x=133, y=137, z=152
- PC: x=132, y=108, z=150
- IH: x=133, y=163, z=196 (anywhere on the midsagittal plane)
At the end of the process, make sure that the file "cortex_15000V" is selected (downsampled pial surface, that will be used for the source estimation). If it is not, double-click on it to select it as the default cortex surface.
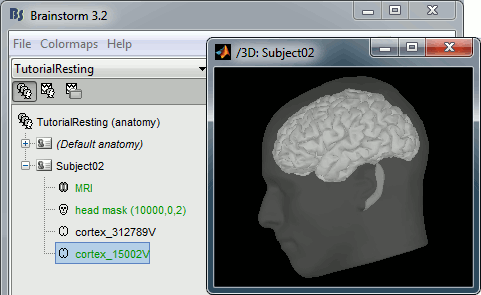
Access the recordings
Resting state recordings: 10min
- The sample_resting download contains two 10 minute resting state runs. We are going to use the first one (run 01).
- Switch to the "functional data" view, the middle button in the toolbar above the database explorer.
Right click on the Subject02 > Review raw file > Pick the file:
sample_resting/Data/subj002_spontaneous_20111102_01_AUX.dsRefine registration now? YES
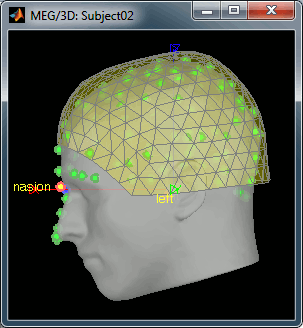
Empty room recordings: 90s
- We are also going to link to the database 20 seconds of empty room measurements that were acquired before the subject entered the MEG room. During the source estimation process, we will use this file to estimate the noise related with the MEG sensors.
Right click on the Subject02 > Review raw file > Pick the file:
sample_resting/Data/subj002_noise_20111104_02.dsIgnore the misplaced MEG helmet you see after the file is link to the database. These are noise measurements, with no subject in the MEG, so there is no head position in this file. We are not going to use the sensor positions from this file.
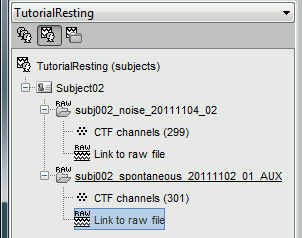
Important note: From now on, all the pre-processing steps that re-write new continuous files (bandpass filter, notch filter) applied to the resting state MEG recordings should be applied as well to the noise recordings. We will calculate the noise covariance from the noise recordings and then use them to whiten the resting state recordings to estimate the sources. For this operation to be valid, the two datasets must be processed exactly in the same way. The SSP are not a problem as they are applied dynamically to the noise covariance when estimating the sources.
Pre-processing
All data should be pre-processed and checked for artifacts prior to doing analyses such as PAC (including marking bad segments and correcting for artifacts). For the purposes of this tutorial, we will correct for blinks and heartbeats with SSPs but will not go through marking out bad sections. When using your own data reviewing the raw data for bad segments and using clean data is of the utmost importance.
Bad segments
Some noisy segments have been marked manually. We are going to import these segments now:
- Double-click on the "Link to raw file" to display the resting state MEG recordings.
In the Record tab, select the menu File > Add events from file
Select the file format: Brainstorm (events*.mat)
Pick the file: Data/subj002_spontaneous_20111102_01_AUX.ds/events_BAD.mat
- This will load 26 bad segments, which will be ignored from most of the upcoming computations.
Heartbeats and eye blinks
Signal Space Projection (SSP) is a method for projecting the recordings away from stereotyped artifacts, such as eye blinks and heartbeats.
- Double-click on the "Link to raw file" to display the resting state MEG recordings.
- From the SSP menu in the Record tab, run the automatic detection of the blinks and the heartbeats:
Detect heartbeats: select channel EEG057 (ECG channel), event name "cardiac".
Detect eye blinks: select channel EEG058 (EOG channel), event name "blink".
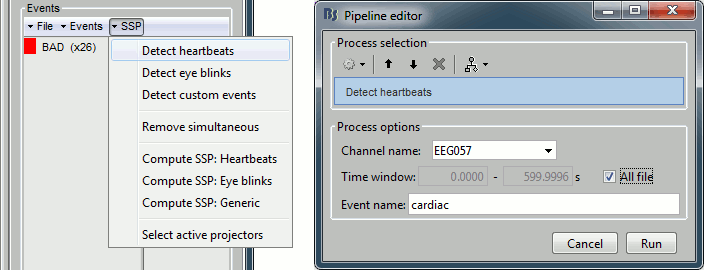
Review the MISC channels and make sure the events detected make sense.
