|
Size: 5769
Comment:
|
Size: 5559
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 9: | Line 9: |
| Reviewers ''[current reviewing status] '' | Reviewers: ''[current reviewing status] '' |
| Line 11: | Line 11: |
| * '''Sylvain Baillet''': Montreal Neurological Institute ''[overview 1-14, no proofreading]'' * '''Richard Leahy''': University of Southern California ''[validated 1-19]'' * '''John Mosher''': Cleveland Clinic ''[not started]'' |
* '''Sylvain Baillet''': Montreal Neurological Institute ''[overview 1-14, edited 20-21]'' * '''Richard Leahy''': University of Southern California ''[validated 1-20]'' * '''John Mosher''': Cleveland Clinic ''[edited 22 only]'' |
| Line 15: | Line 15: |
Expected timeline (2016) * '''March''': Tutorials #1-#21 (interface and pre-processing) * '''April''': Tutorials #22-#24 (source estimation and time-frequency) * '''May-June''': Tutorials #25-#28 (statistics and workflows) * '''July-Sept''': Cross-validation of the pipeline and the results with MNE, FieldTrip and SPM * '''October''': Presentation during a satellite meeting at the Biomag 2016 conference * '''Nov-Dec''': Connectivity tutorial |
|
| Line 27: | Line 18: |
| == Tutorial 10: Power spectrum and frequency filters == | == Inverse models == === Tutorial 22: Source estimation === |
| Line 30: | Line 24: |
| * Richard, John, Sylvain: Finish the evaluation of the band-pass filters used in Brainstorm * Francois: Mark with an extended event the time segments that we cannot trust (edge effect) * Francois: Add warning if recordings are not long enough |
* John: Inverse code: Mixed head models * John: Explain the new (incorrect) results obtained with the epilepsy tutorial (see below) * John: Discuss and validate all the modifications with Alex and Matti * John: Cannot process GRAD and MAG at the same time ? * John, Richard, Sylvain: Why are dSPM values 2x lower than Z-score ? * The tutorial says "Z-normalized current density maps are also easy to interpret. They represent explicitly a "deviation from experimental baseline" as defined by the user. In contrast, dSPM indicates the deviation from the data that was used to define the noise covariance used in computing the min norm map. " * Therefore should we expect the dSPM values to deviate more from the noise recordings, than the Z-score from the pre-stim baseline? Instead of this we observe much lower values. Is there a scaling issue here? <<BR>><<BR>> {{attachment:diff_zscore_dspm.gif||width="385",height="138"}} * Francois: Make the "median" option the default? (changes a lot the range of values) * Francois: Call FieldTrip headmodels and beamformers * Francois: Add note in Elekta tutorial: Process MAG and GRAD separately |
| Line 36: | Line 37: |
| * Richard, John, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsFilter#What_filters_to_apply.3F_.5BTODO.5D|What filters to apply?]] * Richard, John, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsFilter#Filters_specifications_.5BTODO.5D|Filters specifications]] |
* John: Fix all the missing links * John, Richard, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/SourceEstimation#References_.5BTODO.5D|References]] * John, Richard, Sylvain: What do with [[http://neuroimage.usc.edu/brainstorm/Tutorials/Beamformers|Hui-Ling Beamformers]]? |
| Line 39: | Line 41: |
| == Tutorial 12: Artifact detection == [ONLINE DOC] |
=== Tutorial: Dipole scanning === * Add the description of all the measures in [[http://neuroimage.usc.edu/brainstorm/Tutorials/TutDipScan#Significance_mesures_.5BTODO.5D|this section]]. |
| Line 42: | Line 44: |