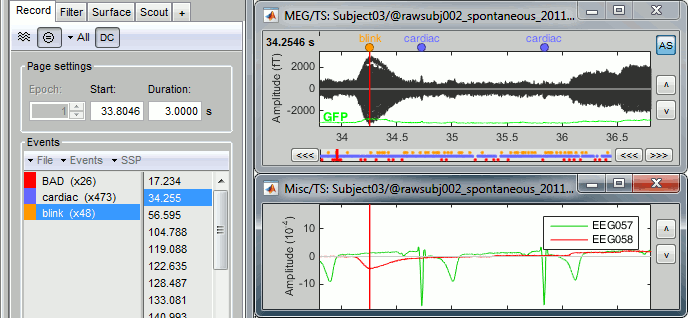
- From the same SSP menu, run the following processes:
Remove simultaneous: Remove events "cardiac" that are less than 250ms from a "blink"
Compute SSP: Heartbeats: Enter event name "cardiac", sensor types="MEG".
Compute SSP: Eyeblinks: Enter event name "blink", sensor types="MEG".
Here you can UNCHECK the option "Computing using existing SSP", it gives better results.
The first component of each category should be automatically selected. Leave the default selection as it is but make sure that the corresponding topographies are really representative of blinks and heartbeats.
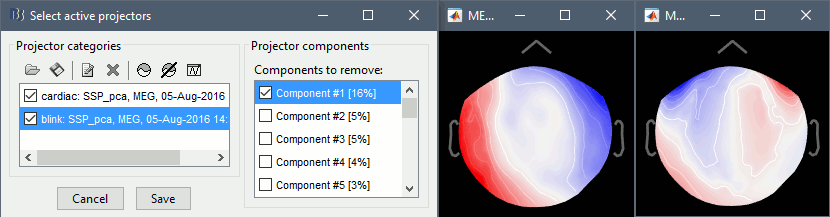
For more information regarding this SSP method, refer to this tutorial: ?Detect and remove artifacts.
Power line contamination
- PAC analysis involves examining a wide band of frequencies, often the entire range of 2-150Hz or more. This band contains the frequencies contaminated by line noise, of either 50 or 60Hz and their harmonics. Brainstorm offers tools to remove line noise from functional data and it is most of the time a recommended pre-processing step. However, here we will not run the notch filter for time efficiency and because it is not required for accurate PAC analysis.
- The PAC function looks for high frequencies occurring specifically at certain phases of low signals such that the ubiquitous nature of line contamination effectively cancels it out for being identified as PAC. Similarly, running the notch filter results in no 60Hz anywhere, such that the function also identifies no PAC.
Source estimation
We need now to calculate a source model for the resting state recordings, using a noise covariance matrix calculated from the noise recordings.
Head model
Right-click on the recordings node ("spontaneous") > Compute head model.
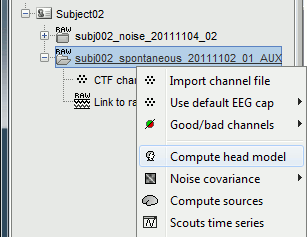
Use the overlapping spheres model and keep all of the options at their default values.
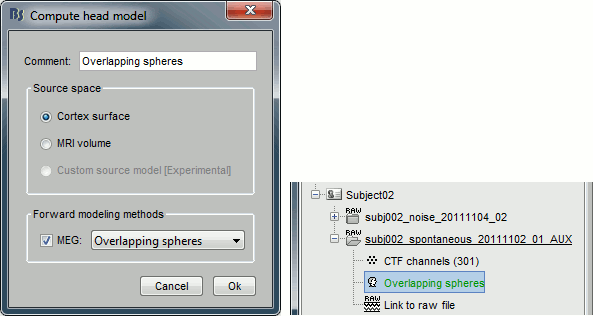
For more information: ?Head model tutorial.
Noise covariance
Right-click on the noise recordings > Noise covariance > Compute from recordings.
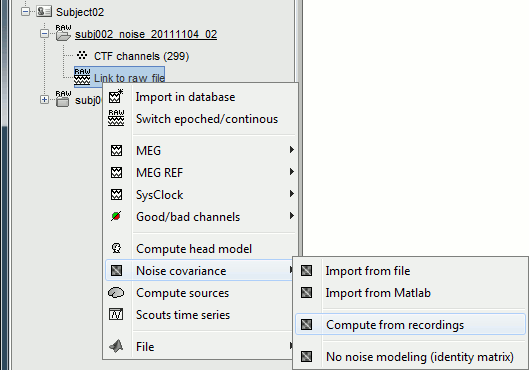
- Leave all the default options and click [OK].
Right-click on the noise covariance file > Copy to other conditions.
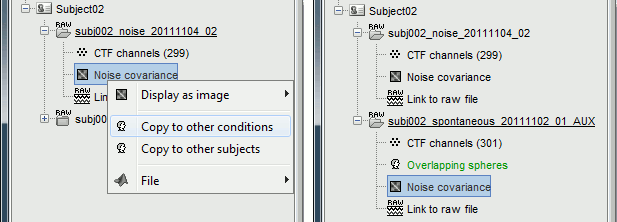
For more information: ?Noise covariance tutorial.
Inverse model
Right-click on the "Link to raw file" > Compute sources.
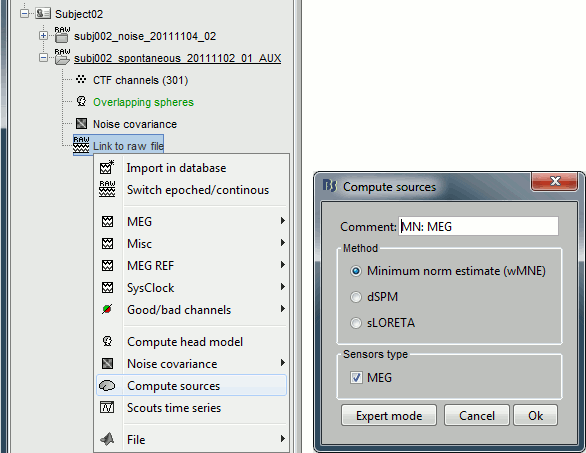
The inverse kernel is saved in a new file in the database.
For more information: ?Source estimation tutorial.
Scouts
Double-click on the source file and review it briefly with the keyboard shortcut indicated for the [>>] button in the time panel, just to get a sense of what these continuous resting state recordings look like.
Create a scout "RO1" with 20 vertices at the pole of the right occipital lobe.
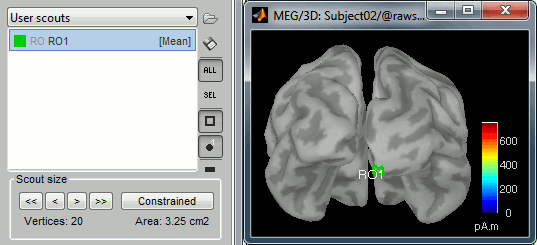
Phase-amplitude coupling
We are now ready to run the PAC analysis on the source signals. This PAC function in Brainstorm is not time resolved, but will analyze the given time series for any stable occurrence of PAC over a time segment you give it. For more information about the PAC measure used here, please refer to the online tutorial
Phase-amplitude coupling.
PAC estimation
- Drag and drop the sources for the spontaneous recordings to the Process1 tab.
Select the process "Frequency > Phase-amplitude coupling"
Set the options as follows (400s-600s, all the sources of scout RO1):
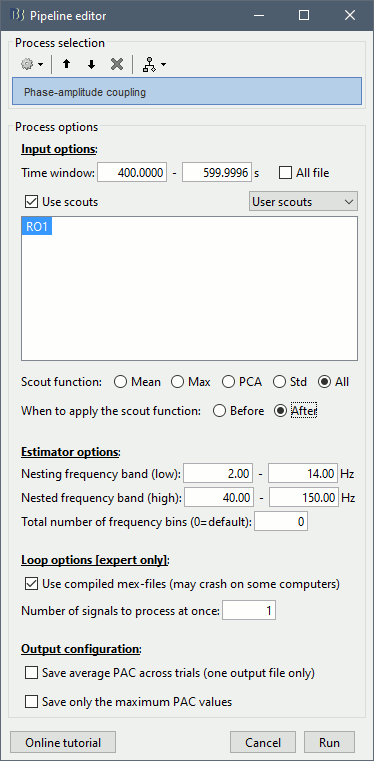
All the options are described in the PAC tutorial, except for the scout options:
- The input options allow you to select whether to process the full brain (15000 sources) or just a few scouts. Processing the full source maps might require a tremendous amount of memory and days of computation, so we recommend you run your analysis on a few small regions of interest. Select the "Use scouts" option, and select the scouts to process.
Scout function: Operation to apply to group the multiple vertices into one, or "All" to keep all the sources.
Before: Applies the scout function before the process, so that only the resulting scout time series is evaluated for PAC coupling. This is usually not a recommended option for more than a few sources, as the fine temporal dynamics are sometimes observable only on raw source signals and become less clear in averaged signals.
After: Applies the scout function after the process. First the full PAC comodulograms are calculated for all the individual sources within the scout, then these comodulograms are grouped together using the scout function. Only one resulting averaged PAC map is evaluated for maxPAC and/or saved in the database.
- In the case of the scout function "All", the before/after option is not relevant.
Note that you need at least 5Gb of memory to run this process. If the process is taking more than two minutes or if your computer becomes very slow or unresponsive, you might be going over your memory limits. At this point, you should monitor the memory usage on your computer using the task manager or the "ps" command from a terminal (google for help). If you notice that the total memory usage is reaching 99%, you should stop the computation and start over with a smaller time window (400s-500s) or a scout with fewer vertices (10).
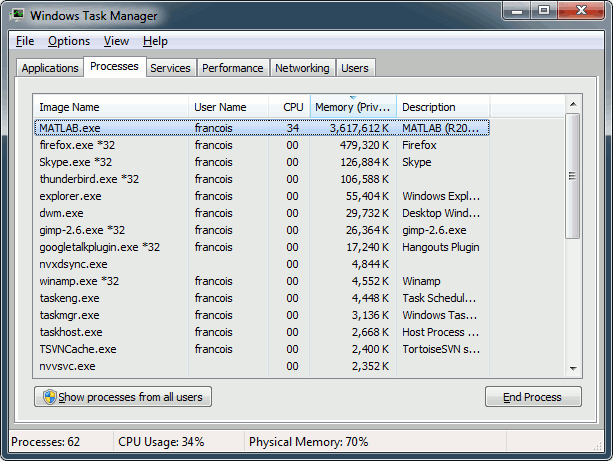
To stop a running Brainstorm process from Matlab: click on the Matlab command window and press CTRL+C. This will terminate the current script execution once the current process is over. If it doesn't stop after a short while, you may have to kill the Matlab process itself (using your task manager or the "kill" command from a terminal).