| * Beth: Adding examples of the different detection processes. Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsDetect#Other_detection_processes|Other detection processes]] | === Tutorial EEG/Epilepsy === * Why sLORETA? * Location issues with the new code: <<BR>>dSPM is now localizing the spike in a much deeper spot (top=old version, bottom=new version) <<BR>><<BR>> {{attachment:epilepsy_dspm.gif||width="272",height="233"}} * '''Marcel Heers''': "Looking at the findings from the intracranial EEG in Matthias Dümpelmann's article (figure 1 panel a) and b)) it is '''very likely the new sources are wrong'''. Additionally, the older sources are much more in agreement with Matthias' sLORETA results and with cMEM findings. Sohrabopour et al. reported as well that their IRES method found results in agreement with sources shown in the tutorial." * Imported data for testing can be downloaded here:<<BR>> https://www.dropbox.com/s/42d9indpjr8ac1y/TutorialEpilepsy.zip?dl=0 |
| Line 44: | Line 50: |
| == Tutorial 13: Artifact cleaning with SSP == | == Filters == === Tutorial 10: Power spectrum and frequency filters === |
| Line 47: | Line 54: |
| * Francois: Check the length needed to filter the recordings (after finishing #10) <<BR>>Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsSsp#SSP_Algorithm_.5BTODO.5D|SSP Algorithm]] == Tutorial 15: Import epochs == [ONLINE DOC] * Richard, Sylvain, John, Francois: Define recommendations for epoch lengths (after finishing #10) <<BR>>Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/Epoching#Epoch_length_.5BTODO.5D|Epoch length]] == Tutorial 20: Head modelling == [ONLINE DOC] * John, Richard, Sylvain: In-depth review of this sensitive tutorial * John, Richard, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/AllIntroduction#Tutorials.2FHeadModel.References_.5BTODO.5D|References]] == Tutorial 22: Source estimation == [CODE] * John: Dipole scanning: Wrong orientations * John: Inverse code: sLORETA * John: Inverse code: NAI * John: Inverse code: Mixed head models * John, Richard, Sylvain: How to normalize unconstrained sources ? * Francois: Remove the warning messages in the interface * Francois: Call FieldTrip headmodels and beamformers * All: Unconstrained: PCA vs Norm?<<BR>>http://www.ncbi.nlm.nih.gov/pubmed/25230358<<BR>>[[http://www.nature.com/neuro/journal/v15/n6/full/nn.3101.html|Hipp et al (2012)]] |
* Richard, Hossein: Define reasonable transient durations * Richard, Hossein: Describe the band-stop and notch filters in the same way * Richard, Hossein: Frequency resolution in the "freqz" plots for high-pass filters under 0.5Hz * Richard, John, Hossein, Sylvain: Weird PSD plots for Elekta recordings |
| Line 74: | Line 61: |
| * John, Richard, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/SourceEstimation#Source_estimation_options_.5BTODO.5D|Source estimation options]] * John, Richard, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/SourceEstimation#Advanced_options_.5BTODO.5D|Advanced options]] * John, Richard, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/SourceEstimation#Equations_.5BTODO.5D|Equations]] * John, Richard, Sylvain: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/SourceEstimation#References_.5BTODO.5D|References]] * John, Richard, Sylvain: Why are dSPM values 2x lower than Z-score ? * John, Richard, Sylvain: In-depth review of this sensitive tutorial |
* Richard, Hossein: Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsFilter#Filters_specifications_.5BTODO.5D|Filters specifications]] for band-stop and notch filters === Tutorial 13: Artifact cleaning with SSP === * Francois: Check the length needed to filter the recordings (after finishing #10) <<BR>>Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsSsp#SSP_Algorithm_.5BTODO.5D|SSP Algorithm]] === Tutorial 15: Import epochs === * Richard, Sylvain, John, Francois: Define recommendations for epoch lengths (after finishing #10) <<BR>>Section [[http://neuroimage.usc.edu/brainstorm/Tutorials/Epoching#Epoch_length_.5BTODO.5D|Epoch length]] === Tutorial 22: Source estimation === * Francois: Update screen capture in section [[http://neuroimage.usc.edu/brainstorm/Tutorials/SourceEstimation#Averaging_in_source_space|Averaging in source space]] |
| Line 84: | Line 75: |
| * Enable option "Hide edge effects" for Hilbert (after finishing #10) | * Francois: Enable option "Hide edge effects" for Hilbert (after finishing #10) |
| Line 88: | Line 79: |
| * Hilbert: Link back to the filters tutorial (after finishing #10) * ERS/ERD on Hilbert: Squared signal only? * Shouldn't we always use power?... Right now, even the averaging process for trials give inconsistent results across methods. == Tutorial 26: Statistics == * Not evaluated yet |
* Francois: Hilbert: Link back to the filters tutorial (after finishing #10) |
| Line 96: | Line 82: |
| * Not evaluated yet == Tutorial 28: Scripting == * Not evaluated yet |
* Add Chi2(log) ? |
| Line 102: | Line 85: |
| * Francois: Check that the script tutorial_introduction.m produces the same output as tutorials | * Francois: Remove all the wiki pages that are not used |
| Line 104: | Line 87: |
| * Francois: Check the number of pages for each tutorial | * Francois: Remove useless images from all tutorials |
| Line 106: | Line 89: |
| * Francois: Remove useless images from all tutorials <<BR>><<BR>><<BR>><<BR>><<BR>>[Additional important stuff] == Phantom tutorial (for validation) == * John: Prepare Elekta recordings to distribute on the [[Tutorials/PhantomElekta|tutorial page]] * Francois: Prepare the Elekta phantom anatomy * Francois: Prepare the analysis script * Francois: Edit the tutorial page == Other analysis scenarios == * Francois: Update all the tutorials (100+ pages) * Francois: Add number of pages in the tutorials |
|
| Line 123: | Line 93: |
| * Richard: How to assess significance from connectivity matrices * Francois: Preparation of a tutorial |
* Richard: How to deal with unconstrained sources?<<BR>> http://neuroimage.usc.edu/forums/showthread.php?2401 * Richard: How to assess significance from connectivity matrices? * Francois, Richard, Hossein: Preparation of a tutorial |
Introduction tutorials: Editing process
http://neuroimage.usc.edu/brainstorm/Tutorials
Redactors:
Francois Tadel: Montreal Neurological Institute
Elizabeth Bock: Montreal Neurological Institute
Reviewers: [current reviewing status]
Sylvain Baillet: Montreal Neurological Institute [overview 1-14, edited 20-21]
Richard Leahy: University of Southern California [validated 1-20]
John Mosher: Cleveland Clinic [edited 22 only]
Dimitrios Pantazis: Massachusetts Institute of Technology [validated 1-15]
Inverse models
Tutorial 22: Source estimation
[CODE]
- John: Inverse code: Mixed head models
- John: Explain the new (incorrect) results obtained with the epilepsy tutorial (see below)
- John: Discuss and validate all the modifications with Alex and Matti
- John: Cannot process GRAD and MAG at the same time ?
- John, Richard, Sylvain: Why are dSPM values 2x lower than Z-score ?
- The tutorial says "Z-normalized current density maps are also easy to interpret. They represent explicitly a "deviation from experimental baseline" as defined by the user. In contrast, dSPM indicates the deviation from the data that was used to define the noise covariance used in computing the min norm map. "
Therefore should we expect the dSPM values to deviate more from the noise recordings, than the Z-score from the pre-stim baseline? Instead of this we observe much lower values. Is there a scaling issue here?
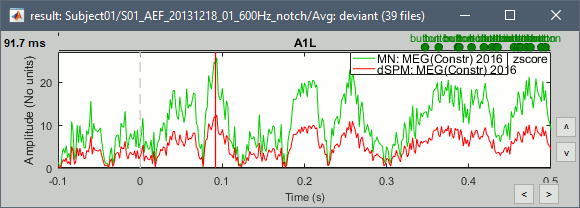
- Francois: Make the "median" option the default? (changes a lot the range of values)
Francois: Call FieldTrip headmodels and beamformers
- Francois: Add note in Elekta tutorial: Process MAG and GRAD separately
[ONLINE DOC]
- John: Fix all the missing links
John, Richard, Sylvain: Section References
John, Richard, Sylvain: What do with Hui-Ling Beamformers?