Visual exploration of the comodulogram
Double-click on the PAC file to display the comodulograms (X=freq for phase, Y=freq for amplitude).
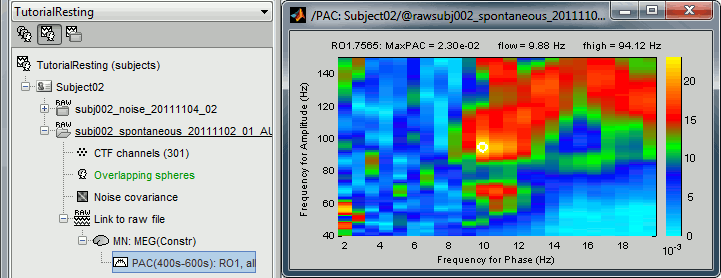
By default it shows the PAC map for the first signal only. This file contains one map for each of the 20 sources, because we selected the scout function "All" in the process options. To switch to a different source, use the drop-down menu in the Display tab, or the up and down arrows of your keyboard. The name of each map is "ScoutName.VertexIndex".
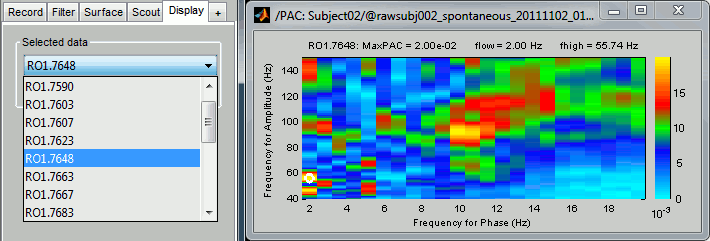
The spots with the yellow and red colors indicate the frequency pairs that show a higher coupling. Some sources seem to show some strong coupling patterns, some other don't. Isolated values are not necessarily meaningful, we are more looking for extended colored blobs that are observable across multiple frequency bins. Manipulating the colormap can help you identifying the coupling values of interest. Click on the colorbar and move the mouse left/right to adjust the contrast and up/down to adjust the brightness. Alternatively, right-click on the figure > Colormap: PAC > Contrast/Brightness. Double-click on the colorbar to reset its default configuration.
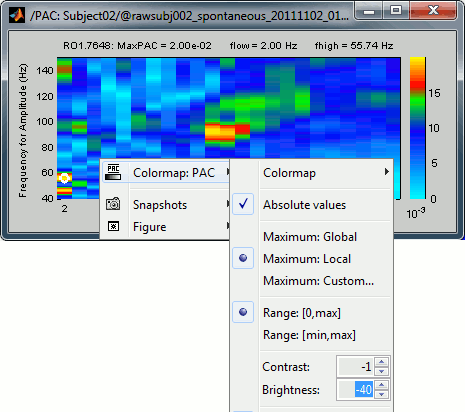
- A small white circle indicates the PAC pairing with the strongest coupling that was identified for the file (maxPAC). The values for this pairing are displayed on the top of the comodulogram:
MaxPAC: the strength of the coupling
flow: the low frequency (nesting)
fhigh: the high frequency (nested)
The maximum is not a very stable estimator and the identified maximal coupling does not necessarily represent the effect we are interested in. In some cases, we would not want to use the default maxPAC but some other low-frequency value that we identify visually. You can click on the figure to see the coupling frequencies and measure at other points: it shows a red circle and the PAC information at the bottom of the figure. This allows you to explore the values represented in the comodulogram.
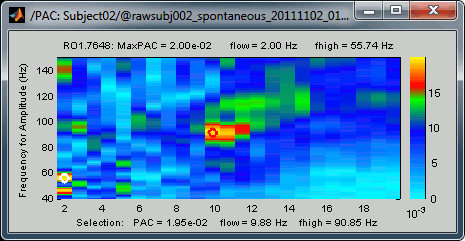
- As a general observation we notice that many sources seem to show a strong coupling around 10Hz/90Hz, others seem to show coupling around 12Hz/110Hz. Both could be relative to relation between the alpha rhythms and the high-frequency activity in the primary visual areas of the cortex (V1). However, it is difficult from this first computation to estimate whether these two observations are relative to the same brain process, or due to two different mechanisms.
Scout functions
- You can run again the same process but selecting a different way to process the region of interest. In the previous example, we have saved all the sources independently. Now, run again the same process but using the following scouts configurations:
Mean / Before: Calculate the PAC map for the average scout time series
Mean / After: Calculate the average of the PAC maps obtained for each source
Max / After: Extract the maximum value across the PAC maps calculated for each source
Note that to open again the pipeline editor with the same options on the same input files, you can use the menu File > Reload last pipeline. This is a faster way to run again the same process twice but changing just one parameter, this way you don't have to enter the time window again.