Tutorial: Dipole scanning
Add the description of all the measures in this section.
Tutorial EEG/Epilepsy
- Why sLORETA?
Location issues with the new code:
dSPM is now localizing the spike in a much deeper spot (top=old version, bottom=new version)
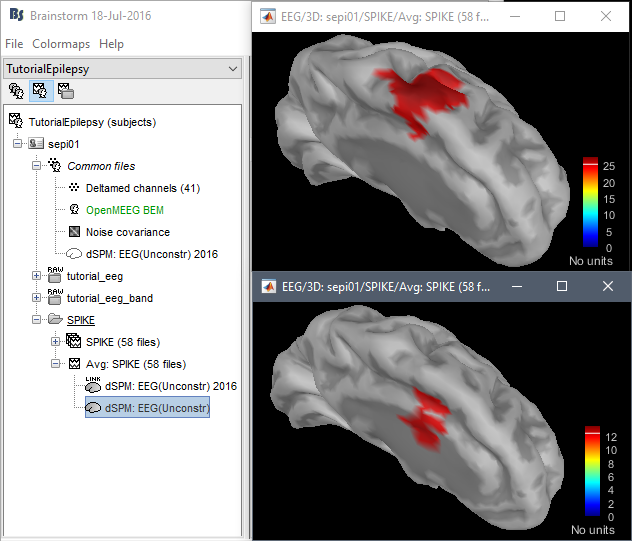
Marcel Heers: "Looking at the findings from the intracranial EEG in Matthias Dümpelmann's article (figure 1 panel a) and b)) it is very likely the new sources are wrong. Additionally, the older sources are much more in agreement with Matthias' sLORETA results and with cMEM findings. Sohrabopour et al. reported as well that their IRES method found results in agreement with sources shown in the tutorial."
Imported data for testing can be downloaded here:
https://www.dropbox.com/s/42d9indpjr8ac1y/TutorialEpilepsy.zip?dl=0
Filters
Tutorial 10: Power spectrum and frequency filters
[CODE]
- Richard, Hossein: Define reasonable transient durations
- Richard, Hossein: Describe the band-stop and notch filters in the same way
- Richard, Hossein: Frequency resolution in the "freqz" plots for high-pass filters under 0.5Hz
- Richard, John, Hossein, Sylvain: Weird PSD plots for Elekta recordings
[ONLINE DOC]
Richard, Hossein: Section Filters specifications for band-stop and notch filters
Tutorial 13: Artifact cleaning with SSP
Francois: Check the length needed to filter the recordings (after finishing #10)
Section SSP Algorithm
Tutorial 15: Import epochs
Richard, Sylvain, John, Francois: Define recommendations for epoch lengths (after finishing #10)
Section Epoch length
Tutorial 22: Source estimation
Francois: Update screen capture in section Averaging in source space
Tutorial 24: Time-frequency
[CODE]
- Francois: Enable option "Hide edge effects" for Hilbert (after finishing #10)
[ONLINE DOC]
- Francois: Hilbert: Link back to the filters tutorial (after finishing #10)
Tutorial 27: Workflows
- Add Chi2(log) ?
Final steps
- Francois: Remove all the wiki pages that are not used
- Francois: Check all the links in all the pages
- Francois: Remove useless images from all tutorials
Francois: Reference on ResearchGate, Academia and Google Scholar
http://neuroimage.usc.edu/brainstorm/Tutorials/AllIntroduction
Connectivity
- Not documented at all
- Richard, Sylvain: Define example dataset and precise results to obtain from them
Richard: How to deal with unconstrained sources?
http://neuroimage.usc.edu/forums/showthread.php?2401- Richard: How to assess significance from connectivity matrices?
- Francois, Richard, Hossein: Preparation of a tutorial