- Open all these files, and observe that there is now only one comodulogram saved per file, for the RO1 scout. The first one (before) shows only the most robust effects that are still present in the signal after the averaging of all the sources. The two others (after) show a combination of all the coupling effects that were identified individually for each source, the resulting maps are a lot noisier, with more intermediate values.
Comodulograms for RO1 scouts with the default colormap configuration:
1) mean/before, 2) mean/after, 3) max/after
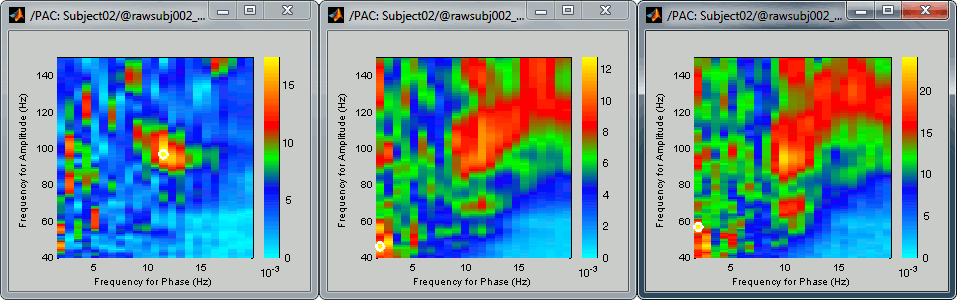
After changing the brightness/contrast of the colormap:
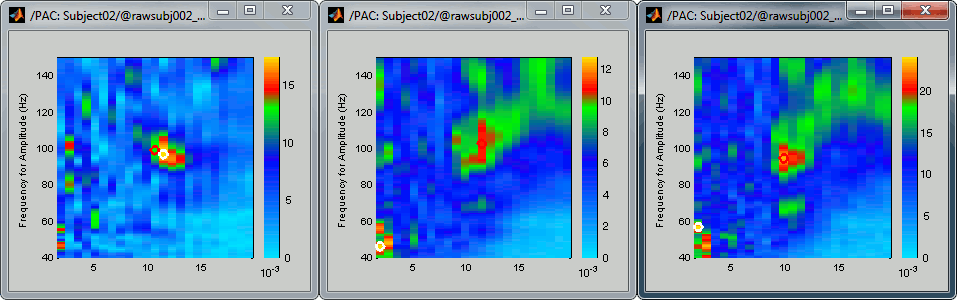
After setting the same maximum for all the three views:
- All three maps seem to show stronger coupling between the range [9-13]Hz and [90-110]Hz. Averaging the values for the region of interest groups into one mode the two separate coupling ranges we were observing with the maps of the individual sources in the previous section (10/90Hz and 12/110Hz). We lose some resolution but probably extract more reproducible results. The purpose of this tutorial is not to discuss whether one result is more correct than the other, but rather to illustrate the tools available in Brainstorm to estimate the phase-amplitude coupling within brain signals.
- Double-click on the colorbar to reset it to its defaults.
Canolty maps
Introduction to the method
Canolty maps are a type of time-frequency decomposition that offer another way to visualize the data and serve as a complimentary tool to visualize and assess phase-amplitude coupling. The process lines up the data to a specific low frequency so as to visualize what happens in the power spectrum related to the phase of the low frequency. Currently there are no significance tests within Brainstorm that can give a measure if PAC is significant in a given time series, but the Canolty maps provide an important way to verify and corroborate the results of the PAC process. We name these maps after the author of the paper in which they were first introduced:
Canolty RT, Edwards E, Dalal SS, Soltani M, Nagarajan SS, Kirsch HE, Berger MS, Barbaro NM, Knight RT
High gamma power is phase-locked to theta oscillations in human neocortex
Science, 2006 Sep 15;313(5793):1626-8.
The procedure to obtain a Canolty map for a signal is the following:
- The signal of interest is filtered at the low frequency of interest using a narrow band-pass filter.
- The amplitude troughs of the low frequency are detected from the filtered signal.
- A small time window is extracted around each of these marked troughs.
- A time-frequency decomposition of these short epochs is performed using a collection of narrow band-pass filters.
- The power of all the TF maps is averaged to constitute the final Canolty map.
Computation
- Select the sources for the spontaneous recordings in the Process1 tab (same as previously).
Select the process "Frequency > Canolty maps", set the options following:
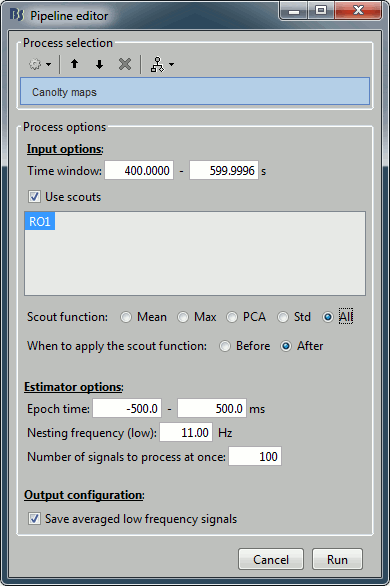
- Explanation of the options:
Time window: Create these maps for the segment [400s-600s] (same as before).
Scouts: Keep all the sources separately (before/after option is not relevant).
Epoch time: How much time we take around each trough of the low frequency amplitude.
Nesting frequency: Let's use 11Hz, which is in the middle of the band we identified in the previous section (9-13Hz).
Save average low frequency signal: Saves the average of the extracted epochs, obtained before the time-frequency decomposition. It will show the event-related signals, locked on the troughs of the low frequency.
- If you get out of memory errors, or if it takes too long to calculate, you can decrease the number of vertices in the scout.
Double-click on the file "Canolty maps": standard TF plot, for help read the ?time-frequency tutorial.
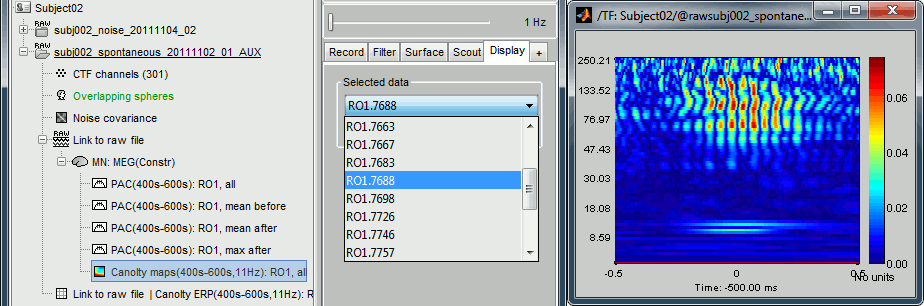
This representation shows whether the power of any high frequencies fluctuates systematically with the phase of the low frequency: basically, we can visualize PAC. If there is PAC present, we see quite stereotyped stripes of the power of certain high frequencies changing consistently with the phase of the low frequency. If there is no PAC there is no discernible pattern. The image above shows some strong coupling between the alpha oscillations (11Hz nesting frequency that we selected in the process options) and the gamma band (60-150Hz, with a peak around 95Hz).
Double-click on the file "Link to raw file | Canolty ERP": it contains the average of the source signals epoched at the trough of the nesting frequency. In the Canolty map itself there is no representation of the low frequency used, it can be useful to visualize the low frequency simultaneously. In this figure, one black line represents one source, you can click on one line to select it then right-click on it to see its name or index.
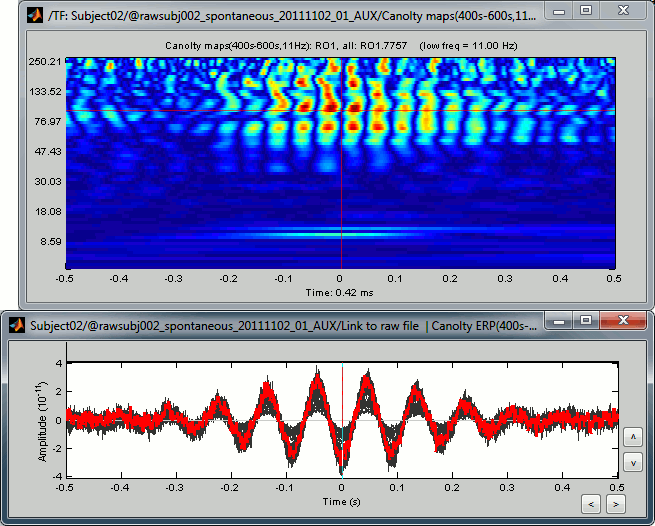
- By arranging the Canolty map and low frequency time series you can get a sense of how the low frequency oscillation and high frequency amplitude relate to each. The time selection on the two is synchronized, so try clicking at particular parts of the low frequency file and examining the power in the Canolty map. You should notice gamma amplitude is low at the peak and trough of the low frequency oscillation and high between them.
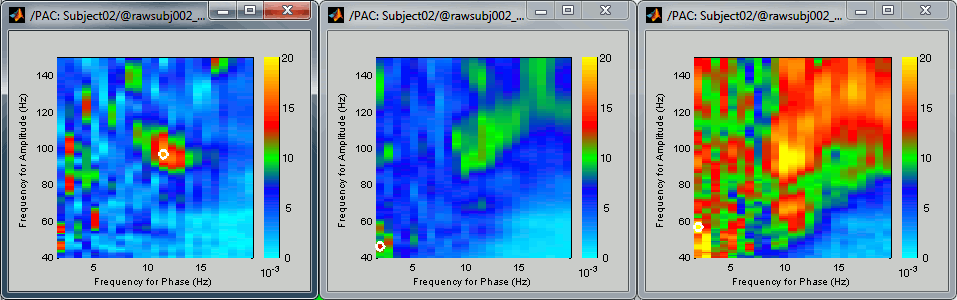
If you cycle through a few sources, you can see that some show a strong coupling and some others don't. If you open at the same time the PAC comodulogram, you can observe that the sources for which we had higher PAC values are also the ones for which with see a better evidence of the alpha-gamma coupling here. Here are a few examples, two good ones and a bad one:
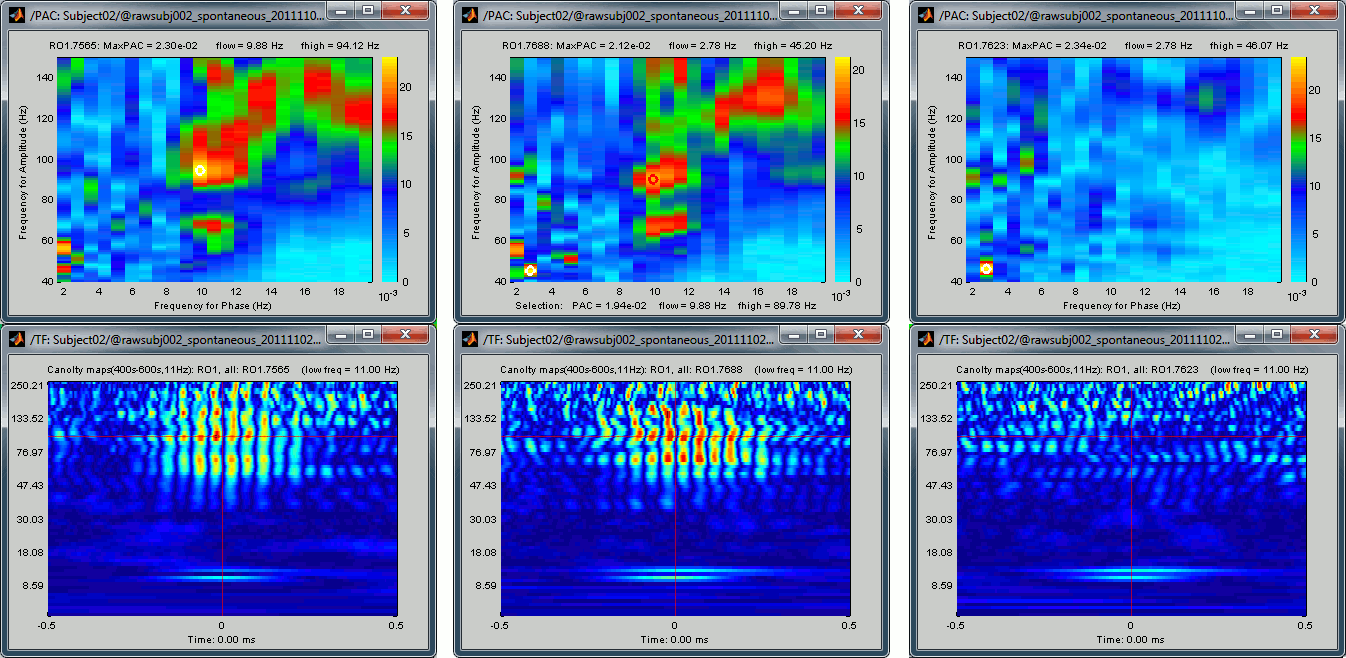
Scout functions
- Similarly to what we've done with the PAC estimation, we can run again the same process but selecting a function to unify the region of interest:
Mean / Before: Calculate the Canolty map for the average scout time series
Mean / After: Calculate the average of the Canolty maps obtained for each source
Max / After: Extract the maximum value across the Canolty maps obtained for each source
Open the three files side by side:
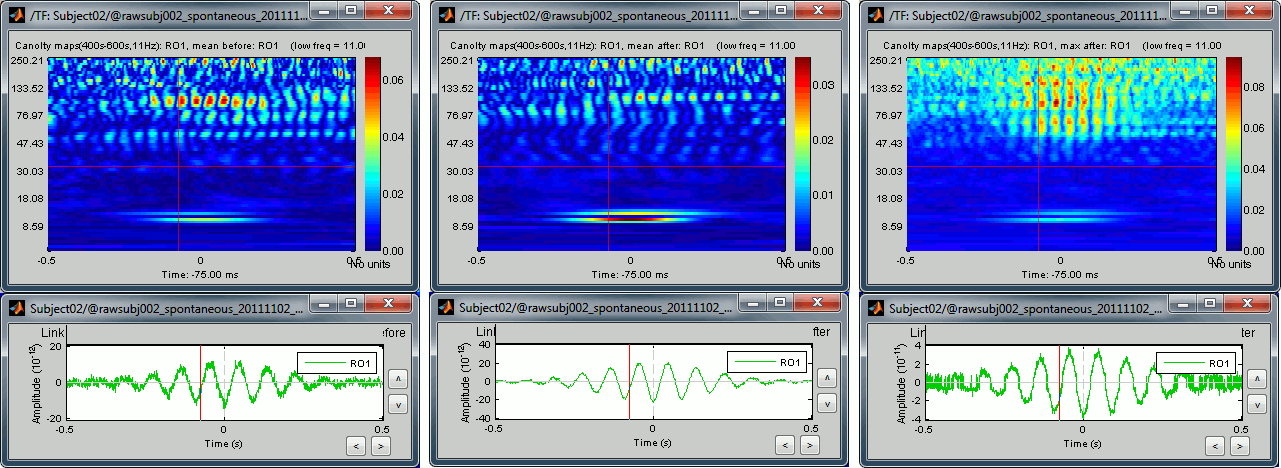
- Same conclusion as with the PAC maps: we get weaker effects if we group multiple sources together, but on the other hand we can extract one meaningful statistic that gets rid of some possibly meaningless variability. With these views, we can restrict our band of interest for the high frequencies to 90-105Hz (instead of 60-150Hz).
Other frequencies
We have calculated Canolty maps for 20 occipital sources, based on a low frequency that was showing strong phase-amplitude coupling with higher frequencies. To illustrate the link between the PAC values and the Canolty maps, let's now take a low-frequency that shows only low values of PAC. We can try 7.5Hz because this frequency bin is showing very little coupling in all the PAC maps we calculated previously.
Run again the same process "Frequency > Canolty maps" for this different low frequency:
Time window: 400s-600s
Scout function: All
Nesting frequency: 7.5Hz
None of the sources in the scout RO1 show any of the clear patterns we observed previously:
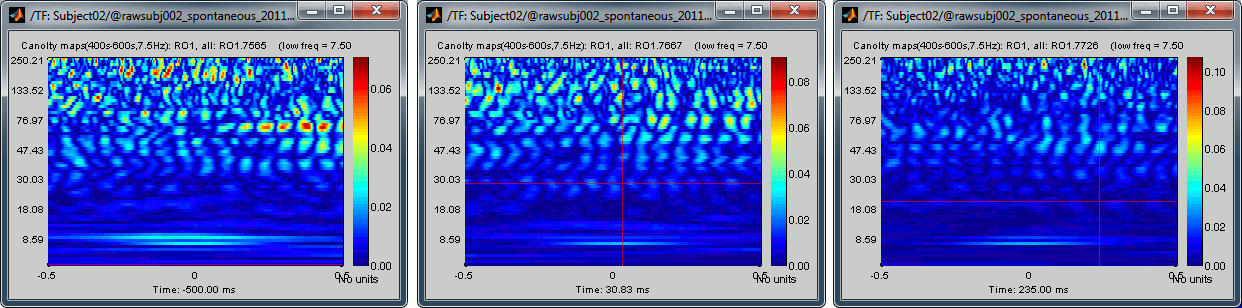
Two inputs
- In the previous example, we have identified visually the optimal low frequency on the PAC comodulograms, and we have entered this value manually in the options of the "Canolty map" process. This methodology is probably the best way to proceed when exploring manually some specific brain regions, but it will not be adapted to process a large number of signals. In the case of a full brain analysis, we cannot reproduce the same steps individually for the 15000 sources.
To automate this, a version of the "Canolty maps" process exists in another flavor, with two inputs. The second input must contains the result of a maxPAC analysis for the file to process. Drop a file of time series into File A and the corresponding maxPAC file into File B.
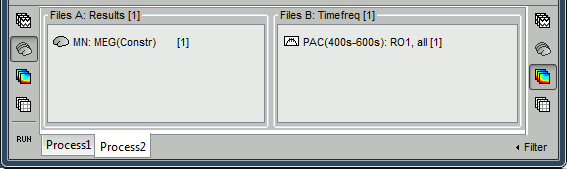
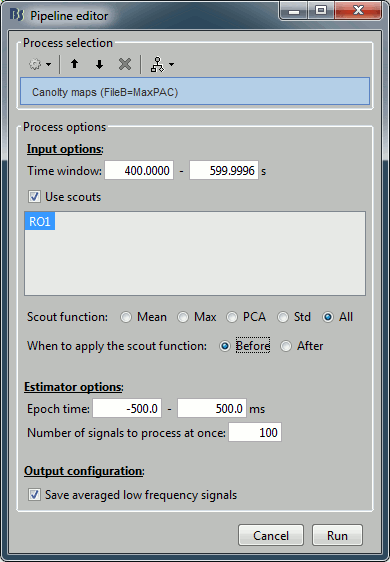
- The only difference in the process options is that there is no edit box for the low frequency. The low-frequency is directly read for each signal from the PAC file (values in the TF field).
- At the current moment, the problem with this approach is that the metric used for estimating the maxPAC is not very stable. It just takes the frequency pair that gives the maximum PAC value, and as we saw earlier in this tutorial, it is most of the time not matching the frequency we are really interested in. This automated version of the low-frequency detection is not recommended for now, but we are currently working on implementing more stable metrics. We will post the updates on these developments on this tutorial page.
Scripting
The following script from the Brainstorm distribution reproduces the analysis presented in this tutorial page: brainstorm3/toolbox/script/tutorial_resting.m
